RUVBL1, CPTAC-268 - CPTAC Assay Portal (original) (raw)
Please include the following statement when referencing the CPTAC Assay Portal
We would like to acknowledge the National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC) Assay Portal (assays.cancer.gov) for developing assays and establishing criteria for the assays described in this publication.
Protein Sequence hover to view complete sequence
| 10 | 20 | 30 | 40 | 50 |
|---|---|---|---|---|
| MKIEEVKSTT | KTQRIASHSH | VKGLGLDESG | LAKQAASGLV | GQENAREACG |
| 60 | 70 | 80 | 90 | 100 |
| VIVELIKSKK | MAGRAVLLAG | PPGTGKTALA | LAIAQELGSK | VPFCPMVGSE |
| 110 | 120 | 130 | 140 | 150 |
| VYSTEIKKTE | VLMENFRRAI | GLRIKETKEV | YEGEVTELTP | CETENPMGGY |
| 160 | 170 | 180 | 190 | 200 |
| GKTISHVIIG | LKTAKGTKQL | KLDPSIFESL | QKERVEAGDV | IYIEANSGAV |
| 210 | 220 | 230 | 240 | 250 |
| KRQGRCDTYA | TEFDLEAEEY | VPLPKGDVHK | KKEIIQDVTL | HDLDVANARP |
| 260 | 270 | 280 | 290 | 300 |
| QGGQDILSMM | GQLMKPKKTE | ITDKLRGEIN | KVVNKYIDQG | IAELVPGVLF |
| 310 | 320 | 330 | 340 | 350 |
| VDEVHMLDIE | CFTYLHRALE | SSIAPIVIFA | SNRGNCVIRG | TEDITSPHGI |
| 360 | 370 | 380 | 390 | 400 |
| PLDLLDRVMI | IRTMLYTPQE | MKQIIKIRAQ | TEGINISEEA | LNHLGEIGTK |
| 410 | 420 | 430 | 440 | 450 |
| TTLRYSVQLL | TPANLLAKIN | GKDSIEKEHV | EEISELFYDA | KSSAKILADQ |
| 456 | ||||
| QDKYMK |
Data source: UniProt
Position of Targeted Peptide Analytes Relative to SNPs, Isoforms, and PTMs
Uniprot Database Entry PhosphoSitePlus ®
Click a point on a node
to view detailed assay information below
All other points link out to UniProt
Phosphorylation Acetylation Ubiquitylation Other
loading

Assay Details for CPTAC-268 Collapse assay details
Data source: Panorama
Official Gene Symbol
RUVBL1
Peptide Sequence
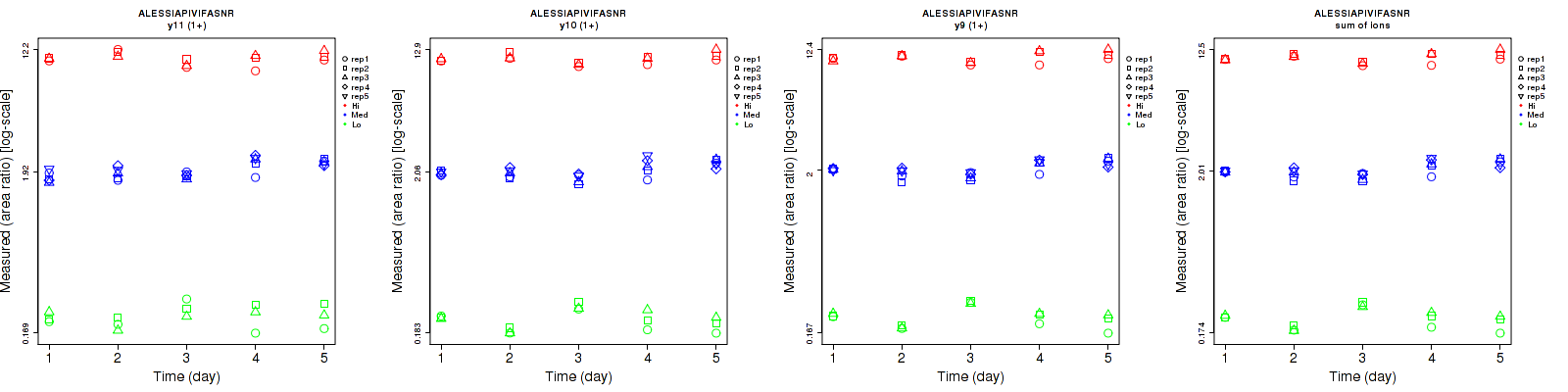
ALESSIAPIVIFASNR
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
318
Peptide End
333
CPTAC ID
CPTAC-268
Peptide Molecular Mass
1,686.9305
Species
Homo sapiens (Human)
Assay Type
Direct MRM
Matrix
cell line lysate pool
Submitting Laboratory
Fred Hutchinson Cancer Research Center
Submitting Lab PI
Amanda Paulovich
Publication
View Details (opens in a new window)
Demonstrating the feasibility of large-scale development of standardized assays to quantify human proteins. Kennedy JJ, Abbatiello SE, Kim K, Yan P, Whiteaker JR, Lin C, Kim JS, Zhang Y, Wang X, Ivey RG, Zhao L, Min H, Lee Y, Yu MH, Yang EG, Lee C, Wang P, Rodriguez H, Kim Y, Carr SA, Paulovich AG. Nat Methods. 2014 Feb;11(2):149-55. doi: 10.1038/nmeth.2763. Epub 2013 Dec 8. PMID: 24317253
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
5500 QTRAP (ABSCIEX)
Internal Standard
peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C,15N
LC
nanoLC-Ultra 2D, cHiPLC-nanoflex (Eksigent)
Column Packing
ChromXP C18-CL, 3 um, 120A
Column Dimensions
150 x 0.075 mm
Flow Rate
500 nL / min
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV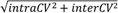 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y11 (1+) | 14.2 | 7.7 | 6.6 | 14.5 | 13.2 | 8.3 | 20.3 | 15.3 | 10.6 | 15 | 25 | 15 |
| y10 (1+) | 7.8 | 7.2 | 4.9 | 15.6 | 11.5 | 6.5 | 17.4 | 13.6 | 8.1 | 15 | 25 | 15 |
| y9 (1+) | 5.9 | 5.8 | 4.8 | 15.4 | 10.6 | 6.9 | 16.5 | 12.1 | 8.4 | 15 | 25 | 15 |
| sum | 6.6 | 6 | 4.6 | 14.8 | 10.9 | 6.5 | 16.2 | 12.4 | 8 | 15 | 25 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y11 (1+) | 14.2 | 7.7 | 6.6 | 14.5 | 13.2 | 8.3 | 20.3 | 15.3 | 10.6 | 15 | 25 | 15 |
| y10 (1+) | 7.8 | 7.2 | 4.9 | 15.6 | 11.5 | 6.5 | 17.4 | 13.6 | 8.1 | 15 | 25 | 15 |
| y9 (1+) | 5.9 | 5.8 | 4.8 | 15.4 | 10.6 | 6.9 | 16.5 | 12.1 | 8.4 | 15 | 25 | 15 |
| sum | 6.6 | 6 | 4.6 | 14.8 | 10.9 | 6.5 | 16.2 | 12.4 | 8 | 15 | 25 | 15 |
Additional Resources and Comments
Assay Details for CPTAC-269 Collapse assay details
Data source: Panorama
Official Gene Symbol
RUVBL1
Peptide Sequence
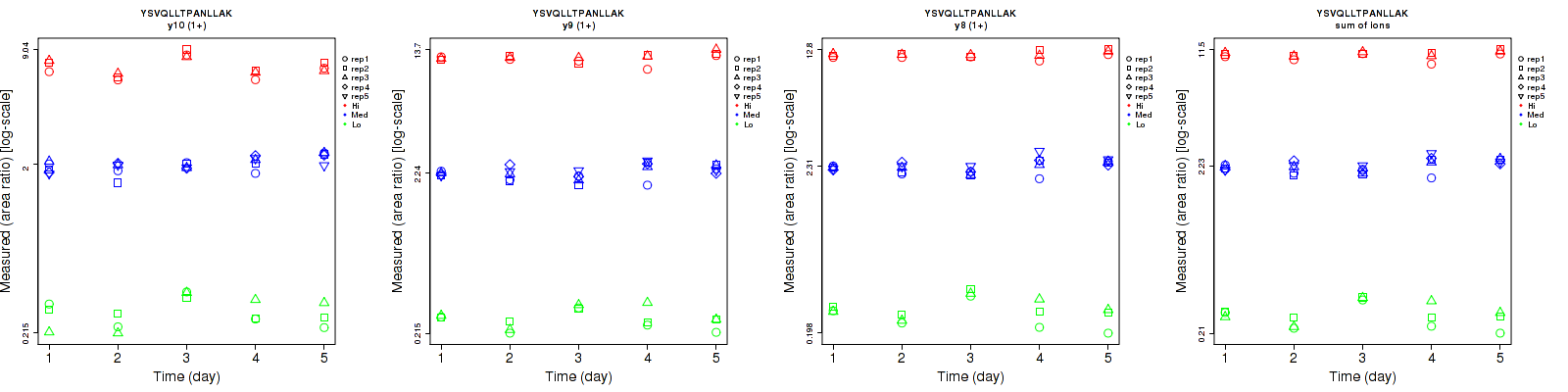
YSVQLLTPANLLAK
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
405
Peptide End
418
CPTAC ID
CPTAC-269
Peptide Molecular Mass
1,529.8817
Species
Homo sapiens (Human)
Assay Type
Direct MRM
Matrix
cell line lysate pool
Submitting Laboratory
Fred Hutchinson Cancer Research Center
Submitting Lab PI
Amanda Paulovich
Publication
View Details (opens in a new window)
Demonstrating the feasibility of large-scale development of standardized assays to quantify human proteins. Kennedy JJ, Abbatiello SE, Kim K, Yan P, Whiteaker JR, Lin C, Kim JS, Zhang Y, Wang X, Ivey RG, Zhao L, Min H, Lee Y, Yu MH, Yang EG, Lee C, Wang P, Rodriguez H, Kim Y, Carr SA, Paulovich AG. Nat Methods. 2014 Feb;11(2):149-55. doi: 10.1038/nmeth.2763. Epub 2013 Dec 8. PMID: 24317253
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
5500 QTRAP (ABSCIEX)
Internal Standard
peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C,15N
LC
nanoLC-Ultra 2D, cHiPLC-nanoflex (Eksigent)
Column Packing
ChromXP C18-CL, 3 um, 120A
Column Dimensions
150 x 0.075 mm
Flow Rate
500 nL / min
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV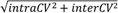 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y10 (1+) | 13.5 | 7.2 | 5.7 | 18.9 | 9.8 | 12.8 | 23.2 | 12.2 | 14 | 15 | 25 | 15 |
| y9 (1+) | 8.4 | 7.8 | 5 | 13.5 | 9.9 | 6.4 | 15.9 | 12.6 | 8.1 | 15 | 25 | 15 |
| y8 (1+) | 10.8 | 6.5 | 3.8 | 18.4 | 9 | 3.5 | 21.3 | 11.1 | 5.2 | 15 | 25 | 15 |
| sum | 9.4 | 6.3 | 3.7 | 16.5 | 9 | 4.2 | 19 | 11 | 5.6 | 15 | 25 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y10 (1+) | 13.5 | 7.2 | 5.7 | 18.9 | 9.8 | 12.8 | 23.2 | 12.2 | 14 | 15 | 25 | 15 |
| y9 (1+) | 8.4 | 7.8 | 5 | 13.5 | 9.9 | 6.4 | 15.9 | 12.6 | 8.1 | 15 | 25 | 15 |
| y8 (1+) | 10.8 | 6.5 | 3.8 | 18.4 | 9 | 3.5 | 21.3 | 11.1 | 5.2 | 15 | 25 | 15 |
| sum | 9.4 | 6.3 | 3.7 | 16.5 | 9 | 4.2 | 19 | 11 | 5.6 | 15 | 25 | 15 |