Spatial organization of chromatin domains and compartments in single chromosomes (original) (raw)
. Author manuscript; available in PMC: 2017 Feb 5.
Published in final edited form as: Science. 2016 Jul 21;353(6299):598–602. doi: 10.1126/science.aaf8084
Abstract
The spatial organization of chromatin critically impacts genome function. Recent chromosome-conformation-capture studies have revealed topologically-associating domains (TADs) as a conserved feature of chromatin organization, but how TADs are spatially organized in individual chromosomes remains unknown. Here we developed an imaging method for mapping the spatial positions of numerous genomic regions along individual chromosomes and traced the positions of TADs in human interphase autosomes and X chromosomes. We observed that chromosome folding deviates from the ideal fractal-globule model at large length scales and that TADs are largely organized into two compartments spatially arranged in a polarized fashion in individual chromosomes. Active and inactive X chromosomes adopt different folding and compartmentalization configurations. These results suggest that the spatial organization of chromatin domains can change in response to regulation.
The spatial organization of chromatin, such as chromatin domains, chromatin loops, associations of chromatin with nuclear structures, and chromosome territories, plays an important role in essential genome functions (1–6). However, many gaps remain in our understanding of the three-dimensional (3D) folding of individual chromosomes in the nucleus. Recently, chromosome-conformation-capture methods such as Hi-C (4, 7) have revealed a wealth of structural insights for interphase chromosomes. For example, chromatin is organized into topologically-associating domains (TADs) or contact domains that are hundreds of kilobases (kb) in size (8–11). These domains tend to spatially segregate from each other (9, 12, 13) and, in Drosophila, correspond to the banding patterns of polytene chromosomes (14, 15). At length scales from several hundred kilobases to several megabases, the power-law scaling of Hi-C contact frequency is consistent with a fractal-globule polymer model (7, 16), whereas, within TADs, Hi-C contact maps are better described by a loop-extrusion model (17, 18). Super-resolution imaging shows that chromatin domains in different epigenetic states adopt distinct folding configurations with different power-law scaling properties (13). Whether the ideal fractal-globule model can describe chromatin at length scales beyond several megabases remains an open question (19, 20). Hi-C analyses have also revealed two multi-TAD compartments, compartments A and B, which are enriched with active and inactive chromatin, respectively (7, 21). However, because the contact maps used to identify these compartments were derived from ensemble averaging of many chromosomes, it is unclear whether the A-B compartments are structures that exist in individual chromosomes inside single cells and, if so, how the two compartments are spatially arranged with respect to each other. To answer questions regarding the spatial organization of individual chromosomes requires methods that directly visualize the conformation of single chromosomes in single cells.
Fluorescence in situ hybridization (FISH) provides a powerful means to directly image the spatial organization of chromosomes, especially when used to simultaneously target two or more genomic loci (e.g. 19, 20, 22–24). In one effort, a three-color barcoding approach has been used to simultaneously label multiple chromatin loci to trace the conformation of a chromosome arm in Drosophila blastoderm embryos (24). Nonetheless, routine tracing of the complex 3D folding path of chromosomes has remained challenging because of the difficulties associated with simultaneously imaging and unambiguously identifying many genomic regions on interphase chromosomes. Here, we report a multiplexed FISH method that enables sequential imaging of many genomic regions for 3D tracing of individual chromosomes in the nucleus and the use of this method to study the spatial arrangements of TADs and compartments in chromosomes 20, 21, 22 and X of human diploid (XX) IMR90 cells.
To map the 3D spatial positions of TADs, delineated here as genomic domains based on ensemble-averaged Hi-C maps (8), along an entire chromosome, we labeled the central 100-kb regions of TADs using a dual-oligonucleotide version of Oligopaints (25, 26), wherein each TAD was targeted with 1000 distinct “primary” oligonucleotide probes and a companion, “secondary” probe (Fig 1A and fig. S1). Each of the 1000 primary probes consisted of a unique targeting sequence complementary to a given sequence within the TAD, and a nongenomic region, called Mainstreet, that contained a readout sequence shared by all 1000 probes but unique for each TAD. The secondary probe contained a sequence complementary to the readout sequence on Mainstreet. We used the genomic coordinates of TADs derived from Hi-C (8) to design the targeting sequences of primary probes and produced these probes with a high-yield enzymatic amplification method (27). We exploited a similar hybridization and imaging protocol as we previously described in multiplexed error robust fluorescence in situ hybridization (MERFISH) (27) but with some modifications. First, we hybridized all primary probes to the chromosome of interest (fig. S1, Hyb 0), imaged the sample, and located the chromosome in the nucleus. We then photobleached the sample and performed a series of secondary hybridizations, separated by photobleaching, in order to sequentially label and image individual TADs (Fig. 1A and fig. S1, Hyb 1, Hyb 2, …). Each round of secondary hybridization employed two different secondary probes, respectively labeled with two spectrally distinct dyes, enabling us to visualize two TADs simultaneously using two-color, 3D fluorescence imaging with z-stepping. The centroid positions of the 3D images of individual TADs were used to approximate their positions in x, y and z.
Fig. 1.
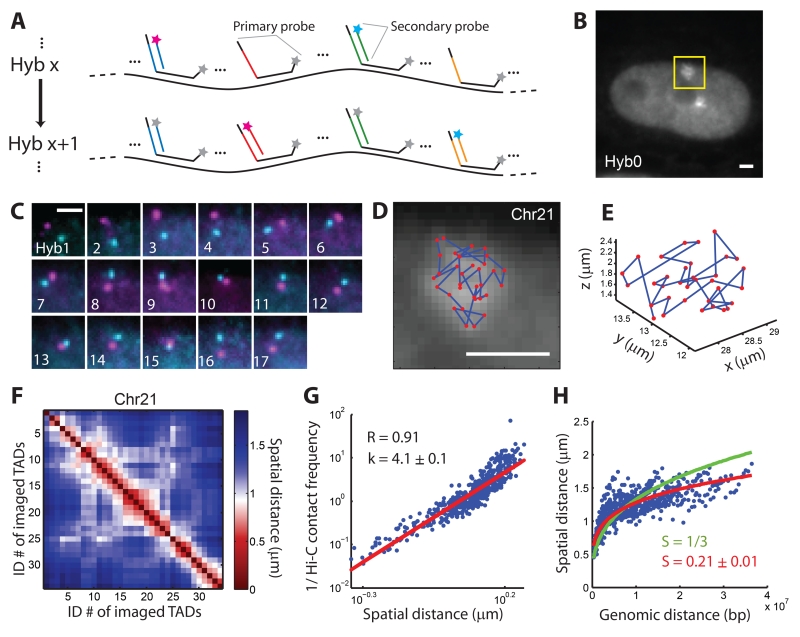
Mapping the spatial organization of the central 100-kb regions of all 34 TADs in chromosome 21 (Chr21) of IMR90 cells. (A) A simplified scheme of the imaging approach. All primary probes are first hybridized to the targeted chromosome, after which secondary probes targeting each TAD are sequentially hybridized to the readout sequences on the primary probes, imaged, and then bleached. In each round of secondary hybridization, two different secondary probes tagged with dyes of different colors allowed simultaneous visualization of two TADs. More details are depicted in fig. S1. (B) Image of an IMR90 cell after the primary hybridization (Hyb 0) with primary probes targeting all TADs in Chr21. The two bright patches, one marked by a yellow box, correspond to the two copies of Chr21 in this diploid cell. (C) Images of the yellow-boxed region in (B) after each round of secondary hybridization (Hyb 1-17). (D) Positions of the 34 TADs of the chromosome were plotted as red dots overlaid on the Hyb 0 image. Scale bars in (B-D): 2 μm. (E) TAD positions plotted in 3D. (F) Mean spatial distance matrix for the 34 TADs, with each element of the matrix corresponding to the mean spatial distance between a pair of TADs. (G) Inverse Hi-C contact frequency between each pair of TADs versus their mean spatial distance. The correlation coefficient (R) and the slope of a fitted line (k) are shown. Contact frequency is calculated as the total Hi-C counts between two TADs normalized to their genomic lengths (8). (H) Mean spatial distance versus genomic distance for all pairs of TADs. The lines are power-law function fits with either a pre-defined scaling exponent (S = 1/3, green) or with S as a fitting parameter (red). Data from 120 individual chromosomes were used to generate (F-H).
We first used this method to image chromosome 21 (Chr21) in interphase IMR90 cells. The Hyb 0 image showed that, when imaged together, the fluorescent signals from all 34 TADs of Chr21 coalesced into a continuous patch (Fig. 1B). The 17 rounds of secondary hybridization then allowed us to image each of the 34 TADs separately (Fig. 1C), determine the 3D position of each TAD, and trace the 3D path of this chromosome at the TAD level (Fig. 1D, E). To characterize the organization of Chr21, we traced 120 copies of Chr21 in many cells, calculated the mean spatial distance between each pair of TADs (averaged over all 120 chromosomes), and constructed a pair-wise mean spatial distance matrix for the 34 TADs (Fig. 1F).
To compare our measurements with previous Hi-C data (8), we correlated the mean spatial distance matrix with the corresponding Hi-C contact frequency matrix of Chr21 (fig. S2). Remarkably, the mean spatial distance showed high correlation with the inverse contact frequency between TADs, with a Pearson correlation coefficient of 0.91 across nearly three orders of magnitude in contact frequency (Fig. 1G). Such a strong correlation between the results from two different methods provided a cross validation for both methods at the TAD-to-chromosome length scales probed in this work. The relationship between the spatial distance and contact frequency also provides a valuable measure for exploring chromosome organization. A mean-field approximation predicts that the contact frequency should be inversely proportional to the 3rd power of the mean spatial distance, whereas the power for real chromatin is expected to be bigger than 3 (19). Our data showed that the Hi-C contact frequency was inversely proportional to the 4th power of the mean spatial distance (Fig. 1G, scaling exponent k = 4.1 ± 0.1, 95% confidence interval (CI), N = 120 chromosomes). We also analyzed the distributions of the spatial distances between pairs of TADs (fig. S3), and found that the Hi-C contact frequency scaled linearly with the probability of two TADs coming into spatial proximity (fig. S4). These results suggest a calibration function to convert Hi-C contact frequencies into mean spatial distances at TAD-to-chromosome length scales, though it remains to be determined whether this calibration extends to sub-TAD scales where the correlation between the Hi-C contact frequency and spatial proximity may be weaker (28).
In addition, our data showed that the mean spatial distance between TADs scaled with their genomic distance to ~1/5th power (Fig. 1H, scaling exponent S = 0.21 ± 0.01, 95% CI, N = 120 chromosomes). This value deviated from the 1/3rd power expected from the ideal fractal-globule polymer model (19). The deviation was most pronounced for large genomic distances, whereas data points with genomic distances less than 7 Mb showed a scaling exponent close to 1/3 (fig. S5), consistent with previous results (7, 19). Interestingly, a previous simulation of confined, unknotted, finite-sized polymers showed a deviation of the scaling exponent from 1/3 at large length scales (29), suggesting a possible physical model to explain our experimental observation.
Next, we determined whether the spatial positions of TADs are partitioned into distinct compartments by implementing a normalization analysis similar to that performed for the Hi-C data (7). First, we normalized the mean spatial distance matrix to the expected spatial distance at each genomic distance as predicted by the power-law scaling shown in Fig. 1H. The normalized spatial distance matrix showed a pattern with alternating regions of large and small values (Fig. 2A), suggesting the existence of two sub-groups of TADs. Next, we calculated the Pearson correlation coefficient between each pair of columns in the normalized distance matrix, defined this coefficient as the correlation between the two corresponding TADs, and constructed a Pearson correlation matrix for all TAD pairs (Fig. 2B). This Pearson correlation matrix showed a plaid pattern, consistent with the existence of two compartments with TADs from the same compartment being positively correlated. For comparison, we used a similar approach (7) to analyze the Hi-C data (8) and obtained a nearly identical Pearson correlation matrix (Fig. 2C), suggesting that the two compartments observed in our imaging data correspond to the A-B compartments identified by Hi-C analysis (7, 21). To assign each TAD to a compartment, we performed a principal component analysis on the Pearson correlation matrix derived from the normalized spatial distances, and assigned TADs with positive and negative values along the first principle component to compartments A and B, respectively (Fig. 2D). Nearly identical assignment was obtained by applying the principal component analysis directly to the normalized spatial distance matrix (fig. S6). We further observed that histone modifications for active chromatin (30, 31) and inactive chromatin (32) were enriched in compartments A and B, respectively (fig. S7), consistent with previous Hi-C analysis (7). We then analyzed the scaling relationship between inverse Hi-C contact frequency and mean spatial distance for pairs of TADs that are either within the same compartment or cross-compartment, and found that cross-compartment TAD pairs gave a moderately higher scaling exponent (fig. S8).
Fig. 2.
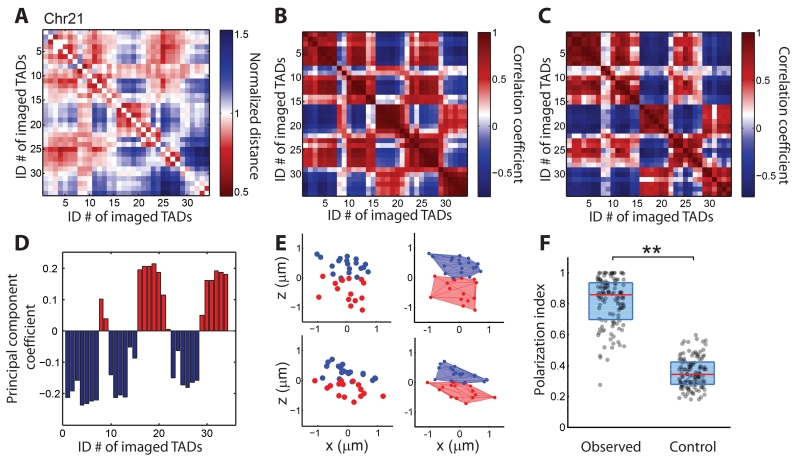
Spatial organization of compartments in individual chromosomes of Chr21. (A) Normalized spatial distance matrix for the 34 TADs, normalized over the expected spatial distances determined by the power-law function fit in Fig. 1H (red line). (B) Pearson correlation matrix of the 34 TADs, determined from the normalized spatial distance matrix in (A). (C) Pearson correlation matrix of the 34 TADs calculated from previous Hi-C data (8). (D) Assignment of TADs to compartment A (red bars) or compartment B (blue bars) based on a principal component analysis of the Pearson correlation matrix shown in (B). (E) Left panels: spatial position maps of compartment-A TADs (red) and compartment-B TADs (blue) in two individual chromosomes. For better visualization, the chromosomes were rotated so that the polarization axis connecting the centroids of compartments A and B is aligned along the z axis. Right panels: corresponding 3D convex hull plots. (F) Polarization index values measured for individual chromosomes (observed) in comparison with those derived from a randomization control where the compartment assignments were randomized while maintaining the total number of TADs in each compartment. The non-zero control values arose from fluctuations associated with the finite number of TADs per chromosome, which provides a baseline for comparison. Each dot corresponds to the polarization index of a single chromosome, the red lines represent the median values, and the blue boxes represent the 25% – 75% quantiles. **: p < 0.001 (Wilcoxon test). Data from 120 individual chromosomes were used to generate (A), (B), (D), and (F).
The above population-averaged analyses cannot reveal whether the higher correlation observed between TADs in the same compartment represents transient proximity between these TADs or whether the two compartments are physical structures that exist in individual chromosomes; nor can they reveal how compartments are spatially arranged, e.g. whether one compartment wraps around the other to form a radial organization within a single chromosome, or whether the two compartments are arranged in a side-by-side, polarized fashion. To address these questions, we examined the spatial positions of the central regions of TADs in single chromosomes. Remarkably, most individual chromosomes in single cells showed a spatially polarized arrangement of compartment-A and compartment-B TADs (Fig. 2E). To quantify the polarized separation of the compartments in individual chromosomes, we defined a polarization index as (1−Vs∕VA)(1−Vs∕VB), where VA and VB are the convex hull volumes of the two compartments and Vs is their shared volume. If the two compartments perfectly overlap with each other, or if one compartment wraps around the other, the polarization index should equal zero; on the other hand, if the two compartments are completely separated in space in a polarized fashion, the polarization index should equal one (fig. S9). The measured polarization index values of Chr21 were indeed close to 1, with a median value of 0.86, substantially larger than the values derived from a randomization control (Fig. 2F).
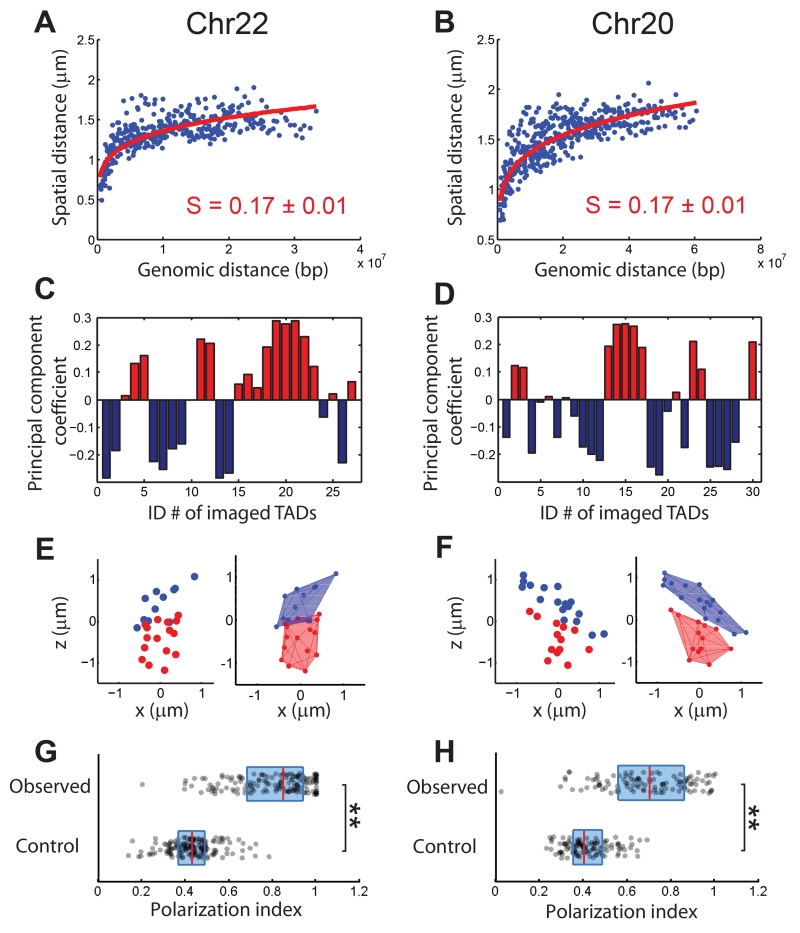
To test whether the above findings were chromosome-specific, we traced the positions of the central 100-kb regions of TADs in Chr22 and Chr20 by imaging all 27 TADs in Chr22 and 30 of the 60 TADs (every other one) in Chr20, and found conclusions similar to those described for Chr21. First, the Hi-C contact frequency was inversely proportional to the 4th power of the mean spatial distance between TADs (figs. S10A and S11A). Second, the mean spatial distance between TADs scaled with genomic distance to a similar albeit slightly smaller power than in Chr21 (Fig. 3A, B), substantially deviating from the 1/3rd power predicted by the ideal fractal-globule model. Third, analysis based on spatial distances showed that TADs in Chr22 and Chr20 were partitioned into two spatial compartments (Fig. 3C, D; figs. S10B-E and S11B-E), with assignments nearly identical to those obtained from our analysis on Hi-C data. These two compartments were again spatially organized in a polarized, side-by-side fashion in individual chromosomes (Fig. 3E-H), though the degree of polarized separation is moderately smaller in Chr20. Whether these findings extend to all other autosomes remains to be determined.
Fig. 3.
Spatial organization of the central 100-kb regions of TADs in chromosome 22 (Chr22) and chromosome 20 (Chr20). (A, B) Mean spatial distance versus genomic distance for Chr22 (A) and Chr20 (B). Power-law function fits are shown as red lines, and the scaling exponents (S) are shown. (C, D) Compartment assignments of TADs based on principal component analyses of the Pearson correlation matrix for Chr22 (fig. S10D) and Chr20 (fig. S11D). Blue bars: compartment B. Red bars: compartment A. (E, F) Spatial position maps of compartment-A TADs (red) and compartment-B TADs (blue) in single chromosomes for Chr22 (E) and Chr20 (F), plotted without (left) or with (right) 3D convex hulls. (G, H) Polarization index values measured for individual chromosomes for Chr22 (G) and Chr20 (H) (observed) in comparison with those of the randomization control (control). The dots, red lines, and blue boxes are defined as in Fig. 2F. **: p < 0.001 (Wilcoxon test). Data from ~150 individual chromosomes were used to generate (A), (C), and (G) and data from ~110 individual chromosomes were used to generate (B), (D), and (H).
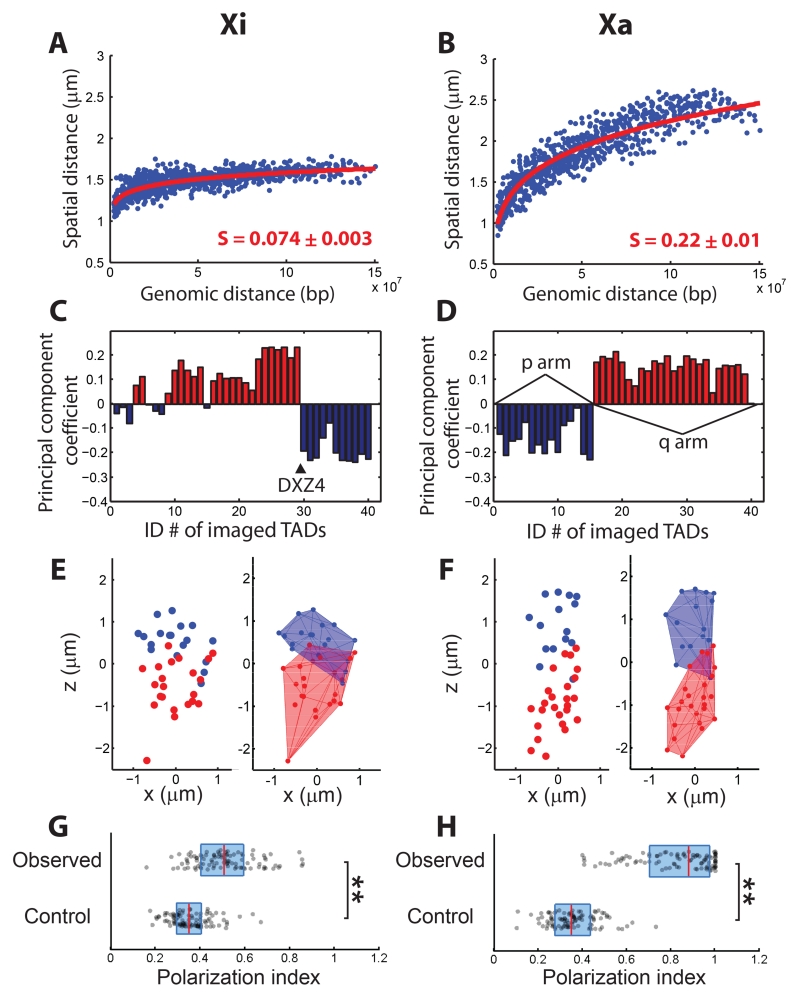
Finally, we traced the positions of the central 100-kb regions of TADs in the X chromosome (ChrX). We imaged 40 TADs (out of 86 total), spanning the whole chromosome at relatively uniform intervals. It is known that one of the two copies of ChrX in female mammalian cells undergoes X-inactivation (33, 34). We used TAD coordinates obtained from the combined Hi-C data (8) of both active and inactive copies of ChrX (Xa and Xi) to determine labeling sites but note that the TAD structures are attenuated or absent on Xi (9, 35). We distinguished Xa and Xi by immunostaining of macroH2A.1 (fig. S12), a histone variant enriched in Xi (35). The mean spatial distance matrices of Xi and Xa were strikingly different with the Xi matrix elements being substantially more homogenous and mostly having smaller values than the Xa matrix elements (Fig. S13A, B). Indeed, fitting a power law function to the spatial versus genomic distance plot yielded a remarkably small scaling exponent of S = 0.074 ± 0.003 (95% CI, N = 95 chromosomes) for Xi (Fig. 4A), whereas the scaling exponent for Xa (S = 0.22 ± 0.01, 95% CI, N = 95 chromosomes) remained similar to those of Chr20, Chr21, and Chr22 (Fig. 4B). These observations suggest that Xi was not only more compact (36) but also adopted a spatially more intermixed chromatin arrangement with more homogeneous inter-loci distances, reminiscent of the chromatin organization observed for Polycomb-repressed domains using super-resolution imaging (13). Given the enrichment of Polycomb group proteins on Xi (33, 34), these observations suggest a potentially general mechanism to induce such a compact and highly intermixed chromatin folding configuration.
Fig. 4.
Spatial organization of the central 100-kb regions of TADs in inactive and active X (Xi and Xa) chromosomes. The TAD structures are attenuated or absent on Xi (9, 35) and hence, for Xi, the term “TAD” simply represents imaged genomic loci. (A, B) Mean spatial distance versus genomic distance for Xi (A) and Xa (B). Power-law function fits are shown as red lines and the scaling exponents (S) are shown. (C, D) Compartment assignments for Xi (C) and Xa (D) based on principal component analyses of the Pearson correlation matrix for Xi (fig. S13C) and Xa (fig. S13D). Positions of the DXZ4 macrosatellite in Xi, and the p and q arms in Xa are indicated. (E, F) Spatial position maps of TADs in single Xi (E) and Xa (F) chromosomes, without (left) or with (right) 3D convex hulls. (G, H) Polarization index measured for Xi (G) and Xa (H) (observed) in comparison with those of the randomization control (control). The dots, red lines, and blue boxes are defined as in Fig. 2F. **: p < 0.001 (Wilcoxon test). Data from 95 individual chromosomes were used to generate (A-D), (G) and (H).
Notably, ChrX also formed two compartments, but the compartmentalization schemes were different for Xi and Xa. Consistent with previous allele-specific Hi-C analyses (21, 35, 37), Xi was largely partitioned into two contiguous compartments (also called superdomains or megadomains) separated on the genomic map by the DXZ4 macrosatellite (Fig. 4C and fig. S13C, E, G). Such a scheme might result from the ability of the DXZ4 element to recruit the chromatin insulator CTCF to Xi but not to Xa (38). The Xa TADs were also partitioned into two spatial compartments, but the two compartments corresponded instead to the p and q arms of the chromosome (Fig. 4D and fig. S13D, F, H). Interestingly, the two compartments in both Xa and Xi were again spatially organized in a polarized, side-by-side manner in individual chromosomes (Fig. 4E-H). However, the degree of polarized segregation was notably smaller for Xi (Fig. 4G), consistent with our observation of more intermixed chromatin in Xi. It is worth noting that within the individual arms of Xa, TADs were further partitioned into two sub-compartments (fig. S14A-C), one of which appeared to be relatively enriched with histone modifications for active chromatin (fig. S14D), implying that these sub-compartments potentially correspond to the A-B compartments.
Our observation that compartments A and B are organized in a spatially polarized manner in single chromosomes in most cells suggest that these are relatively stable physical structures that are present most of the time in individual cells. The fact that we observed this spatial organization for all three examined autosomes (Chr20, 21, and 22) also supports the possibility that these structures are functionally important and maintained by specific mechanisms (2, 4). Since compartments A and B consist mainly of active and inactive chromatin, respectively (7, 21), their presence may serve to locally enrich for transcription machinery and/or epigenetic regulators, and thus enhance the efficient use of these molecular resources. The interactions that maintain these compartments could be direct and/or indirect, i.e. some chromatin binding factors may directly cross-link TADs belonging to the same compartment, or some factors may recruit TADs to pre-defined nuclear areas to form compartments. The large-scale extension of chromatin upon activation (39, 40) may also contribute to the separation of inactive and active chromatin. Finally, we observed distinct compartmentalization schemes for inactive and active X chromosomes, with Xi being partitioned into two contiguous compartments separated by the DXZ4 element (21, 35, 37) and Xa being partitioned according to the p and q arms, though individual arms of Xa may be further partitioned into A and B compartments. Together, these results suggest that the spatial organization of chromatin domains may play an important role in gene regulation and that this organization could be altered to facilitate different chromosomal functions.
Supplementary Material
supplementary material
Acknowledgement
We thank Bryan Harada, Kok Hao Chen, Alistair Boettiger and Guy Nir for helpful discussions. This work is in part supported by the National Institutes of Health (to X.Z. and to C.-t.W.). S.W. is supported in part by a Jane Coffin Childs Fellowship. X.Z. is a Howard Hughes Medical Institute Investigator.
Footnotes
Materials and Methods
References 41-51
Figs. S1 to S15
Tables S1-S8
References
- 1.Misteli T. Beyond the sequence: cellular organization of genome function. Cell. 2007;128:787–800. doi: 10.1016/j.cell.2007.01.028. [DOI] [PubMed] [Google Scholar]
- 2.Gorkin DU, Leung D, Ren B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell. 2014;14:762–75. doi: 10.1016/j.stem.2014.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Levine M, Cattoglio C, Tjian R. Looping back to leap forward: transcription enters a new era. Cell. 2014;157:13–25. doi: 10.1016/j.cell.2014.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dekker J, Mirny L. The 3D Genome as Moderator of Chromosomal Communication. Cell. 2016;164:1110–1121. doi: 10.1016/j.cell.2016.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bickmore WA, van Steensel B. Genome architecture: domain organization of interphase chromosomes. Cell. 2013;152:1270–84. doi: 10.1016/j.cell.2013.02.001. [DOI] [PubMed] [Google Scholar]
- 6.Cremer T, Cremer M. Chromosome territories. Cold Spring Harb. Perspect. Biol. 2010;2:a003889. doi: 10.1101/cshperspect.a003889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lieberman-Aiden E, et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–93. doi: 10.1126/science.1181369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dixon JR, et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature. 2012;485:376–380. doi: 10.1038/nature11082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Nora EP, et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature. 2012;485:381–385. doi: 10.1038/nature11049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sexton T, et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell. 2012;148:458–72. doi: 10.1016/j.cell.2012.01.010. [DOI] [PubMed] [Google Scholar]
- 11.Hou C, Li L, Qin ZS, Corces VG. Gene density, transcription, and insulators contribute to the partition of the Drosophila genome into physical domains. Mol. Cell. 2012;48:471–84. doi: 10.1016/j.molcel.2012.08.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Fabre PJ, et al. Nanoscale spatial organization of the HoxD gene cluster in distinct transcriptional states. Proc. Natl. Acad. Sci. 2015;112:201517972. doi: 10.1073/pnas.1517972112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Boettiger AN, et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature. 2016;529:418–422. doi: 10.1038/nature16496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Eagen KP, Hartl TA, Kornberg RD. Stable Chromosome Condensation Revealed by Chromosome Conformation Capture. Cell. 2015;163:934–46. doi: 10.1016/j.cell.2015.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ulianov SV, et al. Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res. 2016;26:70–84. doi: 10.1101/gr.196006.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mirny L. a. The fractal globule as a model of chromatin architecture in the cell. Chromosom. Res. 2011;19:37–51. doi: 10.1007/s10577-010-9177-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fudenberg G, et al. Formation of Chromosomal Domains by Loop Extrusion. Cell Rep. 2016;15:2038–49. doi: 10.1016/j.celrep.2016.04.085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sanborn AL, et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. 2015;112:201518552. doi: 10.1073/pnas.1518552112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Halverson JD, Smrek J, Kremer K, Grosberg AY. From a melt of rings to chromosome territories: the role of topological constraints in genome folding. Reports Prog. Phys. 2014;77:022601. doi: 10.1088/0034-4885/77/2/022601. [DOI] [PubMed] [Google Scholar]
- 20.Mateos-Langerak J, et al. Spatially confined folding of chromatin in the interphase nucleus. Proc. Natl. Acad. Sci. U. S. A. 2009;106:3812–7. doi: 10.1073/pnas.0809501106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rao SSPSP, et al. A 3D Map of the Human Genome at Kilobase Resolution Reveals Principles of Chromatin Looping. Cell. 2014;159:1665–1680. doi: 10.1016/j.cell.2014.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yokota H, van den Engh G, Hearst JE, Sachs RK, Trask BJ. Evidence for the organization of chromatin in megabase pair-sized loops arranged along a random walk path in the human G0/G1 interphase nucleus. J. Cell Biol. 1995;130:1239–49. doi: 10.1083/jcb.130.6.1239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bystricky K, Heun P, Gehlen L, Langowski J, Gasser SM. Long-range compaction and flexibility of interphase chromatin in budding yeast analyzed by high-resolution imaging techniques. Proc. Natl. Acad. Sci. U. S. A. 2004;101:16495–500. doi: 10.1073/pnas.0402766101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lowenstein MG, Goddard TD, Sedat JW. Long-range interphase chromosome organization in Drosophila: a study using color barcoded fluorescence in situ hybridization and structural clustering analysis. Mol. Biol. Cell. 2004;15:5678–92. doi: 10.1091/mbc.E04-04-0289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Beliveau BJ, et al. Versatile design and synthesis platform for visualizing genomes with Oligopaint FISH probes. Proc. Natl. Acad. Sci. U. S. A. 2012;109:21301–6. doi: 10.1073/pnas.1213818110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Beliveau BJ, et al. Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes. Nat. Commun. 2015;6:7147. doi: 10.1038/ncomms8147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chen KH, Boettiger AN, Moffitt JR, Wang S, Zhuang X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science. 2015;348:aaa6090. doi: 10.1126/science.aaa6090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Williamson I, et al. Spatial genome organization: contrasting views from chromosome conformation capture and fluorescence in situ hybridization. Genes Dev. 2014;28:2778–91. doi: 10.1101/gad.251694.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Imakaev MV, Tchourine KM, Nechaev SK, Mirny LA. Effects of topological constraints on globular polymers. Soft Matter. 2015;11:665–71. doi: 10.1039/c4sm02099e. [DOI] [PubMed] [Google Scholar]
- 30.Hawkins RD, et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell. 2010;6:479–91. doi: 10.1016/j.stem.2010.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bernstein BE, et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 2010;28:1045–8. doi: 10.1038/nbt1010-1045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chandra T, et al. Independence of repressive histone marks and chromatin compaction during senescent heterochromatic layer formation. Mol. Cell. 2012;47:203–14. doi: 10.1016/j.molcel.2012.06.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lessing D, Anguera MC, Lee JT. X chromosome inactivation and epigenetic responses to cellular reprogramming. Annu. Rev. Genomics Hum. Genet. 2013;14:85–110. doi: 10.1146/annurev-genom-091212-153530. [DOI] [PubMed] [Google Scholar]
- 34.Galupa R, Heard E. X-chromosome inactivation: new insights into cis and trans regulation. Curr. Opin. Genet. Dev. 2015;31:57–66. doi: 10.1016/j.gde.2015.04.002. [DOI] [PubMed] [Google Scholar]
- 35.Minajigi A, et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science. 2015;349:aab2276. doi: 10.1126/science.aab2276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Naughton C, Sproul D, Hamilton C, Gilbert N. Analysis of active and inactive X chromosome architecture reveals the independent organization of 30 nm and large-scale chromatin structures. Mol. Cell. 2010;40:397–409. doi: 10.1016/j.molcel.2010.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Deng X, et al. Bipartite structure of the inactive mouse X chromosome. Genome Biol. 2015;16:152. doi: 10.1186/s13059-015-0728-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Horakova AH, Moseley SC, McLaughlin CR, Tremblay DC, Chadwick BP. The macrosatellite DXZ4 mediates CTCF-dependent long-range intrachromosomal interactions on the human inactive X chromosome. Hum. Mol. Genet. 2012;21:4367–77. doi: 10.1093/hmg/dds270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Tumbar T, Sudlow G, Belmont AS. Large-Scale Chromatin Unfolding and Remodeling Induced by VP16 Acidic Activation Domain. J. Cell Biol. 1999;145:1341–1354. doi: 10.1083/jcb.145.7.1341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Levesque MJ, Raj A. Single-chromosome transcriptional profiling reveals chromosomal gene expression regulation. Nat. Methods. 2013;10:246–8. doi: 10.1038/nmeth.2372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Boyle S, Rodesch MJ, Halvensleben HA, Jeddeloh JA, Bickmore WA. Fluorescence in situ hybridization with high-complexity repeat-free oligonucleotide probes generated by massively parallel synthesis. Chromosome Res. 2011;19:901–9. doi: 10.1007/s10577-011-9245-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Yamada NA, et al. Visualization of fine-scale genomic structure by oligonucleotide-based high-resolution FISH. Cytogenet. Genome Res. 2011;132:248–54. doi: 10.1159/000322717. [DOI] [PubMed] [Google Scholar]
- 43.Camacho C, et al. BLAST+: architecture and applications. BMC Bioinformatics. 2009;10:421. doi: 10.1186/1471-2105-10-421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Rouillard J-M, Zuker M, Gulari E. OligoArray 2.0: design of oligonucleotide probes for DNA microarrays using a thermodynamic approach. Nucleic Acids Res. 2003;31:3057–62. doi: 10.1093/nar/gkg426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Cremer M, Müller S, Köhler D, Brero A, Solovei I. Cell Preparation and Multicolor FISH in 3D Preserved Cultured Mammalian Cells. CSH Protoc. 2007;2007:pdb.prot4723. doi: 10.1101/pdb.prot4723. [DOI] [PubMed] [Google Scholar]
- 46.Branco MR, Pombo A. Intermingling of Chromosome Territories in Interphase Suggests Role in Translocations and Transcription-Dependent Associations. PLoS Biol. 2006;4:e138. doi: 10.1371/journal.pbio.0040138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Solovei I, et al. Spatial preservation of nuclear chromatin architecture during three-dimensional fluorescence in situ hybridization (3D-FISH) Exp. Cell Res. 2002;276:10–23. doi: 10.1006/excr.2002.5513. [DOI] [PubMed] [Google Scholar]
- 48.Markaki Y, et al. The potential of 3D-FISH and super-resolution structured illumination microscopy for studies of 3D nuclear architecture: 3D structured illumination microscopy of defined chromosomal structures visualized by 3D (immuno)-FISH opens new perspectives for stud. Bioessays. 2012;34:412–26. doi: 10.1002/bies.201100176. [DOI] [PubMed] [Google Scholar]
- 49.Wang S, Moffitt JR, Dempsey GT, Xie XS, Zhuang X. Characterization and development of photoactivatable fluorescent proteins for single-molecule-based superresolution imaging. Proc. Natl. Acad. Sci. U. S. A. 2014;111:8452–7. doi: 10.1073/pnas.1406593111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Jones SA, Shim S-H, He J, Zhuang X. Fast, three-dimensional super-resolution imaging of live cells. Nat. Methods. 2011;8:499–508. doi: 10.1038/nmeth.1605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yaffe E, Tanay A. Probabilistic modeling of Hi-C contact maps eliminates systematic biases to characterize global chromosomal architecture. Nat. Genet. 2011;43:1059–65. doi: 10.1038/ng.947. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
supplementary material