SERPINA1, CPTAC-651 - CPTAC Assay Portal (original) (raw)
Please include the following statement when referencing the CPTAC Assay Portal
We would like to acknowledge the National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC) Assay Portal (assays.cancer.gov) for developing assays and establishing criteria for the assays described in this publication.
SERPINA1
Protein Sequence hover to view complete sequence
non-CPTAC-1063: View additional LSITGTYDLK data
| 10 | 20 | 30 | 40 | 50 |
|---|---|---|---|---|
| MPSSVSWGIL | LLAGLCCLVP | VSLAEDPQGD | AAQKTDTSHH | DQDHPTFNKI |
| 60 | 70 | 80 | 90 | 100 |
| TPNLAEFAFS | LYRQLAHQSN | STNIFFSPVS | IATAFAMLSL | GTKADTHDEI |
| 110 | 120 | 130 | 140 | 150 |
| LEGLNFNLTE | IPEAQIHEGF | QELLRTLNQP | DSQLQLTTGN | GLFLSEGLKL |
| 160 | 170 | 180 | 190 | 200 |
| VDKFLEDVKK | LYHSEAFTVN | FGDTEEAKKQ | INDYVEKGTQ | GKIVDLVKEL |
| 210 | 220 | 230 | 240 | 250 |
| DRDTVFALVN | YIFFKGKWER | PFEVKDTEEE | DFHVDQVTTV | KVPMMKRLGM |
| 260 | 270 | 280 | 290 | 300 |
| FNIQHCKKLS | SWVLLMKYLG | NATAIFFLPD | EGKLQHLENE | LTHDIITKFL |
| 310 | 320 | 330 | 340 | 350 |
| ENEDRRSASL | HLPKLSITGT | YDLKSVLGQL | GITKVFSNGA | DLSGVTEEAP |
| 360 | 370 | 380 | 390 | 400 |
| LKLSKAVHKA | VLTIDEKGTE | AAGAMFLEAI | PMSIPPEVKF | NKPFVFLMIE |
| 410 | 418 | |||
| QNTKSPLFMG | KVVNPTQK |
Data source: UniProt
Position of Targeted Peptide Analytes Relative to SNPs, Isoforms, and PTMs
Uniprot Database Entry PhosphoSitePlus ®
Click a point on a node
to view detailed assay information below
All other points link out to UniProt
Phosphorylation Acetylation Ubiquitylation Other
loading

Assay Details for CPTAC-651 Collapse assay details
Data source: Panorama
Official Gene Symbol
SERPINA1
Peptide Modified Sequence
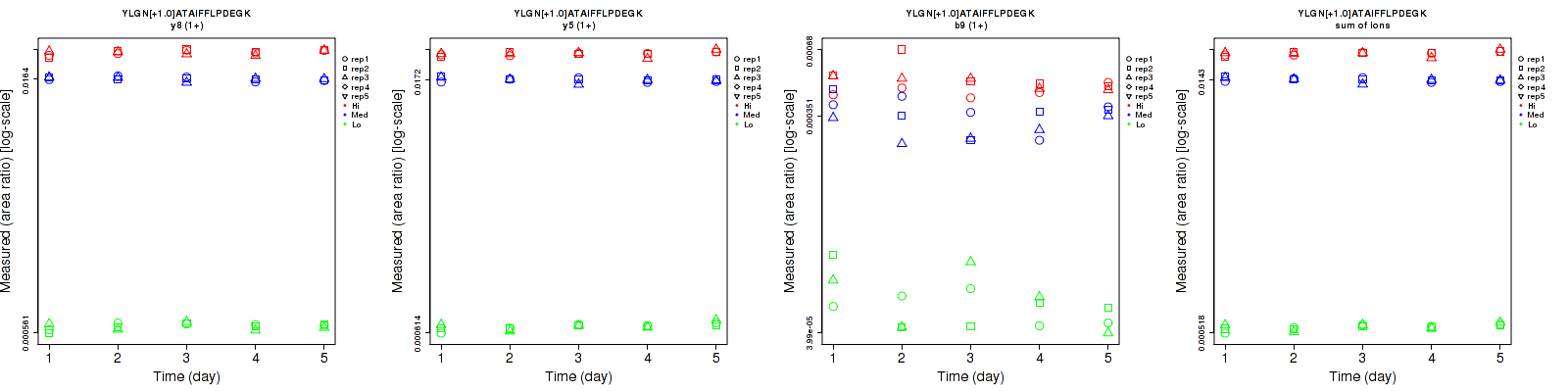
YLGN[+1.0]ATAIFFLPDEGK
Modification Type
Deamidated (NQ)
Protein - Site of Modification
271
Peptide - Site of Modification
4
Peptide Start
268
Peptide End
283
CPTAC ID
CPTAC-651
Peptide Molecular Mass
1,755.8719
Species
Homo sapiens (Human)
Assay Type
Enrichment PRM
Enrichment Method
N-linked glycopeptide solid phase extraction
Matrix
serum
Submitting Laboratory
Johns Hopkins University
Submitting Lab PI
Daniel Chan, Hui Zhang, Zhen Zhang
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
Q-Exactive (ThermoFisher)
Internal Standard
synthetic peptide
Peptide Standard Purity
Crude
Peptide Standard Label Type
13C and 15N at C-terminus K
LC
Dionex UltiMate 3000 RSLCnano LC (ThermoFisher)
Column Packing
Acclaim PepMap100 C18, 5um (Trap); Acclaim PepMap RSLC C18, 2um (Analytical)
Column Dimensions
300um x 5mm (Trap); 75um x 25cm (Analytical)
Flow Rate
0.5 uL/min
Assay Multiplexing Expand assay panel
Johns Hopkins University-A
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV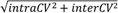 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y8 (1+) | 3.6 | 2.2 | 2.5 | 4.4 | 2.5 | 3.3 | 5.7 | 3.3 | 4.1 | 15 | 15 | 15 |
| y5 (1+) | 2.6 | 2.4 | 2.1 | 4.2 | 2.8 | 2.9 | 4.9 | 3.7 | 3.6 | 15 | 15 | 15 |
| b9 (1+) | 20.6 | 14.6 | 9.9 | 26.6 | 15.1 | 9.6 | 33.6 | 21 | 13.8 | 15 | 15 | 15 |
| sum | 2.5 | 2.3 | 2 | 3.8 | 2.7 | 2.9 | 4.5 | 3.5 | 3.5 | 15 | 15 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y8 (1+) | 3.6 | 2.2 | 2.5 | 4.4 | 2.5 | 3.3 | 5.7 | 3.3 | 4.1 | 15 | 15 | 15 |
| y5 (1+) | 2.6 | 2.4 | 2.1 | 4.2 | 2.8 | 2.9 | 4.9 | 3.7 | 3.6 | 15 | 15 | 15 |
| b9 (1+) | 20.6 | 14.6 | 9.9 | 26.6 | 15.1 | 9.6 | 33.6 | 21 | 13.8 | 15 | 15 | 15 |
| sum | 2.5 | 2.3 | 2 | 3.8 | 2.7 | 2.9 | 4.5 | 3.5 | 3.5 | 15 | 15 | 15 |
Additional Resources and Comments
Comments
Assay Details for non-CPTAC-1063 Collapse assay details
Data source: Panorama
Official Gene Symbol
SERPINA1
Peptide Sequence
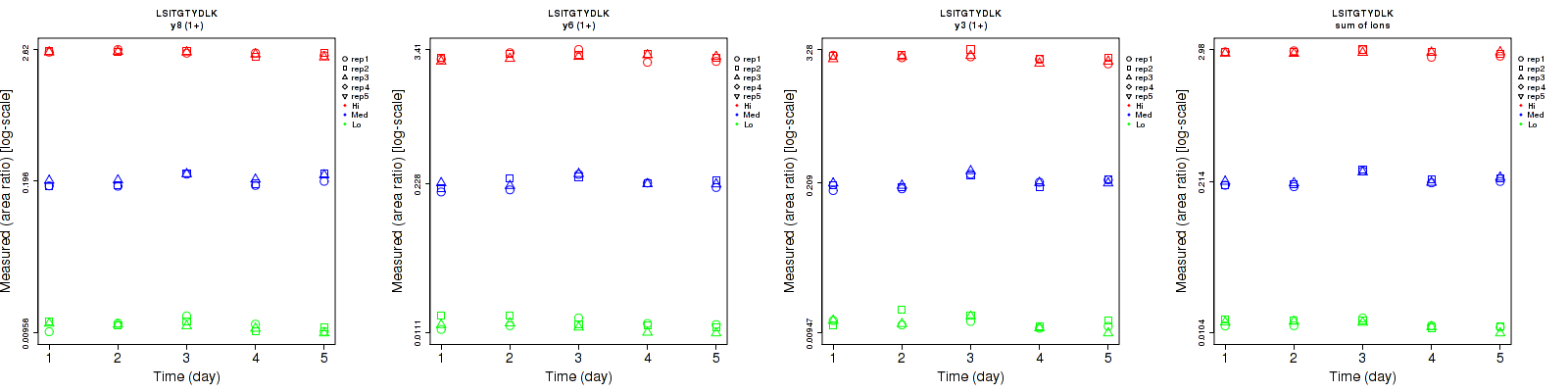
LSITGTYDLK
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
315
Peptide End
324
CPTAC ID
non-CPTAC-1063
Peptide Molecular Mass
1,109.5968
Species
Homo sapiens (Human)
Assay Type
Direct PRM
Matrix
plasma
Submitting Laboratory
UVic-Genome BC Proteomics Centre
Submitting Lab PI
Christoph Borchers
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
6490 Triple Quad (Agilent)
Internal Standard
synthetic peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C and 15N at C-terminus K
LC
1290 LC (Agilent)
Column Packing
Zorbax Eclipse Plus C18, 1.8 µm
Column Dimensions
2.1 x 150 mm
Flow Rate
400 µL/min
Assay Multiplexing Expand assay panel
Johns Hopkins University-A
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV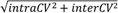 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y8 (2+) | 5.7 | 3.6 | 3.5 | 8.5 | 13.3 | 4.6 | 10.2 | 13.8 | 5.8 | 15 | 15 | 15 |
| y3 (1+) | 9.1 | 5 | 5.5 | 11.6 | 12.6 | 6.9 | 14.7 | 13.6 | 8.8 | 15 | 15 | 15 |
| y6 (1+) | 10.5 | 6.3 | 6.3 | 9.6 | 11.1 | 6.7 | 14.2 | 12.8 | 9.2 | 15 | 15 | 15 |
| y7 (1+) | 6.2 | 6.5 | 5.7 | 9.5 | 14.2 | 6.3 | 11.3 | 15.6 | 8.5 | 15 | 15 | 15 |
| y8 (1+) | 7.1 | 5.6 | 2.9 | 9.7 | 9.9 | 4.5 | 12 | 11.4 | 5.4 | 15 | 15 | 15 |
| sum | 5.2 | 3.9 | 3.6 | 8.2 | 11.8 | 3.7 | 9.7 | 12.4 | 5.2 | 15 | 15 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y8 (2+) | 5.7 | 3.6 | 3.5 | 8.5 | 13.3 | 4.6 | 10.2 | 13.8 | 5.8 | 15 | 15 | 15 |
| y3 (1+) | 9.1 | 5 | 5.5 | 11.6 | 12.6 | 6.9 | 14.7 | 13.6 | 8.8 | 15 | 15 | 15 |
| y6 (1+) | 10.5 | 6.3 | 6.3 | 9.6 | 11.1 | 6.7 | 14.2 | 12.8 | 9.2 | 15 | 15 | 15 |
| y7 (1+) | 6.2 | 6.5 | 5.7 | 9.5 | 14.2 | 6.3 | 11.3 | 15.6 | 8.5 | 15 | 15 | 15 |
| y8 (1+) | 7.1 | 5.6 | 2.9 | 9.7 | 9.9 | 4.5 | 12 | 11.4 | 5.4 | 15 | 15 | 15 |
| sum | 5.2 | 3.9 | 3.6 | 8.2 | 11.8 | 3.7 | 9.7 | 12.4 | 5.2 | 15 | 15 | 15 |