Markedly different gene expression in wheat grown with organic or inorganic fertilizer (original) (raw)
Abstract
Nitrogen is the major determinant of crop yield and quality and the precise management of nitrogen fertilizer is an important issue for farmers and environmentalists. Despite this, little is known at the level of gene expression about the response of field crops to different amounts and forms of nitrogen fertilizer. Here we use expressed sequence tag (EST)-based wheat microarrays in combination with the oldest continuously running agricultural experiment in the world to show that gene expression is significantly influenced by the amount and form of nitrogenous fertilizer. In the Broadbalk winter wheat experiment at Rothamsted in the United Kingdom and at three other diverse test sites, we show that specific genes have surprisingly different expression levels in the grain endosperm when nitrogen is supplied either in an organic or an inorganic form. Many of the genes showing differential expression are known to participate in nitrogen metabolism and storage protein synthesis. However, others are of unknown function and therefore represent new leads for future investigation. Our observations show that specific gene expression is diagnostic for use of organic sources of nitrogen fertilizer and may therefore have useful applications in defining the differences between organically and conventionally grown wheat.
Keywords: transcriptomics, microarrays, organic farming, wheat, nitrogen
1. Introduction
Organic agriculture, including the growing of organic wheat, is one of the fastest-growing sectors in the food industries of Europe and the United States (Greene & Dobbs 2001). Strict rules apply to the growing of organic crops including a ban on the use of artificial nitrogenous fertilizers; nitrogen inputs must come from animal manures and slurries, green manures, crop residues, legumes and residual soil organic matter. Advocates for organic farming point to savings in fossil fuels, reduced environmental pollution, greater biodiversity and more nutritious and better tasting food as positive reasons for farming organically. Opponents counter with reduced yields and quality, and question the claimed environmental and health benefits (Trewavas 2004). Despite the increasing interest in organic wheat, nothing is known at the molecular level about differences between grain grown solely with organic fertilizer compared to grain grown under conventional inorganic conditions. Grain nitrogen levels depend on two factors: continued nitrate uptake by roots and the remobilization of nitrogenous compounds from leaves and stems during grain filling (Masclaux et al. 2001). The proportion of nitrogen from these two sources is greatly influenced by environmental conditions and plant genotype (Purcino et al. 1998). In addition to the readily available ammonium and nitrate ions, organic fertilizers can contain a wide range of organic-nitrogen compounds such as amino acids, amino sugars and nitrogen heterocyclic compounds which may influence the soil microflora (Mäder et al. 2002; Shannon et al. 2002; Pernes-Debuyser & Tessier 2004). We examined the gene expression profiles of developing wheat grain grown under different nitrogen fertilizer regimes on Broadbalk field at Rothamsted in South East England (figure 1) to determine whether these profiles might be diagnostic for these regimes and prognostic for yield. On Broadbalk, winter wheat is grown under well defined nutrient supply conditions including a range of inorganic nitrogen treatments and two treatments based on farmyard cattle manure (FYM; Poulton 1996; Richards & Webster 1999; http://www.rothamsted.bbsrc.ac.uk/resources/ClassicalExperiments.html).
Figure 1.
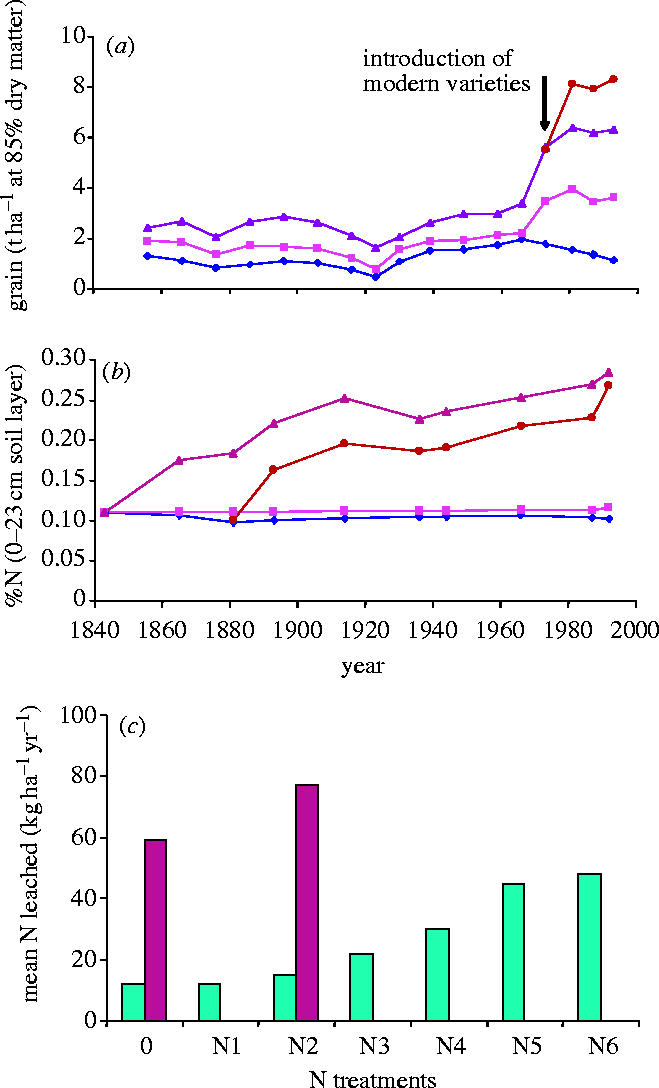
Historical data describing the Broadbalk winter wheat experiment which was established at Rothamsted, England in 1843 by Sir John Bennet Lawes and Sir Henry Gilbert to investigate the influence of inorganic fertilizers and organic manures on crop yield. Various combinations of nitrogen, phosphorus, potassium, magnesium and organic manures have been applied continuously since 1843 giving plots with distinctive nutrient availability profiles. Crop yields have been recorded and plant and soil samples retained since the start of the experiment, providing a unique archive of over 160 years of winter wheat production at this site. (a) Grain yields (pre-1968 data presented are 10 year averages, whereas post-1968 it is an average of the variety used during that period) 1852–1995 (t ha−1 at 85% dry matter): (blue diamond) N0; (red square) N1; (purple triangle) FYM; (brown circle) FYM since 1885+N2 since 1968 (adapted from Poulton 1995). (b) %N in 0–23 cm soil, 1843–1992, symbols as for (a; from Garner & Dyke 1969 and unpublished data). (c) Mean amount of N leached (kg ha−1 yr−1) 1990–1998: different amounts of inorganic N (blue histograms); FYM+different amounts of inorganic N (purple histograms; from Goulding et al. 2000).
2. Methods
(a) Growth of plants and tissue collection
The bread-making wheat cultivar Hereward was grown in 2002, 2003 and 2004 on the Broadbalk winter wheat experiment at Rothamsted in South East England. Grain samples were taken from duplicate plots of continuous winter wheat on seven nitrogen treatments: N0, N1, N1 : 2 : 1, N4, N6, FYM and FYM+N2, where N, is ammonium nitrate in increments of 48 kg N ha−1. Nitrogen was applied in a single dressing in mid-April, except for N1 : 2 : 1, which was applied in three splits in March, April and May. FYM was applied in the autumn at a rate of 35 t ha−1 (which exceeds the levels used in standard organic practice); this contained approximately 250 kg ha−1 of total-N predominantly in organic form. The fertilizer rate N4 (192 kg ha−1) is the typical rate used by UK wheat farmers. None of the plots were limiting for P, K and Mg. For each treatment, 100 plants were harvested per sample from three random areas in each of two duplicate plots. The same cultivar, Hereward, was also sampled from six plots of the ley–arable experiment at Woburn, Bedfordshire in 2004. Samples were collected from two plots given either a split or single dressing of inorganic nitrogen fertilizer (40+170 kg N ha−1 and 210 kg N ha−1), from duplicate plots previously in a clover/grass ley for 8 years and given no inorganic nitrogen and from duplicate plots in a clover/grass ley for 3 years also given no inorganic nitrogen. Wheat samples were also taken from two further farms in 2004; at a farm near Harpenden (UK), cv. Tybalt spring wheat was grown following a lupin crop. This site received annual organic inputs of compost and green manures. Finally crops were sampled from a farm in Shanxi Province (China) in 2004. Here winter wheat, cv. Yang Mei 12 was grown with either 8 t ha−1 pig manure and 7 t ha−1 chicken manure (containing ∼107 kg nitrogen) or 225 kg ha−1 inorganic nitrogen in the form of ammonium nitrate. For all plots all plants were tagged at the start of anthesis and sampled 14 days post-anthesis (dpa) when the grain was at the ‘soft dough’ stage of development.
(b) Expression analysis
For all experiments, total RNA was extracted from six biological replicates (two duplicate plots, three samples per plot). Each sample consisted of 500-pooled grains collected from the middle spikelets of the ear of the main stem (10–15 grains per ear). Tissue samples were stored in liquid nitrogen and RNA was isolated as described previously (Wilson et al. 2004).
All cDNA clones used for the microarray experiments were from the University of Bristol wheat unigene set (Wilson et al. 2004). All sequences within the unigene set have been placed in GenBank. Inserts were prepared and arrayed onto CodeLink microarray slides (Amersham BioSciences/GE Healthcare, Chalfont, UK) using standard procedures in conjunction with a BioRobotics MicroGrid II Microarrayer (Genomic Solutions Ltd, Huntingdon, UK). Fluorescently labelled cDNA probes were prepared from the various total RNA samples using an amino allyl-labelled first-strand reverse transcriptase reaction (Wilson et al. 2004).
(c) Data normalization and quantitative PCR verification
The array data from GenePix Pro (Molecular Devices Corporation, Union City, CA, USA) was normalized using actin (AL825219), and similar results were obtained using tubulin (AL821639; data not shown). Data from each of the two colour dye-swap experiments was normalized using GeneSpring 6.1 (Silicon Genetics, Redwood City, CA, USA) by application of a per spot and per chip intensity-dependent (Lowess) approach. Differentially expressed genes were identified by application of a one way analysis of variance (p<0.05) test. Cluster analysis using Pearson's correlation was based on a similarity matrix calculated from Euclidean distances between genes across conditions using expression values with distances d converted to similarities s as _s_=1/(1+d). Individual candidate genes identified as having expression profiles dependent on the treatment were further analysed by Q-PCR using the LightCycler System (Roche Diagnostics, Lewes, UK). The methods used were according to the manufacturers instructions (QuantiTectTM SYBR Green PCR, Qiagen) based on Dewitte et al. (2003). There were three biological replicates for each treatment.
3. Results and discussion
(a) Global patterns of grain gene expression in response to organic (FYM) and inorganic sources of nitrogen
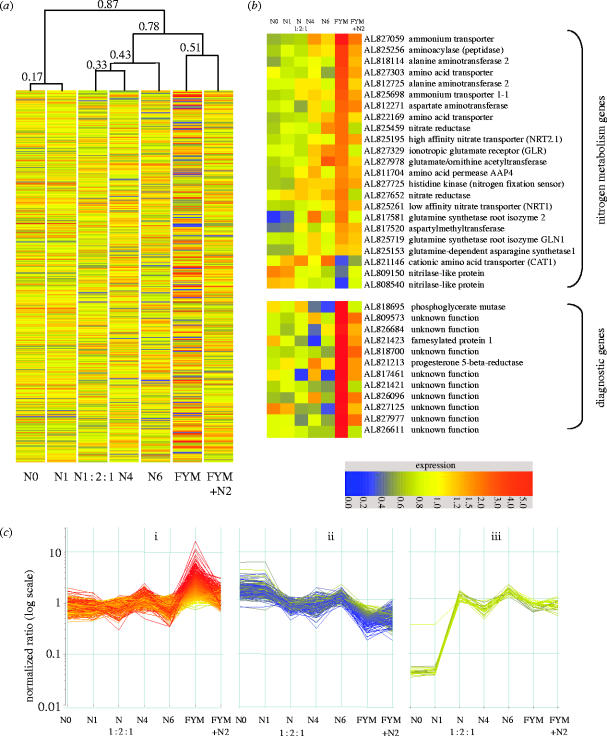
To measure changes in gene expression in developing wheat grain, we hybridized replicate wheat EST-microarrays containing 10 000 features (Wilson et al. 2004) with fluorescently labelled cDNAs derived from endosperm at 14 dpa harvested in 2002 from Broadbalk. Cluster analysis of the global gene expression profiles distinguished grain from plants provided with 0–288 kg ha−1 of inorganic nitrogen, applied either as a single application (N0–N6), or in a 1 : 2 : 1 split application (N1 : 2 : 1) chosen to mimic a slow release scenario, with FYM only or with FYM supplemented with 96 kg inorganic nitrogen (FYM+N2). The global expression patterns were characteristic for each of the treatments (figure 2). Greatest similarity was evident within each of three types of treatments: low nitrogen (N0 and N1), high inorganic nitrogen (N4, N1 : 2 : 1 and N6) and the FYM treatments (figure 2a). This pattern of clustering reflected soil characteristics of the plots: FYM plots had the highest soil organic carbon and total-nitrogen contents, while the N0 and N1 plots had the lowest. The N4, N1 : 2 : 1 and N6 plots had intermediary values for these characteristics (table 1). When global expression patterns for grain derived from FYM and N4 plots in 2002, 2003 and 2004 were compared, the same treatments for the different years clustered together (Electronic Appendix supporting figure 1).
Figure 2.
Gene expression in response to different amounts and forms of nitrogen as determined by microarray analysis. Grain samples were taken from duplicate plots of continuous winter wheat on seven nitrogen treatments: N0, N1, N1 : 2 : 1, N4, N6, FYM and FYM+N2, where N, is ammonium nitrate in increments of 48 kg N ha−1. Nitrogen was applied as a single dressing in mid-April, except N1 : 2 : 1, which was applied in three splits in March, April and May. FYM was applied in the autumn at a rate of 35 t ha−1; this contains approximately 250 kg ha−1 of total-N predominantly in organic form. (a) Hierarchical clustering was used to order the microarray expression profile data from 10 000 wheat unigene ESTs in response to different nitrogen treatments. Values for similarity are indicated. The data presented are the product of at least 12 replicates. (b) The expression profiles of nitrogen metabolism and diagnostic genes. Expression levels are depicted by the colour (see key). GenBank accession numbers (array sequences) and representative annotation are given. (c) K-mean cluster analysis of three different expression profiles. In all cases the data generated are presented following normalization to the expression of actin.
Table 1.
Organic carbon and total nitrogen in Broadbalk and Woburn soils.
| Broadbalk | Woburn | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| N0 | N1 | N1 : 2 : 1 | N4 | N6 | FYM | FYM+N2 | arable | 3y ley | 8y ley | |
| organic C (%) | 0.91 | 1.07 | 1.24 | 1.16 | 1.24 | 2.92 | 2.69 | 0.85 | 1.18 | 1.02 |
| total N (%) | 0.092 | 0.106 | 0.122 | 0.117 | 0.123 | 0.277 | 0.256 | 0.074 | 0.104 | 0.088 |
(b) Expression of specific genes in response to organic and inorganic sources of nitrogen
The expression profiles for two categories of genes are presented: genes encoding proteins involved in nitrogen metabolism and genes, which on the basis of their expression profiles, appear to be markers for the FYM treatment (figure 2b), referred to as ‘diagnostic’ genes. As expected, increased expression of several genes involved in nitrogen metabolism was observed in the higher nitrogen treatments. For many of these, greatest expression was seen in the FYM and FYM+N2 treatments. Induction of nitrate, ammonium and amino acid transporters and nitrate reductase has been reported in array experiments for tomato (Wang et al. 2001) and Arabidopsis (Wang et al. 2000, 2003). A notable down-regulation of two nitrilases was observed in the FYM treatment; nitrilases are involved in indole-3-acetic acid biosynthesis (Normanly et al. 1997).
The 12 diagnostic genes in figure 2b had expression profiles up-regulated in the presence of FYM. For the purpose of this analysis only those genes that showed greater than fivefold up-regulation in the FYM compared to the N4 treatment are presented. Of the 12 genes in this category, nine could not be assigned a function. Of the three sequences for which a function could be assigned no overall link could be identified. One gene (Al818695; phosphoglycerate mutase) inter-converts 3-phosphoglycerate and 2-phosphoglycerate and is one of the many steps in glycolysis (Grana et al. 1992), while a second (AL821423) shows sequence similarities to a stress induced farnesylated protein isolated from barley (Barth et al. 2004). A third sequence, AL821213, shows similarities to a progesterone 5-beta-reductase (BAC20032), which catalyses the reduction of progesterone to beta-pregnane-3, 20-dione.
The cluster analysis enabled the classification of groups of genes with similar expression profiles across the various treatments. In figure 2c we present three representative profiles. The first group (figure 2c(i) and Electronic Appendix supporting table 1) contained 474 genes with highest expression in the FYM treatment and included the diagnostic genes. Curiously, for this group the FYM+N2 treatments often resulted in a decrease in expression. One hundred and fifty-five of the genes within this group had insufficient sequence similarity with other sequences to assign a putative function. However, the remaining 319 genes encoded a wide variety of proteins including proteins known to be involved in amino acid, lignin and flavonoid biosynthesis and glycolysis. The second group of 190 genes (figure 2c(ii) and Electronic Appendix supporting table 1) showed highest expression in N0 and N1 and lowest expression in the FYM treatment. A third group consisting of twenty genes showed low expression in the low nitrogen treatments (N0 and N1) and uniformly high expression at higher nitrogen treatments irrespective of the form of nitrogen (figure 2c(iii) and Electronic Appendix supporting table 1).
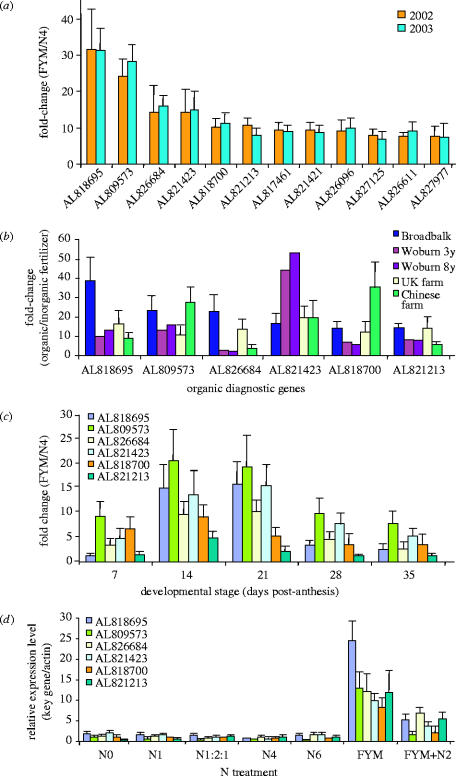
The expression profiles of the 12 diagnostic genes were further verified by quantitative-RT-PCR (Q-PCR) using RNA derived from material grown on Broadbalk during 2002, 2003 or 2004 (figure 3). The expression of the 12 diagnostic genes during 2002 and 2003 is shown (figure 3a). Of these, six genes showed greater than 10 fold change in the level of expression in the FYM versus N4 treatments on Broadbalk in 2002. The expression of these six diagnostic genes was further examined on four separate farms during 2004. In all four locations these six genes showed higher expression in response to a variety of organic inputs compared to controls that had received inorganic nitrogen (figure 3b).
Figure 3.
Expression profiles of the ‘diagnostic genes’ determined by Q-PCR. (a) Analysis of the 12 sequences with the highest ratio of expression following FYM treatment when compared to N4 treatment (data from figure 2). (b) Q-PCR analysis of the six highest ratio of expressions sequences from ‘a’, using samples from four sites during 2004: Broadbalk, Woburn farm (winter wheat after 3 or 8 years clover/grass ley compared to 210 kg N ha−1), a certified organic wheat crop from the Harpenden area, UK (cv. Tybalt spring wheat, compared to Broadbalk N4) and an uncertified organic wheat crop from Shanxi Province, China (winter wheat, cv. Yang Mei 12 compared to plots with 225 kg N ha−1). (c) Relative expression (FYM versus N4) of the six diagnostic genes during the five week period post anthesis in 2004. (d) Relative expression levels, compared to actin, of the six diagnostic genes in the Broadbalk treatments. All data are means±s.d.
To examine if these six genes were more highly expressed in FYM versus N4 treatments throughout grain development, we determined their expression profile on Broadbalk in the five-week period following anthesis (figure 3c). All six genes showed a continuously elevated level of expression in the FYM treatment compared to the N4 treatment with the maximum level of differential expression observed at 14 and 21 dpa. This time period is significant as it represents the early/middle stage of grain filling and reserve synthesis. Finally, we examined the specificity of expression of these six genes across the Broadbalk treatments (figure 3d). Our Q-PCR results showed that the six diagnostic genes were highly expressed in the FYM treatment, moderately expressed in the FYM+N2 treatment, and had lower expression in all other treatments, confirming the microarray data (figure 2). Taken together, the data indicates that these six genes were expressed most highly when the crop had received FYM or other organic inputs.
(c) Expression of storage protein genes
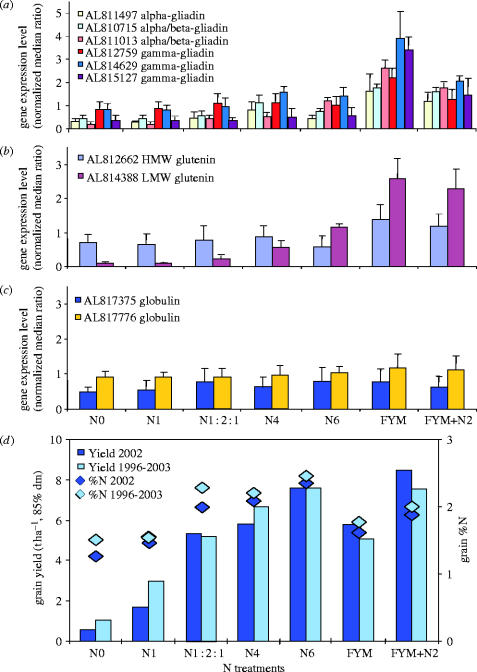
The amount and composition of grain proteins that contribute to grain quality are of practical importance to farmers and millers. As previous studies have shown that the level of storage protein transcripts are an accurate indicator of storage protein levels in the developing grain (Bartel & Thompson 1986; Sorensen et al. 1989), we examined the levels of various transcripts encoding gliadin, glutenin and globulin storage proteins in the Broadbalk treatments (figure 4a_–_c). Our results indicated that in common with previous reports (Wieser & Seilmeier 1998), the expression of gliadins (figure 4a), low molecular weight (LMW) glutenins (figure 4b) and high molecular weight (HMW) glutenins (figure 4b) increased with higher inorganic nitrogen levels, whereas globulin transcripts (figure 4c) were hardly affected by the level of nitrogen input. Interestingly, the application of FYM appeared to further increase the expression of many of the gliadin transcripts even though the FYM treatment contained less nitrogen than the N6 treatment. As for the previously described diagnostic genes, application of FYM+N2 resulted in reduced expression of some of the gliadin transcripts, whereas there was little reduction in the level of the glutenin transcripts. This observation suggests that the gliadin/glutenin ratio would differ between organically grown and inorganically grown wheat with the potential to influence the properties of the resulting flour.
Figure 4.
Q-PCR results for various storage protein sequences. Q-PCR results for the storage-protein sequences including six gliadins (a), one high molecular weight and one low molecular weight glutenin (b) and two globulin related sequences (c). In each case the expression levels of the various storage protein genes are presented as a ratio of the transcript levels to the actin transcript levels. All data are means±s.d. (d), Grain yield and grain %N under different nitrogen treatments for Broadbalk winter wheat (cv. Hereward). The results from 2002 are compared with the mean data for the period in which cv. Hereward has been grown on Broadbalk (1996–2003). The N1 : 2 : 1 treatment was introduced in 2001. The data are presented as a mean of two replicate plots grown under continuous wheat.
The most important factor for farmers is grain yield. The relatively low nitrogen availability in organic farming systems limits yields and grain nitrogen and it is therefore difficult to produce high-protein grain under organic conditions (Gooding et al. 1993). To examine this further we compared the various expression profiles seen in figure 2 with the final grain yields and the percentage of nitrogen in grain dry matter for the corresponding treatments on Broadbalk (figure 4d). In the N0 and N1 treatments, whose gene expression profiles clustered together, total nitrogen uptake, yield and grain nitrogen were all relatively low. Increasing the nitrogen rate (N4, N1 : 2 : 1 and N6) resulted in similar gene expression profiles and increased nitrogen uptake, yield and grain nitrogen. The highest levels of grain nitrogen followed the N6 treatment and this treatment showed the greatest degree of divergence at the expression level from the other inorganic nitrogen treatments. The global gene expression clustering suggested a close relationship between the FYM and FYM+N2 treatments, discrete from the inorganic treatments alone or the N0 treatment, which could be regarded as an organic-N plot since it had not received additional inputs of organic or inorganic nitrogen.
Our results demonstrate that the amount and form of nitrogen available to wheat influences gene expression and grain yield. It is also important to consider what additional factors in the FYM treatments other than the form of nitrogen fertilizer may have contributed to the expression patterns observed. For instance, the soil of the FYM plots had higher organic matter content (table 1), which will have lead to improved water retention properties (Mäder et al. 2002; Pernes-Debuyser & Tessier 2004). However, in the FYM treatments described here inorganic nitrogen application (FYM+N2) down-regulated many of the genes highly expressed in the FYM treatment alone. This suggests that the differences observed between the organic and inorganic nitrogen treatments are not simply due to the increased organic matter content and associated changes in properties such as water retention or soil microbial populations. It is interesting to note that even though many of the genes, including some involved in nitrogen metabolism, were down-regulated in FYM+N2, yields were increased substantially compared to the FYM treatments and final grain nitrogen increased slightly (figure 4d). Our observations suggest that although many genes are expressed at higher levels in fertile organic conditions, their synergistic effect does not result in higher yields or increased grain nitrogen content. It is possible that in both the organic-N treatments and at the higher inorganic nitrogen inputs (N4, N6), the nitrogen in the soil during seedling establishment and the early tillering stages is sufficient to create a high yield potential for the crop. However, in the treatments with solely organic-N inputs (FYM), but not in the inorganic-N treatments or supplemented organic-N treatments (FYM+N2), nitrogen subsequently becomes limiting and in this situation many genes involved in nitrogen uptake, nitrogen metabolism and storage are up-regulated in an attempt to fulfil the initial yield potential. Such a scenario would be consistent with our observation that the addition of inorganic nitrogen to the FYM treated plots resulted in a slightly higher yield and a decrease in the expression level of many of the genes involved in nitrogen metabolism.
4. Conclusion
Numerous reports have discussed the environmental implications and the benefits of consuming organically grown food. Clearly the optimal use of nitrogenous fertilizers will reduce environmental pollution, however there is little evidence that nitrate leaching differs between organic and conventional production systems when following best practice (Stopes et al. 2002; Trewavas 2002). Our results suggest that at the level of gene expression, wheat grain from plants supplied with either organic or inorganic nitrogen is reproducibly different. However, our results also confirm that the limited availability of nitrogen in organic systems tends to limit yields and total grain nitrogen. In this study we have identified several genes whose expression appears to be indicative of the ‘organic status’ of the wheat grain in relation to the type of nitrogenous fertilizer. Such genes, if shown to be causally related to utilisation of organically originating nitrogen, might represent useful starting points for the targeted breeding of varieties that perform better under organic growing conditions. However, we believe that the large number of genes shown to be up-regulated in response to organic nitrogen fertilizer and induced by increased nitrogen supply indicate the complexity of the physiological response and require further experiments to dissect the effects that both the amount and form of nitrogenous fertilizer have on the developing grain.
5. Data deposition
The sequences reported in this paper have been deposited in the GenBank database (accession nos. AL808216–AL831324). The gene expression data have been deposited in the ArrayExpress database in accordance with Microarray Gene Expression Data Society recommendations (accession numbers A-MEXP-151 and E-MEXP-261; http://www.ebi.ac.uk/arrayexpress/Submissions/ index.html).
Acknowledgments
We thank W. Zhou, S. Parmar, S. Shepherd, C. Shepherd, J. Jones, C. Knight, K. Yu, S. White and M. Bond for technical assistance. We thank Howard Roberts for providing access to the Harpenden spring wheat crop during 2004. Rothamsted Research receives grant-aided support from the Biotechnology and Biological Science Research Council (BBSRC) of the UK. This project was supported by a grant from the BBSRC to M.J.H. and K.J.E.
Supplementary Material
References
- Bartel D, Thompson R.D. Synthesis of messenger-RNAs coding for abundant endosperm proteins during wheat grain development. Plant Sci. 1986;46:117–125. [Google Scholar]
- Barth O, Zschiesche W, Siersleben S, Humbeck K. Isolation of a novel cDNA encoding a nuclear protein involved in stress response and leaf senescence. Physiol. Plantarum. 2004;121:282–293. doi: 10.1111/j.0031-9317.2004.00325.x. [DOI] [PubMed] [Google Scholar]
- Dewitte W, Riou-Khamlichi C, Scofield S, Healy J.M.S, Jacqmard A, Kilby N.J, Murray J.A.H. Altered cell cycle distribution, hyperplasia, and inhibited differentiation in Arabidopsis caused by the D-type cyclin CYCD3. Plant Cell. 2003;15:79–92. doi: 10.1105/tpc.004838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner H.V, Dyke G.V. Rothamsted Experimental Station Report for 1968, Part 2. Lawes Agricultural Trust; Harpenden, UK: 1969. The Broadbalk yields; pp. 26–49. [Google Scholar]
- Gooding M.J, Davies W.P, Thompson A.J, Smith S.P. The challenge of achieving breadmaking quality in organic and low input wheat in the UK—a review. Asp. Appl. Biol. 1993;36:189–198. [Google Scholar]
- Goulding K.W.T, Poulton P.R, Webster C.P, Howe M.T. Nitrate leaching from the Broadbalk wheat experiment, Rothamsted, UK, as influenced by fertilizer and manure inputs and the weather. Soil Use Manage. 2000;16:244–250. [Google Scholar]
- Grana X, de Lecea L, el-Maghrabi M.R, Urena J.M, Caellas C, Carreras J, Puigdomenech P, Pilkis S.J, Climent F.J. Cloning and sequencing of a cDNA encoding 2,3-biophosphoglycerate-independent phosphoglycerate mutase from maize. Possible relationship to the alkaline phosphatase family. J. Biol. Chem. 1992;267:12 797–12 803. [PubMed] [Google Scholar]
- Greene C, Dobbs T. Wheat situation and outlook yearbook WHS-2001. Economic Research Service/USDA; Washington, DC: 2001. Organic wheat production in the United States: expanding markets and supplies; pp. 31–37. [Google Scholar]
- Mäder P, Fliessbach A, Dubois D, Gunst L, Fried P, Niggli U. Soil fertility and biodiversity in organic farming. Science. 2002;296:1694–1697. doi: 10.1126/science.1071148. [DOI] [PubMed] [Google Scholar]
- Masclaux C, Quillere I, Gallais A, Hirel B. The challenge of remobilisation in plant nitrogen economy. A survey of physio-agronomic and molecular approaches. Ann. Appl. Biol. 2001;138:69–81. [Google Scholar]
- Normanly J, Grisafi P, Fink G.R, Bartel B. Arabidopsis mutants resistant to the auxin effects of indole-3-acetontrile are defective in the nitrilase encoded by the NIT1 gene. Plant Cell. 1997;9:1781–1790. doi: 10.1105/tpc.9.10.1781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pernes-Debuyser A, Tessier D. Soil physical properties affected by long-term fertilization. Eur. J. Soil Sci. 2004;55:505–512. [Google Scholar]
- Poulton P.R. The importance of long-term trials in understanding sustainable farming systems: the Rothamsted experience. Aust. J. Exp. Agric. 1995:825–834. [Google Scholar]
- Poulton P.R. The Rothamsted long-term experiments: are they still relevant? Can. J. Plant Sci. 1996;76:559–571. [Google Scholar]
- Purcino A.A.C, Arellano C, Athwal G.S, Huber S.C. Nitrate effect on carbon and nitrogen assimilating enzymes of maize hybrids representing seven eras of breeding. Maydica. 1998;43:83–94. [Google Scholar]
- Richards J.E, Webster C.P. Denitrification in the subsoil of the Broadbalk continuous wheat experiment. Soil Biol. Biochem. 1999;31:747–755. [Google Scholar]
- Shannon D, Sen A.M, Johnson D.B. A comparative study of the microbiology of soils managed under organic and conventional regimes. Soil Use Manage. 2002;18(Suppl. S1):274–283. [Google Scholar]
- Sorensen M.B, Cameron-Mills V, Brandt A. Transcriptional and post-transcriptional regulation of gene expression in developing barley endosperm. Mol. Gen. Genet. 1989;217:195–201. [Google Scholar]
- Stopes C, Lord E.I, Philipps L, Woodward L. Nitrate leaching from organic farms and conventional farms following best practice. Soil Use Manage. 2002;18(Suppl. S1):256–263. [Google Scholar]
- Trewavas A. A critical assessment of organic farming-and-food assertions with particular respect to the UK and the potential environmental benefits of no-till agriculture. Crop Prot. 2004;23:757–781. [Google Scholar]
- Wang R, Guegler K, LaBrie S.T, Crawford N.M. Genome analysis of a nutrient response in Arabidopsis reveals diverse expression patterns and novel metabolic and potential regulatory genes induced by nitrate. Plant Cell. 2000;12:1491–1509. doi: 10.1105/tpc.12.8.1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y.H, Garvin D.F, Kochian L.V. Nitrate-induced genes in tomato roots. Array analysis reveals novel genes that may play a role in nitrogen nutrition. Plant Physiol. 2001;127:345–359. doi: 10.1104/pp.127.1.345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang R, Okamoto M, Xing X, Crawford N. Microarray analysis of the nitrate response in Arabidopsis roots and shoots reveals over 1,000 rapidly responding genes and new linkages to glucose, trehalose-6-phosphate, iron, and sulfate metabolism. Plant Physiol. 2003;132:556–567. doi: 10.1104/pp.103.021253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wieser H, Seilmeier W. The influence of nitrogen fertilisation on quantities and proportions of different protein types in wheat flour. J. Sci. Food Agric. 1998;76:49–55. [Google Scholar]
- Wilson I.D, et al. A transcriptomics resource for wheat functional genomics. Plant Biotechol. J. 2004;2:495–506. doi: 10.1111/j.1467-7652.2004.00096.x. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.