Phylogenetic and phyletic studies of informational genes in genomes highlight existence of a 4 domain of life including giant viruses - PubMed (original) (raw)
Phylogenetic and phyletic studies of informational genes in genomes highlight existence of a 4 domain of life including giant viruses
Mickaël Boyer et al. PLoS One. 2010.
Abstract
The discovery of Mimivirus, with its very large genome content, made it possible to identify genes common to the three domains of life (Eukarya, Bacteria and Archaea) and to generate controversial phylogenomic trees congruent with that of ribosomal genes, branching Mimivirus at its root. Here we used sequences from metagenomic databases, Marseillevirus and three new viruses extending the Mimiviridae family to generate the phylogenetic trees of eight proteins involved in different steps of DNA processing. Compared to the three ribosomal defined domains, we report a single common origin for Nucleocytoplasmic Large DNA Viruses (NCLDV), DNA processing genes rooted between Archaea and Eukarya, with a topology congruent with that of the ribosomal tree. As for translation, we found in our new viruses, together with Mimivirus, five proteins rooted deeply in the eukaryotic clade. In addition, comparison of informational genes repertoire based on phyletic pattern analysis supports existence of a clade containing NCLDVs clearly distinct from that of Eukarya, Bacteria and Archaea. We hypothesize that the core genome of NCLDV is as ancient as the three currently accepted domains of life.
Conflict of interest statement
Competing Interests: The authors have declared that no competing interests exist.
Figures
Figure 1. NCLDV proteins involved in DNA, RNA and protein biosynthesis.
The frames on the right represent DNA processing steps with related enzymes found in NCLDVs, and the frames on the left with grey backgrounds show viruses that have the corresponding enzymes. The first frame from the top right corresponds to the dNTP biosynthesis catalyzed by RNR and ThyA. The second frame shows four enzymes involved in DNA replication: DNAP B, TopoIIA, PCNA and FEN. Proteins involved in these two DNA biosynthesis steps were found in diverse large DNA viruses, such as NCLDVs, Herpesviridae and Myoviridae. The third frame represents the DNA transcription step, in which the RNAP II and the transcription factor TFIIB are involved; viral homologs of these proteins are only found in NCLDVs. The last frame shows Mimiviridae amino-acyl tRNA synthetases (aa tRNA synthetase) involved in mRNA translation. Interestingly, viral lineages were consistently less well represented from nucleotide to protein biosynthesis, with the exception of NCLDVs, which were represented in each step. a.a., amino acid.
Figure 2. Phylogenetic tree of the RNA polymerase II beta subunit.
The ML tree of RNAP II was inferred from a cured alignment of 80 sequences from the six supergroups of Eukarya (blue), Bacteria (purple), Archaea (green) and NCLDVs (red). The tree is unrooted, and values near branches are SH-like local supports computed by the FastTree program and are used as confidence values of tree branches. Scale bar represents the number of estimated changes per position for a unit of branch length.
Figure 3. Phylogenetic tree of the Transcription factor II B (TFIIB).
The TFIIB phylogenetic tree is inferred with Bayesian approach from a cured alignment of 32 sequences (155 conserved positions) from the Eukarya (blue), Archaea (green), NCLDVs (red), and metagenomic databases (black). Bayesian posterior probabilities are mentioned near branches as a percentage and are used as confidence values of tree branches. Scale bar represents the number of estimated changes per position for a unit of branch length.
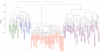
Figure 4. Hierarchical clustering of Eukarya (blue), Bacteria (purple), Archaea (green) and NCLDVs (red) by phyletic pattern.
The phyletic patterns of the putative orthologous sets of informational genes indicating the presence/absence of the respective gene in each cellular organisms and virus were used for the construction of the dendogram tree.
Figure 5. Scenario of NCLDVs emerging from the rhizome of life with roots appearing at the very beginning of life.
This figure represents the living species in the four small pictures according to the current classification of organisms: eukaryotes (represented by yellow cell), bacteria (represented by green cell), Archaea (represented by blue cell) and viruses (represented by magenta colored Mimivirus). An organism classification based on ribosomal proteins allows the discrimination of cellular organisms, but viruses are excluded de facto from historical scenarios for life evolution. Our study shows that eight proteins involved in DNA processing and present both in viruses, particularly in NCLDVs, and in two or three other domains of life could be used for phylogenetic tree reconstruction, displaying schematic topologies for each protein represented by lines colored in light green for RNR, in yellow for ThyA, in blue-green for DNAP B, in blue for TopoIIA, in magenta for PCNA, in orange for FEN, in red for RNAP II and in purple for TFIIB. Topology representations were similar to that based on ribosomes (for RNAP II, FEN, PCNA and TFIIB proteins) and/or with bifurcations (for ThyA, RNR, DNAP B and Topo IIA proteins) representing probable lateral gene transfer. Thus, this figure illustrates that REOs and CEOs (bona fide viruses) share a common set of genes involved in DNA processing that evolved from a common ancestral source of genes. The phenomenon of long-branch attraction makes the identification of the exact deep rooting of each analyzed gene difficult and is therefore hidden by a black mark.
Similar articles
- The giant Cafeteria roenbergensis virus that infects a widespread marine phagocytic protist is a new member of the fourth domain of Life.
Colson P, Gimenez G, Boyer M, Fournous G, Raoult D. Colson P, et al. PLoS One. 2011 Apr 29;6(4):e18935. doi: 10.1371/journal.pone.0018935. PLoS One. 2011. PMID: 21559486 Free PMC article. - Mimiviridae: clusters of orthologous genes, reconstruction of gene repertoire evolution and proposed expansion of the giant virus family.
Yutin N, Colson P, Raoult D, Koonin EV. Yutin N, et al. Virol J. 2013 Apr 4;10:106. doi: 10.1186/1743-422X-10-106. Virol J. 2013. PMID: 23557328 Free PMC article. - The effects of model choice and mitigating bias on the ribosomal tree of life.
Lasek-Nesselquist E, Gogarten JP. Lasek-Nesselquist E, et al. Mol Phylogenet Evol. 2013 Oct;69(1):17-38. doi: 10.1016/j.ympev.2013.05.006. Epub 2013 May 22. Mol Phylogenet Evol. 2013. PMID: 23707703 - Giant viruses and the origin of modern eukaryotes.
Forterre P, Gaïa M. Forterre P, et al. Curr Opin Microbiol. 2016 Jun;31:44-49. doi: 10.1016/j.mib.2016.02.001. Epub 2016 Mar 19. Curr Opin Microbiol. 2016. PMID: 26894379 Review. - Evolutionary genomics of nucleo-cytoplasmic large DNA viruses.
Iyer LM, Balaji S, Koonin EV, Aravind L. Iyer LM, et al. Virus Res. 2006 Apr;117(1):156-84. doi: 10.1016/j.virusres.2006.01.009. Epub 2006 Feb 21. Virus Res. 2006. PMID: 16494962 Review.
Cited by
- Potassium ion channels: could they have evolved from viruses?
Thiel G, Moroni A, Blanc G, Van Etten JL. Thiel G, et al. Plant Physiol. 2013 Jul;162(3):1215-24. doi: 10.1104/pp.113.219360. Epub 2013 May 29. Plant Physiol. 2013. PMID: 23719891 Free PMC article. Review. - Informational gene phylogenies do not support a fourth domain of life for nucleocytoplasmic large DNA viruses.
Williams TA, Embley TM, Heinz E. Williams TA, et al. PLoS One. 2011;6(6):e21080. doi: 10.1371/journal.pone.0021080. Epub 2011 Jun 16. PLoS One. 2011. PMID: 21698163 Free PMC article. - Eukaryotic and Prokaryotic Microbiota Interactions.
Kodio A, Menu E, Ranque S. Kodio A, et al. Microorganisms. 2020 Dec 17;8(12):2018. doi: 10.3390/microorganisms8122018. Microorganisms. 2020. PMID: 33348551 Free PMC article. Review. - Seasonal Dynamics of Algae-Infecting Viruses and Their Inferred Interactions with Protists.
Gran-Stadniczeñko S, Krabberød AK, Sandaa RA, Yau S, Egge E, Edvardsen B. Gran-Stadniczeñko S, et al. Viruses. 2019 Nov 9;11(11):1043. doi: 10.3390/v11111043. Viruses. 2019. PMID: 31717498 Free PMC article. - Zamilon, a novel virophage with Mimiviridae host specificity.
Gaia M, Benamar S, Boughalmi M, Pagnier I, Croce O, Colson P, Raoult D, La Scola B. Gaia M, et al. PLoS One. 2014 Apr 18;9(4):e94923. doi: 10.1371/journal.pone.0094923. eCollection 2014. PLoS One. 2014. PMID: 24747414 Free PMC article.
References
- Moreira D, Lopez-Garcia P. Ten reasons to exclude viruses from the tree of life. Nat Rev Microbiol. 2009;7:306–311. - PubMed
- Krupovic M, Bamford DH. Virus evolution: how far does the double beta-barrel viral lineage extend? Nat Rev Microbiol. 2008;6:941–948. - PubMed
- Raoult D, Audic S, Robert C, Abergel C, Renesto P, et al. The 1.2-megabase genome sequence of Mimivirus. Science. 2004;306:1344–1350. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials