A full-length enriched cDNA library and expressed sequence tag analysis of the parasitic weed, Striga hermonthica (original) (raw)
Abstract
Background
The obligate parasitic plant witchweed (Striga hermonthica) infects major cereal crops such as sorghum, maize, and millet, and is the most devastating weed pest in Africa. An understanding of the nature of its parasitism would contribute to the development of more sophisticated management methods. However, the molecular and genomic resources currently available for the study of S. hermonthica are limited.
Results
We constructed a full-length enriched cDNA library of S. hermonthica, sequenced 37,710 clones from the library, and obtained 67,814 expressed sequence tag (EST) sequences. The ESTs were assembled into 17,317 unigenes that included 10,319 contigs and 6,818 singletons. The S. hermonthica unigene dataset was subjected to a comparative analysis with other plant genomes or ESTs. Approximately 80% of the unigenes have homologs in other dicotyledonous plants including Arabidopsis, poplar, and grape. We found that 589 unigenes are conserved in the hemiparasitic Triphysaria species but not in other plant species. These are good candidates for genes specifically involved in plant parasitism. Furthermore, we found 1,445 putative simple sequence repeats (SSRs) in the S. hermonthica unigene dataset. We tested 64 pairs of PCR primers flanking the SSRs to develop genetic markers for the detection of polymorphisms. Most primer sets amplified polymorphicbands from individual plants collected at a single location, indicating high genetic diversity in S. hermonthica. We selected 10 primer pairs to analyze S. hermonthica harvested in the field from different host species and geographic locations. A clustering analysis suggests that genetic distances are not correlated with host specificity.
Conclusions
Our data provide the first extensive set of molecular resources for studying S. hermonthica, and include EST sequences, a comparative analysis with other plant genomes, and useful genetic markers. All the data are stored in a web-based database and freely available. These resources will be useful for genome annotation, gene discovery, functional analysis, molecular breeding, epidemiological studies, and studies of plant evolution.
Background
Striga hermonthica is an obligate root parasite belonging to the family Orobanchaceae, and is a major constraint of crop production in sub-Saharan Africa. S. hermonthica infests economically important crops such as sorghum, maize, millet, and upland rice, and the yield losses caused by this species have been estimated to cost as much as US$ 7 billion annually [1]. However, methods for controlling S. hermonthica are not well established. Despite its agricultural importance, the molecular mechanisms controlling the establishment of parasitism are poorly understood.
The S. hermonthica life cycle is unique and well adapted to its parasitic lifestyle. The seeds need to be exposed to germination stimulants exudated from the host roots, such as strigolactones and ethylene; otherwise they can remain dormant in the soil for several decades [2]. The seeds are tiny and possess limited amounts of nutrients, and this restricts their growth without a host connection. When a potential host is recognized through the sensing of strigolactones or other germination stimulants, the seeds that are close to the host roots (within 5 mm) can germinate. The germinated seedlings form haustoria, which are round shaped organs specialized in host attachment and penetration [3]. The formation of haustoria also requires host-derived signal compounds. The haustoria penetrate the host roots and finally connect with the vasculature to rob the host plant of water and nutrients. This dramatic developmental transition from an autotrophic to a heterotrophic lifestyle occurs within several days.
Intensive efforts in the scientific community, mainly in the United States during the 1960s, lead to the identification of some germination stimulants. This was followed by the development of a "suicidal germination" strategy to eradicate Striga weeds [4]. By this strategy, a germination stimulant (in this case ethylene) is mixed in the soil to trigger germination in the absence of the hosts. This approach was used successfully to eradicate Striga asiatica infestations in North Carolina. Although suicidal germination was effective for controlling S. asiatica, this approach was not applicable for African farmers due to the high cost of the strategy and the much larger scale of infestation.
Whole genome sequencing is a valuable approach to understanding an organism. The genome sequences of growing numbers of model and crop plant species have been published in recent years, providing new insights in plant biology. The development of new generation sequencing technologies has dramatically accelerated the speed of large-scale sequencing. However, the de novo sequencing of the whole genome of a non-model plant is still a challenging and laborious task [5]. Expressed sequence tags (ESTs) are a less expensive alternative for gaining information about the expressed genes of an organism [6]. In particular, the ESTs from a full-length enriched cDNA library provide the complete sequences of functional proteins [7].
This study aims to provide genome scale molecular resources for understanding the parasitic processes of the obligate parasite, S. hermonthica. We constructed a full-length enriched cDNA library from S. hermonthica and generated a large-scale EST dataset by reading the sequences of individual clones from both ends. The only other genus from the family Orobanchaceae with publically available EST data is Triphysaria [8]. Triphysaria spp. are facultative hemiparasites, which are able to complete their life cycles without hosts. The comparison of our S. hermonthica EST dataset with those of Triphysaria and other non-parasitic plantspecies enabled us to identify the potentially parasite specific genes. Furthermore, our results provide the tools to analyze genetic diversity within S. hermonthica. We found 1,445 putative simple sequence repeats (SSRs) that could be useful as markers. We amplified the genomic regions flanking some of these SSRs from S. hermonthica individuals that were collected in different fields in Africa. The results revealed high sequence divergence in the S. hermonthica genomes. All the sequences and the annotation results are freely available on the internet [9].
Results and Discussion
Genome size of S. hermonthica
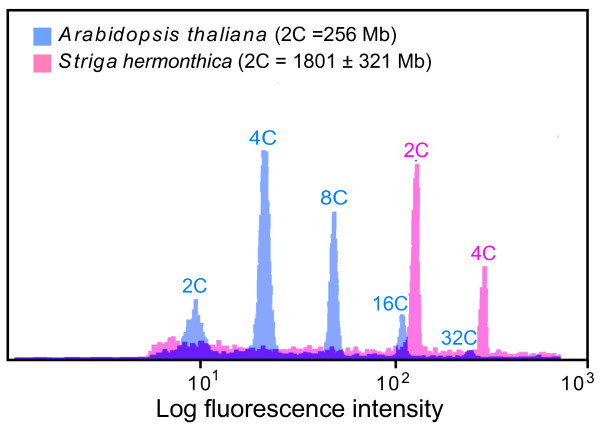
S. hermonthica is likely to be a diploid species with a chromosome number of n = 19 [10]. First, we estimated the genome size of S. hermonthica to gain information about its genome contents. Leaves of S. hermonthica plants parasitizing to rice were harvested and the DNA contents were measured with a flow cytometer. Arabidopsis thaliana, whose genome size is 128 Mbp, was used as a control. Five individual plants were used for the measurements with two or more replicates for each plant. The genome size of S. hermonthica was estimated to be 1,801 Mbp (± 321 Mbp) (Fig. 1), which is approximately 14 times that of Arabidopsis, 4 times those of rice and poplar, and 2 times that of sorghum.
Figure 1.
Genome size of S. hermonthica estimated by flow cytometry. The genome size of S. hermonthica (pink) was estimated by comparison with Arabidopsis (blue). n = 5.
Full-length enriched cDNA library construction
To construct a full-length enriched cDNA library containing highly variable sequences, total RNA was extracted from various S. hermonthica tissues at various developmental stages (Table 1). A full-length enriched normalized cDNA library was constructed using a mixture of these RNAs as starting materials. To assess the quality of the resulting library, the inserts from 90 randomly picked clones were amplified by PCR with primers specific to the library vector, and the insert sizes were estimated by agarose-gel electrophoresis (Table 2). The average insert size was approximately 1.42 kb, which is similar to the average insert size of the RIKEN Arabidopsis Full-Length (RAFL) cDNA clones (estimated at 1,445 bp) [11,12]. This average insert size was similar to that of a poplar full-length cDNA library (Populus nigra, about 1.4 kb) [13], and slightly shorter than those from soybean and wheat (approximately 1.5 kb) [12,14]. The longest insert was estimated at more than 3 kb, suggesting that the library contains relatively long cDNAs.
Table 1.
RNA samples used for the S. hermonthica full-length enriched cDNA library construction.
| Tissue | Growth stage or treatment |
|---|---|
| Seedlings | At 3 d after strigol treamtment |
| Seedlings | At 3 d after co-incubation with rice roots |
| Leaves and stems | From mature plants parasitized on rice |
| Roots (secondary haustoria) | From mature plants parasitized on rice in rhizotron |
| Flowers | From mature plants parasitized on rice |
| Axenically grown plants | Grown axenically for 1 month |
Table 2.
Distribution of insert lengths in the S. hermonthica full-length enriched cDNA library.
| Length (kb) | Clone number | Frequency (%) |
|---|---|---|
| <0.5 | 0 | 0 |
| 0.5-1.0 | 18 | 20.0 |
| 1.0-1.5 | 35 | 38.9 |
| 1.5-2.0 | 23 | 25.6 |
| 2.0-2.5 | 9 | 10.0 |
| 2.5-3.0 | 4 | 4.4 |
| ≥3.0 | 1 | 1.1 |
| Total | 90 | 100 |
To assess the proportion of the library containing full-length cDNA clones, we randomly picked 90 clones and sequenced them from both the 5' and 3' ends. These DNA sequences were analyzed against the Arabidopsis genome database using the blastx program. Of the 90 clones, 79 contained sequences similar to those of Arabidopsis genes (e_value < e-10), while the insert sequences of the other 11 clones did not show any similarity. The 5'- and 3'- sequences of the 79 clones were aligned with the homologous Arabidopsis cDNAs. The 5'-sequences of 62 clones contained ATG start codons at similar positions to those in the corresponding Arabidopsis homologs, and 59 possessed stop codons at the equivalent positions. Therefore, we estimated that approximately 75% of the clones in the S. hermonthica library encode full-length cDNAs. Among the 59 sequenced full-length clones, the average lengths of the 5'- and 3'-untranslated regions (UTRs) were 127 bp and 203 bp, respectively, and the longest 5'-and 3' -UTRs were 486 bp and 480 bp, respectively.
EST sequencing and statistical analysis
Next, we sequenced both the 5'- and 3'-ends of 37,710 clones from the S. hermonthica full-length enriched cDNA library. The sequence chromatograms were analyzed using the EST2uni package [15], which is an automated analysis tool for the clean-up, clustering, and annotation of EST sequences. Among the 75,330 raw sequence reads, we found that 67,814 were of good quality and were deposited in the DNA Databank of Japan [DDBJ: FS438984-FS506797]. The sequences are clustered into 17,137 non-redundant unigenes (10,319 contigs and 6,818 singletons) (Table 3). The average GC content among the unigene sequences is 44.5%. The lengths of the unigenes are distributed between 82 and 3,949 bp, and most of them (11,546 unigenes) have sequence lengths between 601 and 900 bp (Additional file 1), with an average of 810.3 bp. Most (84%) of the unigenes are comprised of fewer than 6 ESTs (Additional file 1), suggesting that the redundancy rate is relatively low in this normalized library.
Table 3.
Summary of the S. hermonthica EST sequence analysis
| Group | Records |
|---|---|
| Number of independent clones | 37,710 |
| Number of raw sequences | 75,330 |
| Number of high quality sequences | 67,814 |
| Number of unigenes | 17,137 |
| singletons | 6,818 |
| contigs | 10,319 |
| Average unigene length | 775.3 bp |
| Minimum unigene length | 101 bp |
| Maximum unigene length | 3,051 bp |
| Average number of ESTs per unigene | 2.9 |
| Maximum number of ESTs per contig | 106 |
| Number of superunigenes | 12,272 |
| with more than one unigene | 2,203 |
| with one unigene | 10,069 |
| Number of putative SNPs (pSNPs) | 9,299 |
| Number of putative SSRs (pSSRs) | 1,445 |
Functional annotation of the unigene sequences
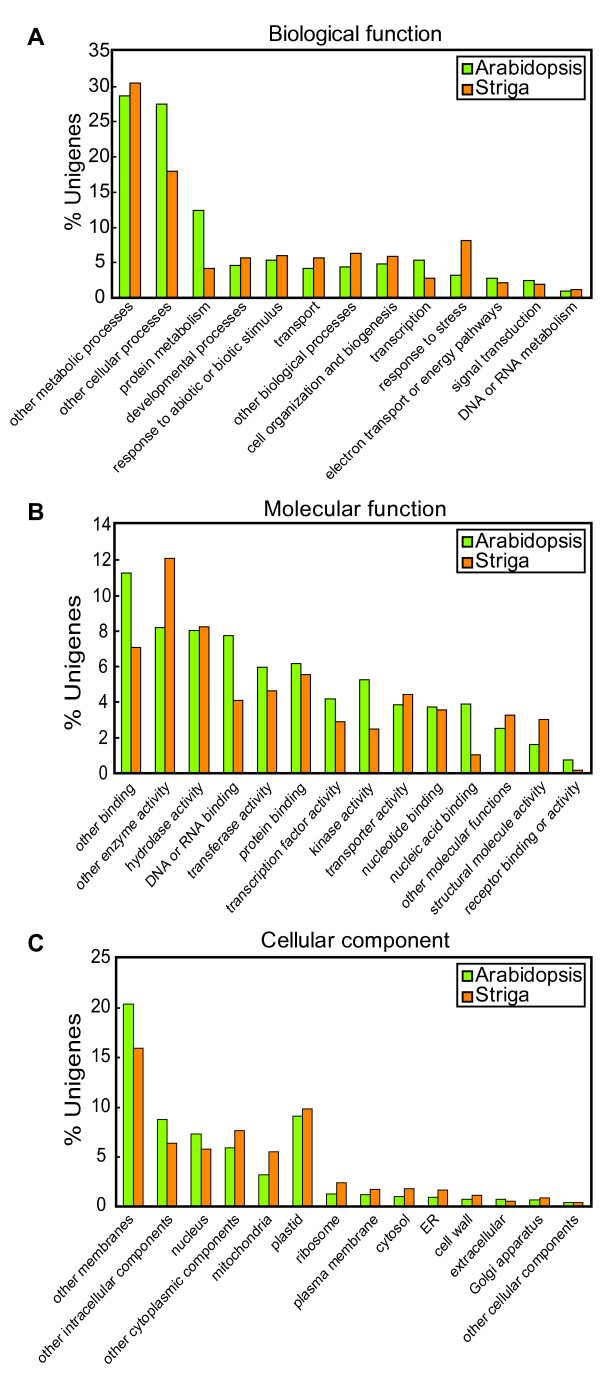
For the functional annotation of the 17,137 unigene sequences, we carried out a blastx analysis against the UniRef90 database [16,17]. About 79% of the S. hermonthica unigenes were annotated as homologs of known proteins. For further functional annotations of the structural domains, the Pfam database [18] was searched using the HMMER program (ver. 2.3.2, [19,20]), and 31% (5367) of the unigenes contained Pfam hits. Then the S. hermonthica unigenes were classified into Gene ontology (GO) groups based on their similarities with the corresponding Arabidopsis genes (Fig. 2). In the classification of genes according to their cellular components, we found that 16% of the unigenes encode putative membrane proteins and 10% encode putative plastid proteins. In the classification of molecular functions, 12% were assigned to catalytic activity. These percentages are similar to those in Arabidopsis [21], indicating that there was no functional bias among the predicted proteins encoded in the S. hermonthica library.
Figure 2.
Gene ontology analysis of S. hermonthica unigene-encoding products. The S. hermonthica unigenes were classified according to their predicted biological functions (A), molecular functions (B), and cellular components (C). The numbers in each category were compared with those in A. thaliana.
Comparative analysis with other plant genes
The S. hermonthica unigenes were compared with genes in other plant genomes, including A. thaliana, poplar (Populus trichocarpa), grape (Vitis vinifera), soybean (Glycine max), rice (Oryza sativa), sorghum (Sorghum bicolor), a moss (Physcomitrella patens), and an algae (Chlamydomonas reindardtii) [22-26]. Seventy-seven to seventy-nine percent of the S. hermonthica unigenes showed similarities with genes from other dicotyledonous plants (Arabidopsis, grape, soybean, and poplar), as detected by blastx (e_value < e-10). Approximately 75% of the unigenes have homologs in monocotyledonous plants (rice and sorghum), and approximately 65% and 38% showed blastx hits in the P. patens and C. reindardtii databases, respectively. These lower percentages of blast hits are consistent with the greater evolutionary distances from those organisms.
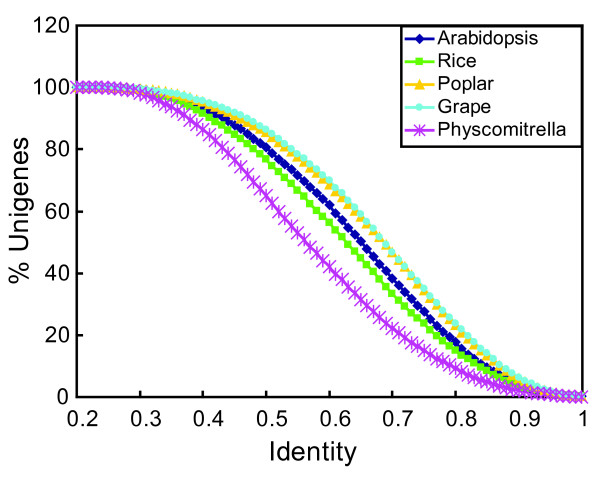
We plotted the percentages of S. hermonthica unigenes against levels of amino acid sequence identity with homologs in the other plant genomes (Fig. 3). Larger percentages of S. hermonthica unigenes showed higher levels of identity with poplar and grape sequences than with sequences from the other plant species. The identity scores corresponding to half the population of S. hermonthica unigenes were 0.68 for grape and poplar, 0.65 for Arabidopsis, 0.62 for rice, and 0.56 for P. patens. These numbers roughly reflect the evolutionary distances between S. hermonthica and these species.
Figure 3.
Cumulative count curves of identity between S. hermonthica unigenes and those from other plant species. All the sequenced S. hermonthica unigenes were used in blastx or tblastx searches against the peptide databases of the indicated plant species. The curves represent the percentages of S. hermonthica unigenes that showed higher levels of identity than the values on the _x_-axis.
Large scale EST sequence datasets have previously been reported for Triphysaria versicolor [8] and Triphysaria pusilla [27], which are hemiparasitic plants belonging to the Orobanchaceae. The assembled EST sequences are available at the plantGDB web site [28]. Althoughthe genus Triphysaria is closely related taxonomically to S. hermonthica, only 74% of the S. hermonthica unigenes showed similarity to Triphysaria sequences (including both T. pusilla and T. versicolor), when analyzed with the tblastx program (Table 4). This is significantly lower than percentages of similarity found with the other dicotyledonous plants, but this is likely due to the lack of saturation of the Triphysaria EST datasets.
Table 4.
Summary of blast search results using S. hermonthica unigenes.
| Species | DB version | Number of hits | % Unigenes |
|---|---|---|---|
| Populus trichocarpa | JGI ver1.1 | 13,573 | 79.2 |
| Glycine max | JGI ver1.1 | 12,716 | 79.0 |
| Vitis vinifera | ver1 | 13,345 | 77.9 |
| Arabidopsis thaliana | TAIR8 | 13,255 | 77.3 |
| Oryza sativa | TIGR ver6 | 12,841 | 74.9 |
| Sorghum bicolor | JGI ver1.1 | 12,803 | 74.7 |
| Triphysaria pusilla | EST | 12,716 | 74.2 |
| Physcomitrella patens | JGI ver1.1 | 11,140 | 65.0 |
| Chlamydomonas reinhardtii | JGI ver1.1 | 6,477 | 37.8 |
| No hit | 2,389 | 13.9 |
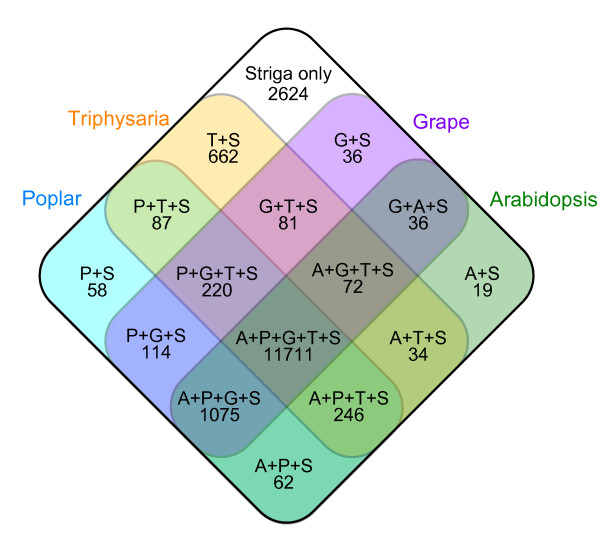
The conservation of the genes between S. hermonthica and Arabidopsis, grape, poplar, or Triphysaria spp. is shown in a Venn diagram (Fig. 4). Among the 17,137 unigenes, 11,711 (68%) are conserved among all five groups. Only 19, 36, and 58 of the S. hermonthica unigenes are conserved specifically in Arabidopsis, grape, and poplar, respectively. Interestingly, we found that 662 (3.9%) of the S. hermonthica unigenes are conserved in Triphysaria spp. but not in Arabidopsis, grape, or poplar.
Figure 4.
Homologous gene groups between S. hermonthica and four other plant species. The numbers of S. hermonthica unigenes that have homologues in the indicated plant species are represented by a Venn diagram. A: A. thaliana, G: V. vinifera, P: P. trichocarpa, T: _T. pusilla_or T. versicolor, and S: S. hermonthica.
Of these 662 sequences, 73 show similarities to sequences in other databases such as rice, sorghum, soybean, Physcomitrella, UniRef90 or nr (the non-redundant peptide database from NCBI). We found no other homologs for the remaining 589 unigenes (Additional file 2). Since T. pusilla and T. versicolor are hemiparasitic plants, these 589 might include genes specific to parasitic plants. The ongoing project to sequence the genome of Mimulus spp. may help to narrow down the number of candidate genes that are involved in parasitism, because Mimulus spp. are non-parasitic members of the family Scrophulariaceae, which is taxonomically close to Orobanchaceae. The 2,389 unigenes (14%) that did not show significant hits with any known peptide sequences in the tested databases (including nr) are also listed in Additional file 2. These unigenes may include sequences that are specific to Striga.
Genetic diversity of the S. hermonthica sequences
S. hermonthica is an obligate outcrossing plant with high levels of morphological and genetic variation [29]. The EST2uni program detected 9,299 putative single nucleotide polymorphisms (SNPs) among the S. hermonthica unigenes. To exclude the misidentification of sequencing errors as SNPs, only polymorphisms confirmed by at least 2 independent sequences were counted, although there is still the possibility that those polymorphisms occurred during cDNA synthesis. The average frequency of SNPs in the unigene sequences is 0.67%, or approximately 1 SNP per 1.5 kbp. Although these SNPs will need to be confirmed, these data will be useful for developing EST-SNP markers for S. hermonthica [30].
We found 1,445 di-, tri- or tetra-nucleotide microsatellites (or SSRs) among the S. hermonthica unigenes. The most frequent of these are the tri-nucleotide repeats (Additional file 3), which is in agreement with previous studies of other plant species [31-33]. The most frequent individual microsatellite repeat is AG (including TC, GA, and TC) (283, 19.6%) and the second most frequent is AC (including TG, CA, and GT) (218, 15.1%). The most frequent tri-nucleotide repeat is ATC (including TCA and CAT) (157, 11.0%) (Additional file 4).
The EST-SSR sequences are good candidates for genetic markers, which can be used for molecular diagnosis, for biotyping weeds, and for investigating the genetic diversity and population structures of S. hermonthica. To investigate whether the SSRs that we identified can be used as such markers, we designed primers using sequences flanking the putative SSRs and looked for polymorphisms by PCR. First, we pooled DNA samples extracted from the leaves of several plants in the same field and used the DNA pools as PCR templates. Of the 64 primer sets tested, 44 successfully amplified DNA bands. However, 26 primer sets (59%) produced smears or multiple bands that were not countable and only 18 primer pairs (41%) amplified clear separate bands (Additional file 5). The smeared bands may indicate heterozygosity and genetic diversity among the individual plants harvested from the same field. Therefore, we tested the individual plants for polymorphisms. Several markers that showed smear patterns from the pooled DNA templates actually amplified clear polymorphic bands from individual plants in the same population (Additional file 6). These data verify that S. hermonthica is a highly adaptable weed that has maintained a high degree of genetic variation and plasticity, to survive in various ecosystems [34].
Genetic distances among S. hermonthica populations with different hosts
Although individual S. hermonthica plants possess highly diversified genomes, 18 of the primer sets we tested showed countable band patterns when using pooled DNA templates. Using those primer sets, we investigated the relationships between different S. hermonthica populations from 6 fields growing sorghum, maize, or pearl millet in various locations in Sudan or Kenya [35]. Of the 18 primer sets, 10 showed clear polymorphisms for different S. hermonthica populations (Table 5, Additional file 5). The analysis of PCR products was carried out using MultiNa® (Shimadzu, Japan), a microchip electrophoresis system that permits the separation of small fragments and that can detect 5 bp differences. The average polymorphism information content (PIC) was 0.463, which confirms that the SSR markers used in this study were highly informative The lowest PIC value was 0.305 for SSR57, and the highest was 0.545 for SSR26 (Table 5). The analyzed loci included 3 di-, 3 tri-, and 4 tetra-nucleotide repeats. A total of 27 alleles were detected, with an average number of alleles per locus of 2.7. The genetic diversity among the six populations was revealed by the gene diversity values, which ranged from 0.375 to 0.625, with an average of 0.549. These results suggest a high level of diversity among the surveyed populations, as was expected for this obligate outcrossing plant [36-38].
Table 5.
Genetic diversity among S. hermonthica populations collected from various locations and host plants.
| SSR ID | Primer name | Repeat unit | No of repeats | No of alleles | Gene diversity | PIC |
|---|---|---|---|---|---|---|
| ShSSR_ShContig8678_1 | SSR17 | AC | 18 | 3 | 0.611 | 0.535 |
| ShSSR_ShContig6892_1 | SSR26 | AG | 15 | 3 | 0.625 | 0.545 |
| ShSSR_ShSHAA-aai51d05.b1_c_s_1 | SSR33 | AG | 13 | 3 | 0.611 | 0.535 |
| ShSSR_ShContig9253_1 | SSR43 | CCG | 10 | 2 | 0.486 | 0.368 |
| ShSSR_ShContig5481_1 | SSR50 | AAG | 9 | 3 | 0.569 | 0.477 |
| ShSSR_ShContig5198_1 | SSR53 | ACC | 8 | 2 | 0.486 | 0.368 |
| ShSSR_ShContig5533_1 | SSR57 | AACT | 6 | 2 | 0.375 | 0.305 |
| ShSSR_ShContig10128_1 | SSR58 | AAAC | 7 | 3 | 0.569 | 0.505 |
| ShSSR_ShSHAA-aab89e01.b1_c_s_1 | SSR59 | AAAC | 6 | 3 | 0.542 | 0.460 |
| ShSSR_ShContig9110_1 | SSR63 | AAAG | 5 | 3 | 0.611 | 0.535 |
| Average | 2.700 | 0.549 | 0.463 |
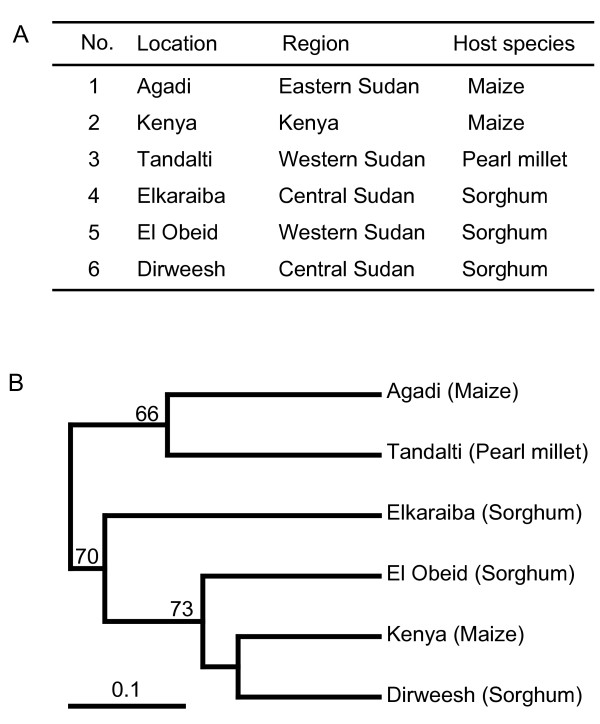
We also looked for correlations between host species and S. hermonthica biotypes, using the Unweighted Pair Group Method with Arithmetic mean (UPGMA) clustering analysis. The populations from El Obeid (host: sorghum), Dirweesh (host: sorghum), and Kenya (host: maize) clustered in one group, while the population from Elkaraiba (host: sorghum) was in a distant branch of the same group. Those from Tandalti (host: pearl millet) and Agadi (host: maize) formed another cluster (Fig. 5). Thus, we did not detect any correlations between genetic distance and host specificity in this study. This result is consistent with previous epidemiological reports [35,38-40]. In summary, our results suggest that the SSRs found in our study could be useful tools for further investigations of genetic diversity in S. hermonthica.
Figure 5.
Clustering analysis of S. hermonthica populations using SSR polymorphisms. A. S. hermonthica populations used in this study. B. A UPGMA dendrogram constructed using polymorphisms at 10 SSR loci with a total of 27 alleles. Bootstrap values are indicated at supporting nodes when the values are greater than 50.
Web-based database
The results of the sequencing and analysis of the S. hermonthica ESTs are freely available online from our web-based database [9]. The web interface was based on the original EST2uni web site [15]. The database contains features for complex query searches and a blast search. A page for each unigene consists of its sequence, contig images, results of blast similarity searches, lists of detected SSRs and SNPs, and GO categorizations. In addition, the homologs of each unigene are linked to outside databases such as The Arabidopsis Information Resource (TAIR) [41]. This web-based database will be a powerful tool for the detailed analysis of S. hermonthica genes.
Conclusions
This paper provides large scale EST information about S. hermonthica, which can be used in studies of parasitic plants, plant-plant interactions, weed management, and plant evolution. Comparative analyses between S. hermonthica and other plant genomes should allow us to identify genes responsible for plant parasitism. These genes are of particular interest as potential targets for future pest management strategies against noxious parasitic weeds. Our analysis also highlights the intra-species genetic diversity of S. hermonthica. A more detailed analysis might contribute to future breeding programs to develop resistant crops, since genetic variation in the weed population could be the main factor allowing the quick breakdown of resistance. In summary, our study provides powerful analytical tools for the molecular analysis of the parasitic weed S. hermonthica. Our data will alsocontribute to the annotation of genes identified by the on-going genome-scale sequencing of the parasitic genera from Orobanchaceae.
Methods
Plant materials and growth conditions
S. hermonthica seeds collected from a sorghum field in 1994 in Kenya were provided by Dr. A. G. Babiker (Univ. of Sudan, Khartoum, Sudan). Rice seeds (Oryza sativa L. subspecies japonica, cultivar Koshihikari) were originally obtained from the National Institute of Agricultural Sciences (NIAS, Tsukuba, Japan). S. hermonthica plants parasitizing rice were grown in rhizotrons as described previously [42] or in soil (1:1 mixture of vermiculite: clay). For the axenic culture of S. hermonthica, seeds were sterilized with 20% bleach solution (approx. 6% NaOCl) for 5 min and washed thoroughly with sterile water. The sterile seeds were preconditioned on MS medium with 1% sucrose and 0.5% phytagel (Sigma) at 26°C for 7 to 10 days in the dark and germination was stimulated by the exogenous application of 5 μl 1 μM Strigol per plate. Sterile S. hermonthica plants were grown on the same medium at 26°C with a 16-h photoperiod, and the medium was renewed every 3 weeks.
Determination of nuclear DNA content
The nuclear DNA content was analyzed with a flow cytometer (Partec PA, Tokyo, Japan). Soil-grown S. hermonthica (host: rice) leaves were chopped with a razor blade into small pieces and analyzed according to the previously published method [43]. Leaves of Arabidopsis (ecotype Col -0) were used as the control.
RNA extraction
The S. hermonthica tissues and developmental stages used for RNA extraction are listed in Table 1. S. hermonthica RNAs were extracted using a modified cetyl trimethylammonium bromide (CTAB) method. Briefly, plant tissues were ground under liquid nitrogen and suspended in 5 × volumes of CTAB solution (2% CTAB, 2% polybinylpyrrolidone (PVP), 25 mM ethylenediaminetetraacetic acid(EDTA), 2 M NaCl, 1% beta-mercaptoethanol, 100 mM Tris-HCl (pH 8.0)) and phenol:chloroform (5:1, pH 4.7, Sigma). The mixtures were shaken at 55°C for 5 min. After 10 min centrifugation, the aqueous phase was extracted with an equal volume of phenol:chloroform, and subsequently with chloroform. The RNAs were precipitated by adding 0.25 volumes of 10 M LiCl. The RNA pellet was washed with 70% ethanol and then dissolved in nuclease-free water. Samples were subsequently purified using the PureLink RNA mini kit (Invitrogen) according to the manufacture's instructions. To obtain mRNA for library construction, total RNAs from each tissue and developmental stage were mixed and purified using an mRNA purification kit (GE) according to the manufacture's instructions. The quality and quantity of the total RNA and the mRNA were assessed by measurements of OD230, OD260, and OD280, followed by visual checking by electrophoresis.
Library construction and EST sequencing
The construction of the normalized, full-length enriched library was carried out in Evrogen (Russia). The cDNA normalization was conducted using a Duplex-specific nuclease (DSN)-based method, and full-length cDNAs were enriched using the SMART™ technology (Clontech). Each cDNA was inserted into the pAL17.3 vector. Sequencing of randomly picked clones was performed in the Genome Center at Washington University using the ABI3730 capillary sequencer.
Computational analysis
The EST sequences were automatically trimmed, clustered and annotated using the EST2uni analysis pipeline [15]. Sequence assembly was performed using the CAP3 program with the default parameter settings [44]. Blast searches were performed with NCBI blast program against the databases shown in Table 4. The S. hermonthica online database was constructed based on the EST2uni web program with slight modifications.
SSR markers and genetic diversity analysis
Genomic DNA was extracted from about 10 g of S. hermonthica seeds using the modified CTAB method described previously [35]. Primers flanking the microsatellites were designed using the PRIMER 3 program [45]. The PCRs were performed in 10 μl volumes with one initial denaturation step of 1 min at 95°C, followed by 40 cycles of 15 sec at 94°C, 30 sec at 60°C and 30 sec at 72°C, anda final extension step of 5 min at 72°C. The PCR products were analyzed either by 4% agarose gel electrophoresis (Additional file 6) or using the MCE-202 MultiNa Microchip Electrophoresis System for DNA/RNA analysis (Shimadzu, Japan) using the DNA-500 kit (Table 5 and Fig. 5). The data were analyzedusing the PowerMarker program version 3.25 [46], and the genetic diversity was estimated based on allelic numbersand the gene diversity value:
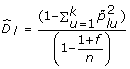
where n is the number of populations sampled, p lu is the frequency of _u_th allele at the _l_th locus, and f is the inbreeding coefficient (association between alleles) at the l_th locus. The Polymorphism Information Content (PIC) was estimated as 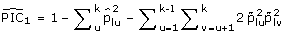 , where the p_lv is the frequency of the _v_th allele at the _l_th locus. The phylogenetic UPGMA tree was generated based on a matrix of the frequencies and distances using the LogSharedAllele algorithm with the PowerMarker v.3.25 program. Bootstrap analysis was performed using the software package WINBOOT [47].
, where the p_lv is the frequency of the _v_th allele at the _l_th locus. The phylogenetic UPGMA tree was generated based on a matrix of the frequencies and distances using the LogSharedAllele algorithm with the PowerMarker v.3.25 program. Bootstrap analysis was performed using the software package WINBOOT [47].
Authors' contributions
SY carried out the data collection and bioinformatic analyses, and drafted the manuscript. JKI performed the SSR marker analyses. NMK and AMA collected S. hermonthica seeds and extracted genomic DNAs. SN participated in the design and coordination of the study. KS conceived of the study, contributed to designing the experiments, and drafted the manuscript. All authors read and approved the final manuscript.
Supplementary Material
Additional file 1
Distribution of unigene lengths and EST numbers per unigene. (A) Distribution of unigene lengths in the entire S. hermonthica unigene dataset. (B) Distribution of EST numbers per unigene.
Additional file 2
Lists of S. hermonthica unigenes that are potentially specific to parasitic plants. Sheet 1- The list of S. hermonthica unigenes that have homologs in T. pusilla or T. versicolor but not in other species databases. Sheet 2- The list of S. hermonthica unigenes that do not have homologs in other known sequences.
Additional file 3
Distribution of SSR patterns detected in S. hermonthica ESTs.
Additional file 4
Distribution of SSR motifs detected in S. hermonthica ESTs.
Additional file 5
SSR information. Sheet1- The list of SSRs analyzed in this study, with SSR ID, primer sequences, and PCR results. The yellow colored linesindicate the markers used in this study.
Additional file 6
Examples of PCR results from the amplification of SSR-containing regions in S. hermonthica. (A) Agarose gel images of PCR results using the indicated primer sets and pooled genomic DNAs from the populations listed in Fig. 5. The population numbers correspond to the numbers in Fig. 5A. (B) An agarose gel image showing PCR results using the SSR8 primer set and genomic DNAs extracted from individual plantsfrom the population in Kenya.
Contributor Information
Satoko Yoshida, Email: satokoy@psc.riken.jp.
Juliane K Ishida, Email: jkishida@psc.riken.jp.
Nasrein M Kamal, Email: renokamal@yahoo.com.
Abdelbagi M Ali, Email: abdmali@yahoo.com.
Shigetou Namba, Email: anamba@mail.ecc.u-tokyo.ac.jp.
Ken Shirasu, Email: ken.shirasu@psc.riken.jp.
Acknowledgements
We thank Dr K. Mochida for advice on bioinformatics, K. Akiyama and T. Sakurai for web-server maintenance, and Dr A. G. Babiker for providing the S. hermonthica seeds. This work was funded by grants from the Gatsby Charitable Foundation, the RIKEN president fund, and KAKENHI (19780040 and 21780044 to SY and 19678001 to KS). JKI is supported by the MEXT scholarship program.
References
- Parker C. Observations on the current status of Orobanche and Strigaproblems worldwide. Pest management science. 2009;65(5):453–459. doi: 10.1002/ps.1713. [DOI] [PubMed] [Google Scholar]
- Bouwmeester HJ, Roux C, Lopez-Raez JA, Becard G. Rhizosphere communication of plants, parasitic plants and AM fungi. Trends in plant science. 2007;12(5):224–230. doi: 10.1016/j.tplants.2007.03.009. [DOI] [PubMed] [Google Scholar]
- Yoder JI. Host-plant recognition by parasitic Scrophulariaceae. Current Opinion in Plant Biology. 2001;4(4):359–365. doi: 10.1016/S1369-5266(00)00185-0. [DOI] [PubMed] [Google Scholar]
- Rispail N, Dita MA, Gonzalez-Verdejo C, Perez-de-Luque A, Castillejo MA, Prats E, Roman B, Jorrin J, Rubiales D. Plant resistance to parasitic plants: molecular approaches to an old foe. The New phytologist. 2007;173(4):703–712. doi: 10.1111/j.1469-8137.2007.01980.x. [DOI] [PubMed] [Google Scholar]
- Varshney RK, Nayak SN, May GD, Jackson SA. Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends in Biotechnology. 2009;27(9):522–530. doi: 10.1016/j.tibtech.2009.05.006. [DOI] [PubMed] [Google Scholar]
- Rudd S. Expressed sequence tags: alternative or complement to whole genome sequences? Trends in plant science. 2003;8(7):321–329. doi: 10.1016/S1360-1385(03)00131-6. [DOI] [PubMed] [Google Scholar]
- Sakurai T, Plata G, Rodriguez-Zapata F, Seki M, Salcedo A, Toyoda A, Ishiwata A, Tohme J, Sakaki Y, Shinozaki K, Ishitani M. Sequencing analysis of 20,000 full-length cDNA clones from cassava reveals lineage specific expansions in gene families related to stress response. BMC Plant Biol. 2007;7:66. doi: 10.1186/1471-2229-7-66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torres MJ, Tomilov AA, Tomilova N, Reagan RL, Yoder JI. Pscroph, a parasitic plant EST database enriched for parasite associated transcripts. BMC Plant Biol. 2005;5:24. doi: 10.1186/1471-2229-5-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Striga hermonthica EST database. http://striga.psc.riken.jp
- Aigbokhan EI, Berner DK, Musselman LJ. Reproductive Ability of Hybrids of Striga aspera and Striga hermonthica. Phytopathology. 1998;88(6):563–567. doi: 10.1094/PHYTO.1998.88.6.563. [DOI] [PubMed] [Google Scholar]
- Seki M, Narusaka M, Kamiya A, Ishida J, Satou M, Sakurai T, Nakajima M, Enju A, Akiyama K, Oono Y, Muramatsu M, Hayashizaki Y, Kawai J, Carninci P, Itoh M, Ishii Y, Arakawa T, Shibata K, Shinagawa A, Shinozaki K. Functional annotation of a full-length Arabidopsis cDNA collection. Science. 2002;296(5565):141–145. doi: 10.1126/science.1071006. [DOI] [PubMed] [Google Scholar]
- Umezawa T, Sakurai T, Totoki Y, Toyoda A, Seki M, Ishiwata A, Akiyama K, Kurotani A, Yoshida T, Mochida K, Kasuga M, Todaka D, Maruyama K, Nakashima K, Enju A, Mizukado S, Ahmed S, Yoshiwara K, Harada K, Tsubokura Y, Hayashi M, Sato S, Anai T, Ishimoto M, Funatsuki H, Teraishi M, Osaki M, Shinano T, Akashi R, Sakaki Y. Sequencing and analysis of approximately 40,000 soybean cDNA clones from a full-length-enriched cDNA library. DNA Res. 2008;15(6):333–346. doi: 10.1093/dnares/dsn024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nanjo T, Sakurai T, Totoki Y, Toyoda A, Nishiguchi M, Kado T, Igasaki T, Futamura N, Seki M, Sakaki Y, Shinozaki K, Shinohara K. Functional annotation of 19,841 Populus nigra full-length enriched cDNA clones. BMC genomics. 2007;8:448. doi: 10.1186/1471-2164-8-448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogihara Y, Mochida K, Kawaura K, Murai K, Seki M, Kamiya A, Shinozaki K, Carninci P, Hayashizaki Y, Shin IT, Kohara Y, Yamazaki Y. Construction of a full-length cDNA library from young spikelets of hexaploid wheat and its characterization by large-scale sequencing of expressed sequence tags. Genes & genetic systems. 2004;79(4):227–232. doi: 10.1266/ggs.79.227. [DOI] [PubMed] [Google Scholar]
- Forment J, Gilabert F, Robles A, Conejero V, Nuez F, Blanca JM. EST2uni: an open, parallel tool for automated EST analysis and database creation, with a data mining web interface and microarray expression data integration. BMC bioinformatics. 2008;9:5. doi: 10.1186/1471-2105-9-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzek BE, Huang H, McGarvey P, Mazumder R, Wu CH. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinfomatics. 2007;23(10):1282–1288. doi: 10.1093/bioinformatics/btm098. [DOI] [PubMed] [Google Scholar]
- Uniref. http://www.uniprot.org/help/uniref
- Pfam. http://pfam.sanger.ac.uk/
- hmmer. http://hmmer.janelia.org/
- Bateman A, Birney E, Durbin R, Eddy SR, Finn RD, Sonnhammer EL. Pfam 3.1: 1313 multiple alignments and profile HMMs match the majority of proteins. Nucleic acidsresearch. 1999;27(1):260–262. doi: 10.1093/nar/27.1.260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berardini TZ, Mundodi S, Reiser L, Huala E, Garcia-Hernandez M, Zhang P, Mueller LA, Yoon J, Doyle A, Lander G, Moseyko N, Yoo D, Xu I, Zoeckler B, Montoya M, Miller N, Weems D, Rhee SY. Functional annotation of the Arabidopsis genome using controlled vocabularies. Plant physiology. 2004;135(2):745–755. doi: 10.1104/pp.104.040071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arabidopsis-Genome-Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408(6814):796–815. doi: 10.1038/35048692. [DOI] [PubMed] [Google Scholar]
- Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyere C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Del Fabbro C, Alaux M, Di Gaspero G, Dumas V. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature. 2007;449(7161):463–467. doi: 10.1038/nature06148. [DOI] [PubMed] [Google Scholar]
- Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J, Spannagl M, Tang H, Wang X, Wicker T, Bharti AK, Chapman J, Feltus FA, Gowik U, Grigoriev IV, Lyons E, Maher CA, Martis M, Narechania A, Otillar RP, Penning BW, Salamov AA, Wang Y, Zhang L, Carpita NC. The Sorghum bicolor genome and the diversification of grasses. Nature. 2009;457(7229):551–556. doi: 10.1038/nature07723. [DOI] [PubMed] [Google Scholar]
- Rensing SA, Lang D, Zimmer AD, Terry A, Salamov A, Shapiro H, Nishiyama T, Perroud PF, Lindquist EA, Kamisugi Y, Tanahashi T, Sakakibara K, Fujita T, Oishi K, Shin IT, Kuroki Y, Toyoda A, Suzuki Y, Hashimoto S, Yamaguchi K, Sugano S, Kohara Y, Fujiyama A, Anterola A, Aoki S, Ashton N, Barbazuk WB, Barker E, Bennetzen JL, Blankenship R. The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science. 2008;319(5859):64–69. doi: 10.1126/science.1150646. [DOI] [PubMed] [Google Scholar]
- Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray) Science. 2006;313(5793):1596–1604. doi: 10.1126/science.1128691. [DOI] [PubMed] [Google Scholar]
- Triphysaria EST database. http://www.plantsciences.ucdavis.edu/yoder/lab/Sequence_index.html
- PlantGDB. http://www.plantGDB.org
- Mohamed KI, Musselman LJ, Riches CR. The genus Striga (Scrophulariaceae) in Africa. Ann Mo Bot Gard. 2001;88(1):60–103. doi: 10.2307/2666132. [DOI] [Google Scholar]
- Deleu W, Esteras C, Roig C, Gonzalez-To M, Fernandez-Silva I, Gonzalez-Ibeas D, Blanca J, Aranda M, Arus P, Nuez F, Monforte A, Pico M, Garcia-Mas J. A set of EST-SNPs for map saturation and cultivar identification in melon. BMC Plant Biology. 2009;9(1):90. doi: 10.1186/1471-2229-9-90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kantety RV, La Rota M, Matthews DE, Sorrells ME. Data mining for simple sequence repeats in expressed sequence tags from barley, maize, rice, sorghum and wheat. Plant Molecular Biology. 2002;48(5-6):501–510. doi: 10.1023/A:1014875206165. [DOI] [PubMed] [Google Scholar]
- Liang X, Chen X, Hong Y, Liu H, Zhou G, Li S, Guo B. Utility of EST-derived SSR in cultivated peanut (Arachis hypogaea L.) and Arachis wild species. BMC Plant Biol. 2009;9(1):35. doi: 10.1186/1471-2229-9-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moccia M, Oger-Desfeux C, Marais G, Widmer A. A White Campion (Silene latifolia) floral expressed sequence tag (EST) library: annotation, EST-SSR characterization, transferability, and utility for comparative mapping. BMC genomics. 2009;10(1):243. doi: 10.1186/1471-2164-10-243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao Q, Lu BR, Xia H, Rong J, Sala F, Spada A, Grassi F. Genetic diversity and origin of weedy rice (Oryza sativa f. spontanea) populations found in North-eastern China revealed by simple sequence repeat (SSR) markers. Annals of botany. 2006;98(6):1241–1252. doi: 10.1093/aob/mcl210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abdelbagi MA, Yasir SA, Ahmed AE, Dawoud AD, Yabuta-Miyamoto S, Sugimoto Y. Molecular diversity of Striga hermonthica collected from different locations and host plant species. Sudan Journal of Agricultural Research. 2007;10:121–126. [Google Scholar]
- Mohamed KI, Bolin JF, Musselman LJ, Peterson AT. In: Integrating new technologies for Striga control: Towards Ending the Witch-Hunt. Ejeta G, Gressel J, editor. World Scientific Publishing Company; 2007. Genetic Diversity of Striga and Implications for 71 Control and Modeling Future Distributions; pp. 71–84. full_text. [Google Scholar]
- Koyama ML. In: Application of molecular markers in plant breedingbreeding Training manual for a seminar held at IITA, Ibadan, Nigeria, from 16-17 August 1999. Haussmann BIG, Geiger HH, Hess DE, Hash CT, Bramel-Cox P, editor. Patancheru 502 324, Andhra Pradesh, India: International Crops Research Institute for the Semi-Arid Tropics (ICRISAT); 2000. Molecular markers for the study of pathogen variability: implications for breeding resistance to Striga hermonthica; pp. 133–152. [Google Scholar]
- Olivier A, Glaszmann JC, Lanaud C, Leroux GD. Population structure, genetic diversity and host specificity of the parasitic weed Striga hermonthica (Scrophulariaceae) in Sahel. Plant Systematics and Evolution. 1998;209(1-2):33–45. doi: 10.1007/BF00991522. [DOI] [Google Scholar]
- Bharathalakshmi, Werth CR, Musselman LJ. A Study of Genetic Diversity among Host-Specific Populations of the Witchweed Striga-Hermonthica (Scrophulariaceae) in Africa. Plant Systematics and Evolution. 1990;172(1-4):1–12. doi: 10.1007/BF00937794. [DOI] [Google Scholar]
- Gethi JG, Smith ME, Mitchell SE, Kresovich S. Genetic diversity of Striga hermonthica and Striga asiatica populations in Kenya. Weed Research. 2005;45(1):64–73. doi: 10.1111/j.1365-3180.2004.00432.x. [DOI] [Google Scholar]
- The Arabidopsis Information Resource (TAIR) http://www.arabidopsis.org/
- Yoshida S, Shirasu K. Multiple layers of incompatibility to the parasitic witchweed, Striga hermonthica. The New phytologist. 2009;183(1):180–189. doi: 10.1111/j.1469-8137.2009.02840.x. [DOI] [PubMed] [Google Scholar]
- Yoshizumi T, Tsumoto Y, Takiguchi T, Nagata N, Yamamoto YY, Kawashima M, Ichikawa T, Nakazawa M, Yamamoto N, Matsui M. Increased level of polyploidy1, a conserved repressor of CYCLINA2 transcription, controls endoreduplication in Arabidopsis. The Plant cell. 2006;18(10):2452–2468. doi: 10.1105/tpc.106.043869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang X, Madan A. CAP3: A DNA Sequence Assembly Program. Genome research. 1999;9(9):868–877. doi: 10.1101/gr.9.9.868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Primer3. http://frodo.wi.mit.edu/primer3/
- Liu K, Muse SV. PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics (Oxford, England) 2005;21(9):2128–2129. doi: 10.1093/bioinformatics/bti282. [DOI] [PubMed] [Google Scholar]
- Yap IV, Nelson R. WINBOOT: A program for performing bootstrap analysis of binary data to determine the confidence limits of UPGMA-based dendrograms. IRRI Discussion Paper Series. 1996;14 [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1
Distribution of unigene lengths and EST numbers per unigene. (A) Distribution of unigene lengths in the entire S. hermonthica unigene dataset. (B) Distribution of EST numbers per unigene.
Additional file 2
Lists of S. hermonthica unigenes that are potentially specific to parasitic plants. Sheet 1- The list of S. hermonthica unigenes that have homologs in T. pusilla or T. versicolor but not in other species databases. Sheet 2- The list of S. hermonthica unigenes that do not have homologs in other known sequences.
Additional file 3
Distribution of SSR patterns detected in S. hermonthica ESTs.
Additional file 4
Distribution of SSR motifs detected in S. hermonthica ESTs.
Additional file 5
SSR information. Sheet1- The list of SSRs analyzed in this study, with SSR ID, primer sequences, and PCR results. The yellow colored linesindicate the markers used in this study.
Additional file 6
Examples of PCR results from the amplification of SSR-containing regions in S. hermonthica. (A) Agarose gel images of PCR results using the indicated primer sets and pooled genomic DNAs from the populations listed in Fig. 5. The population numbers correspond to the numbers in Fig. 5A. (B) An agarose gel image showing PCR results using the SSR8 primer set and genomic DNAs extracted from individual plantsfrom the population in Kenya.