HLA-B-associated transcript 3 (Bat3)/Scythe is essential for p300-mediated acetylation of p53 - PubMed (original) (raw)
HLA-B-associated transcript 3 (Bat3)/Scythe is essential for p300-mediated acetylation of p53
Toru Sasaki et al. Genes Dev. 2007.
Abstract
In response to DNA damage, p53 undergoes post-translational modifications (including acetylation) that are critical for its transcriptional activity. However, the mechanism by which p53 acetylation is regulated is still unclear. Here, we describe an essential role for HLA-B-associated transcript 3 (Bat3)/Scythe in controlling the acetylation of p53 required for DNA damage responses. Depletion of Bat3 from human and mouse cells markedly impairs p53-mediated transactivation of its target genes Puma and p21. Although DNA damage-induced phosphorylation, stabilization, and nuclear accumulation of p53 are not significantly affected by Bat3 depletion, p53 acetylation is almost completely abolished. Bat3 forms a complex with p300, and an increased amount of Bat3 enhances the recruitment of p53 to p300 and facilitates subsequent p53 acetylation. In contrast, Bat3-depleted cells show reduced p53-p300 complex formation and decreased p53 acetylation. Furthermore, consistent with our in vitro findings, thymocytes from Bat3-deficient mice exhibit reduced induction of puma and p21, and are resistant to DNA damage-induced apoptosis in vivo. Our data indicate that Bat3 is a novel and essential regulator of p53-mediated responses to genotoxic stress, and that Bat3 controls DNA damage-induced acetylation of p53.
Figures
Figure 1.
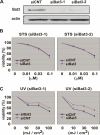
Bat3 is necessary for DNA damage response. (A) Efficient Bat3 KD by siRNA oligo-duplexes. U2OS cells were transfected with control (siCNT) or Bat3 (siBat3-1 and siBat3-2) RNA oligo-duplexes, and Bat3 protein levels were assessed 48 h later by immunoblotting. (actin) Loading control. (B,C) Reduced DNA damage-induced apoptosis. The cells in A were treated with staurosporine (STS) (B) or UV (C) as indicated. Percentage of cell viability was assayed at 16 h post-treatment by annexin-V staining. Data shown are one trial representative of four independent experiments.
Figure 2.
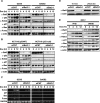
Impaired induction of p53 target genes in Bat3-depleted cells. (A,B) Impaired protein expression of p53 target genes. U2OS and SaOS2 cells (A) and p53 wild-type and p53-null HCT116 cells (B) were transfected with siCNT or siBat3-1 RNA oligo-duplexes and treated with Dox (0.4 μg/mL) for the indicated times. Protein levels of p53, Hdm2, p21, Bax, and Puma were determined by Western blotting. (C) Impaired p53 target gene transcription. U2OS and SaOS2 cells were transfected and treated as in A, and levels of Hdm2, p21, Bax, and Puma transcripts were determined by semiquantitative RT–PCR analysis. (GAPDH) Loading control. (D) Establishment of a Bat3 mutant resistant to RNAi. U2OS cells subjected to Bat3 KD with siBat3-1 RNA oligo-duplexes were transfected with WT-B3 and ΔRNAi-B3. Protein levels of Bat3 were determined as in A. (E) Rescue of the defective induction of p53 target genes by introducing ΔRNAi-B3. U2OS cells subjected to Bat3 KD with siBat3-1 RNA oligo-duplexes were transfected with WT-B3 and ΔRNAi-B3. The cells were then treated with Dox (0.4 μg/mL) for the indicated times. Protein levels of p53 and its targets were determined as in A.
Figure 3.
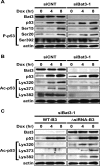
Bat3 is necessary for DNA damage-induced p53 acetylation. (A) Normal p53 phosphorylation. U2OS cells were transfected with siCNT or siBat3 RNA oligo-duplexes and treated 48 h later with Dox (0.4 μg/mL). Protein samples were prepared at the indicated time points post-Dox. Phosphorylation of p53 on Ser15, Ser20, and Ser392 (P-p53) was detected using phospho-specific antibodies. (B) Impaired DNA damage-induced p53 acetylation. U2OS cells were transfected and treated as in A. Extracts were immunoprecipitated with anti-p53 antibody (DO-1) and immunoblotted with antibodies specific for acetylated Lys320, Lys373, and Lys382 (Ac-p53). (C) Rescue of the impaired DNA damage-induced p53 acetylation by the ΔRNAi-B3. U2OS cells were subjected to Bat3 KD and 24 h later they were transfected with WT-B3 and ΔRNAi-B3. The cells were then treated with Dox as in A. Protein levels of acetylated p53 were examined as in B. Data shown are representative of four independent preparations.
Figure 4.
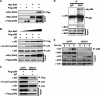
Bat3 binds to p300 and enhances p300-mediated p53 acetylation. (A) Binding of Bat3 to p300. 293T cells were transfected with empty vector (−), Flag-p300, Myc-Bat3, or Flag-p300 plus Myc-Bat3 as indicated. Extracts prepared at 48 h post-transfection were immunoprecipitated using beads conjugated to anti-Flag antibody. The immunoprecipitated samples were analyzed by Western blotting using anti-Myc antibody. (B) Bat3-mediated enhancement of p53–p300 binding and p53 acetylation. H1299 cells were transfected with expression vectors for p53 (0.3 μg), p300-Flag (0.3 μg), and Bat3-Myc (0.1, 0.2, 0.3, or 0.6 μg). The total amount of DNA transfected was equalized by adding empty vector. At 48 h post-transfection, extracts were immunoprecipitated as in A, followed by immunoblotting with anti-p53 antibody (polyclonal). The extracts were also immunoprecipitated with anti-p53 antibody (DO-1), followed by immunoblotting with anti-acetylated p53 (Lys382). (C) Suppression of p53–p300 binding and p53 acetylation by Bat3 KD. H1299 cells were transfected with control or Bat3-specific siRNA oligo-duplexes and cotransfected 24 h later with Flag-p300 (0.6 μg) and p53 (0.6 μg). After another 24 h, extracts were immunoprecipitated to examine p53–p300 interaction and p53 acetylation as in B. (D) Binding of endogenous Bat3 and p300. Extract prepared from U2OS cells was immunoprecipitated using anti-Bat3 antibody or control IgG. The immunoprecipitated samples were analyzed by Western blotting using anti-p300 antibody. (E) Reduced interaction of endogenous p53 and p300 following Bat3 KD. U2OS cells were transfected with control or Bat3-specific siRNA oligo-duplexes and treated 48 h later with Dox (0.4 μg/mL) for the indicated times. Extracts were immunoprecipitated with anti-p300 antibody followed by immunoblotting with anti-p53 antibody. For A_–_E, “Input” represents the protein levels of the indicated molecules in the original extracts as determined by conventional Western blotting.
Figure 5.
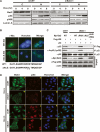
Nuclear localization of Bat3 is necessary for its activity. (A) Biochemical demonstration. U2OS cells transfected with control or Bat3-specific siRNA oligo-duplexes were subjected to Dox (0.4 μg/mL) treatment. After 0, 4, or 8 h, cytosolic (C) and nuclear (N) fractions were prepared from extracts. Protein levels of Bat3, p53, and p300 were determined by immunoblotting. (Lamin A) Loading control for nuclear samples. (B) Establishment of a nuclear-localization-defective mutant of Bat3. U2OS cells transfected with Myc-tagged WT-B3 and ΔNLS-B3 were stained with anti-myc-FITC antibody. Nuclei were visualized using Hoechst 33258 (blue), merge of myc and Hoechst 33258 staining. The point mutations introduced in ΔNLS-B3 are indicated by asterisk. (C) Failure of the enhancement of p53–p300 binding and p53 acetylation by ΔNLS-B3. H1299 cells were transfected with control or Bat3-specific siRNA oligo-duplexes and cotransfected 24 h later with WT-B3, ΔRNAi-B3, ΔNLS-B3, or ΔNLS/ΔRNAi-B3. At 48 h post-transfection, p53–p300 binding and p53 acetylation were examined as in Figure 3. (D) Cellular demonstration. U2OS cells were transfected as in A and treated with Dox for 0 or 8 h, at which times the cells were fixed, permeabilized, and stained with anti-Bat3 (panels a,e,i,m) or anti-p53 (DO-1; panels b,f,j,n) antibodies, followed by anti-rabbit IgG (green) or anti-mouse IgG (red). Nuclei were visualized using Hoechst 33258 (blue; panels c,g,k,o). (Panels d,h,l,p) Merge of Bat3 and Hoechst 33258 staining. Data shown are representative of four independent preparations.
Figure 6.
Bat3 is required for p53-mediated transcription. Impaired recruitment of p53 to the p21, Puma, and Hdm2 promoters in the absence of Bat3. U2OS cells transfected with control or Bat3-specific siRNA oligo-duplexes were treated with Dox (0.4 μg/mL) for 8 h, and extracts were immunoprecipitated with anti-p53, anti-acetylated p53 (Lys382 and 320), anti-p300, or Bat3 antibodies. The p53-binding regions of the p21, Puma, and Hdm2 promoters were amplified by PCR. Data shown are representative of four independent preparations.
Figure 7.
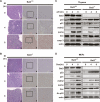
Impaired γ-IR-induced apoptosis in Bat3-deficient thymus. (A,B) Histological determination of impaired apoptosis. Six-week-old Bat3+/+ (A) and Bat3−/− (B) littermate mice were left untreated or subjected to 10 Gy whole-body γ-IR. After 4 or 8 h, the thymus was removed and cut in half. One half was fixed and subjected to histological analysis. The other half was used for biochemical analysis, as shown in C. Serial sections of paraffin-embedded thymus were stained with H&E (_A_-,B, panels -a,d,g) and subjected to in situ end-labeling (IEL) of fragmented DNA (_A_-,B, panels b,c,e,f,h,i). For each IEL preparation, two magnifications are shown (_A_-,B, panels b,e,h: 10 × 10; _A_-,B, panels c,f,i: 10 × 25). Data shown are representative of three independent preparations. (C) Impaired induction of p53 target genes and apoptosis. Half of the thymic lysates prepared in A were analyzed by Western blotting for protein levels of the indicated proteins. (casp 3) Activated caspase-3. (D) Impaired induction of p53 target genes and p53 acetylation in Dox-treated Bat3-deficient MEFs. Protein lysates prepared from Dox-treated Bat3+/+ and Bat3−/− MEFs were analyzed by Western blotting for protein levels of the indicated proteins. Data shown are representative of three independent preparations.
Similar articles
- E6 oncoprotein represses p53-dependent gene activation via inhibition of protein acetylation independently of inducing p53 degradation.
Thomas MC, Chiang CM. Thomas MC, et al. Mol Cell. 2005 Jan 21;17(2):251-64. doi: 10.1016/j.molcel.2004.12.016. Mol Cell. 2005. PMID: 15664194 - HIPK2 contributes to PCAF-mediated p53 acetylation and selective transactivation of p21Waf1 after nonapoptotic DNA damage.
Di Stefano V, Soddu S, Sacchi A, D'Orazi G. Di Stefano V, et al. Oncogene. 2005 Aug 18;24(35):5431-42. doi: 10.1038/sj.onc.1208717. Oncogene. 2005. PMID: 15897882 - DNA damage activates p53 through a phosphorylation-acetylation cascade.
Sakaguchi K, Herrera JE, Saito S, Miki T, Bustin M, Vassilev A, Anderson CW, Appella E. Sakaguchi K, et al. Genes Dev. 1998 Sep 15;12(18):2831-41. doi: 10.1101/gad.12.18.2831. Genes Dev. 1998. PMID: 9744860 Free PMC article. - To die or not to die: a HAT trick.
Tyteca S, Legube G, Trouche D. Tyteca S, et al. Mol Cell. 2006 Dec 28;24(6):807-8. doi: 10.1016/j.molcel.2006.12.005. Mol Cell. 2006. PMID: 17189182 Review. - The impact of acetylation and deacetylation on the p53 pathway.
Brooks CL, Gu W. Brooks CL, et al. Protein Cell. 2011 Jun;2(6):456-62. doi: 10.1007/s13238-011-1063-9. Epub 2011 Jul 12. Protein Cell. 2011. PMID: 21748595 Free PMC article. Review.
Cited by
- HLA-B-associated transcript 3 (Bat3/Scythe) negatively regulates Smad phosphorylation in BMP signaling.
Goto K, Tong KI, Ikura J, Okada H. Goto K, et al. Cell Death Dis. 2011 Dec 1;2(12):e236. doi: 10.1038/cddis.2011.114. Cell Death Dis. 2011. PMID: 22130070 Free PMC article. - BAG Family Members as Mitophagy Regulators in Mammals.
Pattingre S, Turtoi A. Pattingre S, et al. Cells. 2022 Feb 15;11(4):681. doi: 10.3390/cells11040681. Cells. 2022. PMID: 35203329 Free PMC article. Review. - An optimized genome-wide, virus-free CRISPR screen for mammalian cells.
Xiong K, Karottki KJC, Hefzi H, Li S, Grav LM, Li S, Spahn P, Lee JS, Ventina I, Lee GM, Lewis NE, Kildegaard HF, Pedersen LE. Xiong K, et al. Cell Rep Methods. 2021 Aug 23;1(4):100062. doi: 10.1016/j.crmeth.2021.100062. Epub 2021 Aug 4. Cell Rep Methods. 2021. PMID: 34935002 Free PMC article. - Exploitation of the Host Ubiquitin System: Means by Legionella pneumophila.
Luo J, Wang L, Song L, Luo ZQ. Luo J, et al. Front Microbiol. 2021 Dec 22;12:790442. doi: 10.3389/fmicb.2021.790442. eCollection 2021. Front Microbiol. 2021. PMID: 35003021 Free PMC article. Review. - A soluble fragment of the tumor antigen BCL2-associated athanogene 6 (BAG-6) is essential and sufficient for inhibition of NKp30 receptor-dependent cytotoxicity of natural killer cells.
Binici J, Hartmann J, Herrmann J, Schreiber C, Beyer S, Güler G, Vogel V, Tumulka F, Abele R, Mäntele W, Koch J. Binici J, et al. J Biol Chem. 2013 Nov 29;288(48):34295-303. doi: 10.1074/jbc.M113.483602. Epub 2013 Oct 16. J Biol Chem. 2013. PMID: 24133212 Free PMC article.
References
- Ahn J., Prives C., Prives C. The C-terminus of p53: The more you learn the less you know. Nat. Struct. Biol. 2001;8:730–732. - PubMed
- Appella E., Anderson C.W., Anderson C.W. Post-translational modifications and activation of p53 by genotoxic stresses. Eur. J. Biochem. 2001;268:2764–2772. - PubMed
- Avantaggiati M.L., Ogryzko V., Gardner K., Giordano A., Levine A.S., Kelly K., Ogryzko V., Gardner K., Giordano A., Levine A.S., Kelly K., Gardner K., Giordano A., Levine A.S., Kelly K., Giordano A., Levine A.S., Kelly K., Levine A.S., Kelly K., Kelly K. Recruitment of p300/CBP in p53-dependent signal pathways. Cell. 1997;89:1175–1184. - PubMed
- Ayed A., Mulder F.A., Yi G.S., Lu Y., Kay L.E., Arrowsmith C.H., Mulder F.A., Yi G.S., Lu Y., Kay L.E., Arrowsmith C.H., Yi G.S., Lu Y., Kay L.E., Arrowsmith C.H., Lu Y., Kay L.E., Arrowsmith C.H., Kay L.E., Arrowsmith C.H., Arrowsmith C.H. Latent and active p53 are identical in conformation. Nat. Struct. Biol. 2001;8:756–760. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous