mbs: modifying Hudson's ms software to generate samples of DNA sequences with a biallelic site under selection - PubMed (original) (raw)
mbs: modifying Hudson's ms software to generate samples of DNA sequences with a biallelic site under selection
Kosuke M Teshima et al. BMC Bioinformatics. 2009.
Abstract
Background: The pattern of single nucleotide polymorphisms, or SNPs, contains a tremendous amount of information with respect to the mechanisms of the micro-evolutionary process of a species. The inference of the roles of these mechanisms, including natural selection, relies heavily on computer simulations. A coalescent simulation is extremely powerful in generating a large number of samples of DNA sequences from a population (species) when all mutations are neutral, and Hudson's ms software is frequently used for this purpose.However, it has been difficult to incorporate natural selection into the coalescent framework.
Results: We herein present a software application to generate samples of DNA sequences when there is a biallelic site targeted by selection. This software application, referred to as mbs, is developed by modifying Hudson's ms. The mbs software is so flexible that it can incorporate any arbitrary histories of population size changes and any mode of selection as long as selection is operating on a biallelic site.
Conclusion: mbs provides opportunities to investigate the effect of any mode of selection on the pattern of SNPs under various demography.
Figures
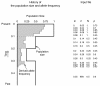
Figure 1
An example of the allele frequency and demographic change. Illustration of an example of the allele frequency and demographic changes (left), and their input format (right).
Similar articles
- msABC: a modification of Hudson's ms to facilitate multi-locus ABC analysis.
Pavlidis P, Laurent S, Stephan W. Pavlidis P, et al. Mol Ecol Resour. 2010 Jul;10(4):723-7. doi: 10.1111/j.1755-0998.2010.02832.x. Epub 2010 Feb 2. Mol Ecol Resour. 2010. PMID: 21565078 - msHOT: modifying Hudson's ms simulator to incorporate crossover and gene conversion hotspots.
Hellenthal G, Stephens M. Hellenthal G, et al. Bioinformatics. 2007 Feb 15;23(4):520-1. doi: 10.1093/bioinformatics/btl622. Epub 2006 Dec 6. Bioinformatics. 2007. PMID: 17150995 - PopGenome: an efficient Swiss army knife for population genomic analyses in R.
Pfeifer B, Wittelsbürger U, Ramos-Onsins SE, Lercher MJ. Pfeifer B, et al. Mol Biol Evol. 2014 Jul;31(7):1929-36. doi: 10.1093/molbev/msu136. Epub 2014 Apr 16. Mol Biol Evol. 2014. PMID: 24739305 Free PMC article. - Discoal: flexible coalescent simulations with selection.
Kern AD, Schrider DR. Kern AD, et al. Bioinformatics. 2016 Dec 15;32(24):3839-3841. doi: 10.1093/bioinformatics/btw556. Epub 2016 Aug 24. Bioinformatics. 2016. PMID: 27559153 Free PMC article. - Coalescence 2.0: a multiple branching of recent theoretical developments and their applications.
Tellier A, Lemaire C. Tellier A, et al. Mol Ecol. 2014 Jun;23(11):2637-52. doi: 10.1111/mec.12755. Epub 2014 May 20. Mol Ecol. 2014. PMID: 24750385 Review.
Cited by
- Automatic inference of demographic parameters using generative adversarial networks.
Wang Z, Wang J, Kourakos M, Hoang N, Lee HH, Mathieson I, Mathieson S. Wang Z, et al. Mol Ecol Resour. 2021 Nov;21(8):2689-2705. doi: 10.1111/1755-0998.13386. Epub 2021 May 3. Mol Ecol Resour. 2021. PMID: 33745225 Free PMC article. - Searching for footprints of positive selection in whole-genome SNP data from nonequilibrium populations.
Pavlidis P, Jensen JD, Stephan W. Pavlidis P, et al. Genetics. 2010 Jul;185(3):907-22. doi: 10.1534/genetics.110.116459. Epub 2010 Apr 20. Genetics. 2010. PMID: 20407129 Free PMC article. - Simulation of genes and genomes forward in time.
Carvajal-Rodríguez A. Carvajal-Rodríguez A. Curr Genomics. 2010 Mar;11(1):58-61. doi: 10.2174/138920210790218007. Curr Genomics. 2010. PMID: 20808525 Free PMC article. - Distinguishing positive selection from neutral evolution: boosting the performance of summary statistics.
Lin K, Li H, Schlötterer C, Futschik A. Lin K, et al. Genetics. 2011 Jan;187(1):229-44. doi: 10.1534/genetics.110.122614. Epub 2010 Nov 1. Genetics. 2011. PMID: 21041556 Free PMC article. - Genetics and evidence for balancing selection of a sex-linked colour polymorphism in a songbird.
Kim KW, Jackson BC, Zhang H, Toews DPL, Taylor SA, Greig EI, Lovette IJ, Liu MM, Davison A, Griffith SC, Zeng K, Burke T. Kim KW, et al. Nat Commun. 2019 Apr 23;10(1):1852. doi: 10.1038/s41467-019-09806-6. Nat Commun. 2019. PMID: 31015412 Free PMC article.
References
- Hudson RR. Gene genealogies and the coalescent process. In: Futuyma D, Antonovics J, editor. Oxford Surveys in Evolutionary Biology. Vol. 7. New York: Oxford University Press; 1990. pp. 1–44.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources