F12, CPTAC-1100 - CPTAC Assay Portal (original) (raw)
Please include the following statement when referencing the CPTAC Assay Portal
We would like to acknowledge the National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC) Assay Portal (assays.cancer.gov) for developing assays and establishing criteria for the assays described in this publication.
Protein Sequence hover to view complete sequence
non-CPTAC-2650: View additional EQPPSLTR data
non-CPTAC-3707: View additional TGPGGVAR data
non-CPTAC-1100: View additional VVGGLVALR data
| 10 | 20 | 30 | 40 | 50 |
|---|---|---|---|---|
| MRALLLLGFL | LVSLESTLSI | PPWEAPKEHK | YKAEEHTVVL | TVTGEPCHFP |
| 60 | 70 | 80 | 90 | 100 |
| FQYHRQLYHK | CTHKGRPGPQ | PWCATTPNFD | QDQRWGYCLE | PKKVKDHCSK |
| 110 | 120 | 130 | 140 | 150 |
| HSPCQKGGTC | VNMPSGPHCL | CPQHLTGNHC | QKEKCFEPQL | LRFFHKNEIW |
| 160 | 170 | 180 | 190 | 200 |
| YRTEQAAVAR | CQCKGPDAHC | QRLASQACRT | NPCLHGGRCL | EVEGHRLCHC |
| 210 | 220 | 230 | 240 | 250 |
| PVGYTGAFCD | VDTKASCYDG | RGLSYRGLAR | TTLSGAPCQP | WASEATYRNV |
| 260 | 270 | 280 | 290 | 300 |
| TAEQARNWGL | GGHAFCRNPD | NDIRPWCFVL | NRDRLSWEYC | DLAQCQTPTQ |
| 310 | 320 | 330 | 340 | 350 |
| AAPPTPVSPR | LHVPLMPAQP | APPKPQPTTR | TPPQSQTPGA | LPAKREQPPS |
| 360 | 370 | 380 | 390 | 400 |
| LTRNGPLSCG | QRLRKSLSSM | TRVVGGLVAL | RGAHPYIAAL | YWGHSFCAGS |
| 410 | 420 | 430 | 440 | 450 |
| LIAPCWVLTA | AHCLQDRPAP | EDLTVVLGQE | RRNHSCEPCQ | TLAVRSYRLH |
| 460 | 470 | 480 | 490 | 500 |
| EAFSPVSYQH | DLALLRLQED | ADGSCALLSP | YVQPVCLPSG | AARPSETTLC |
| 510 | 520 | 530 | 540 | 550 |
| QVAGWGHQFE | GAEEYASFLQ | EAQVPFLSLE | RCSAPDVHGS | SILPGMLCAG |
| 560 | 570 | 580 | 590 | 600 |
| FLEGGTDACQ | GDSGGPLVCE | DQAAERRLTL | QGIISWGSGC | GDRNKPGVYT |
| 610 | 615 | |||
| DVAYYLAWIR | EHTVS |
Data source: UniProt
Position of Targeted Peptide Analytes Relative to SNPs, Isoforms, and PTMs
Uniprot Database Entry PhosphoSitePlus ®
Click a point on a node
to view detailed assay information below
All other points link out to UniProt
Phosphorylation Acetylation Ubiquitylation Other
loading

Assay Details for non-CPTAC-3541 Collapse assay details
Data source: Panorama
Official Gene Symbol
F12
Peptide Sequence
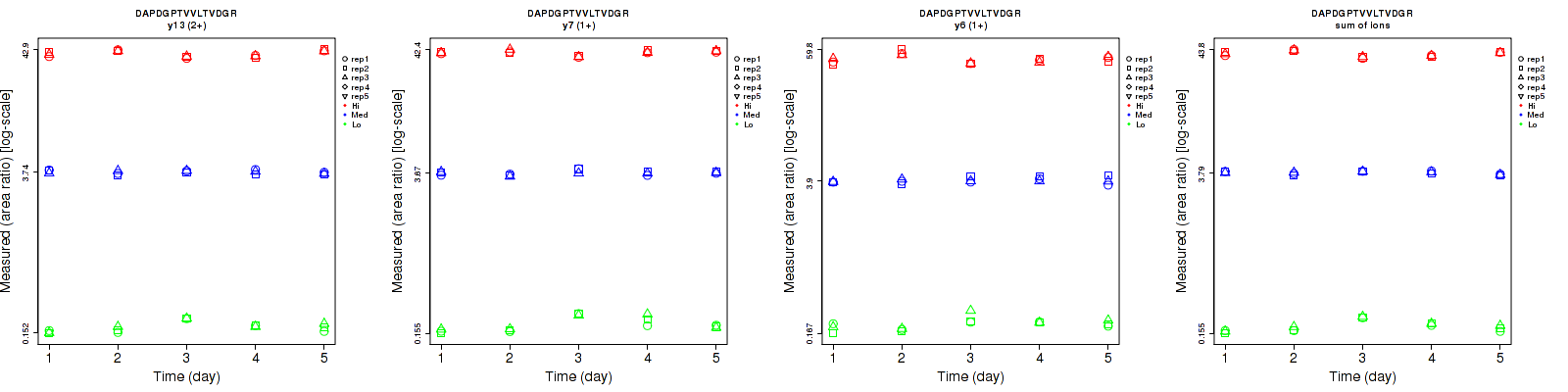
DAPDGPTVVLTVDGR
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
31
Peptide End
45
CPTAC ID
non-CPTAC-3541
Peptide Molecular Mass
1,510.7627
Species
Mus musculus (Mouse)
Assay Type
Direct MRM
Matrix
Plasma
Submitting Laboratory
UVic-Genome BC Proteomics Centre
Submitting Lab PI
Christoph Borchers
Publication
View Details (opens in a new window)
Molecular phenotyping of laboratory mouse strains using 500 multiple reaction monitoring mass spectrometry plasma assays. Michaud SA, Sinclair NJ, Petrošová H, Palmer AL, Pistawka AJ, Zhang S, Hardie DB, Mohammed Y Eshghi A1, Richard VR, Sickmann A, Borchers CH. Commun Biol. 2018 Jun 27;1:78. doi: 10.1038/s42003-018-0087-6. eCollection 2018.
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
Agilent 6490/6495 QQQ
Internal Standard
synthetic peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C and 15N at C-terminus R
LC
1290 LC (Agilent)
Column Packing
Zorbax Eclipse Plus C18, 1.8 µm
Column Dimensions
2.1 x 150 mm
Flow Rate
400 µL/min
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV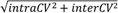 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y13 (2+) | 3.6 | 3.2 | 2.5 | 11.4 | 3.2 | 6.7 | 12 | 4.5 | 7.2 | 15 | 15 | 15 |
| y2 (1+) | 4.9 | 2.9 | 2.8 | 19 | 7.9 | 11.5 | 19.6 | 8.4 | 11.8 | 15 | 15 | 15 |
| y6 (1+) | 6.6 | 5.3 | 4.2 | 10.9 | 4.9 | 9.7 | 12.7 | 7.2 | 10.6 | 15 | 15 | 15 |
| y7 (1+) | 4.4 | 3.2 | 2.2 | 15.7 | 4.4 | 4.7 | 16.3 | 5.4 | 5.2 | 15 | 15 | 15 |
| y10 (1+) | 6.2 | 5.4 | 4.6 | 12.2 | 8 | 12.1 | 13.7 | 9.7 | 12.9 | 15 | 15 | 15 |
| sum | 3.1 | 1.6 | 1.7 | 12 | 3 | 6 | 12.4 | 3.4 | 6.2 | 15 | 15 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y13 (2+) | 3.6 | 3.2 | 2.5 | 11.4 | 3.2 | 6.7 | 12 | 4.5 | 7.2 | 15 | 15 | 15 |
| y2 (1+) | 4.9 | 2.9 | 2.8 | 19 | 7.9 | 11.5 | 19.6 | 8.4 | 11.8 | 15 | 15 | 15 |
| y6 (1+) | 6.6 | 5.3 | 4.2 | 10.9 | 4.9 | 9.7 | 12.7 | 7.2 | 10.6 | 15 | 15 | 15 |
| y7 (1+) | 4.4 | 3.2 | 2.2 | 15.7 | 4.4 | 4.7 | 16.3 | 5.4 | 5.2 | 15 | 15 | 15 |
| y10 (1+) | 6.2 | 5.4 | 4.6 | 12.2 | 8 | 12.1 | 13.7 | 9.7 | 12.9 | 15 | 15 | 15 |
| sum | 3.1 | 1.6 | 1.7 | 12 | 3 | 6 | 12.4 | 3.4 | 6.2 | 15 | 15 | 15 |
Additional Resources and Comments
Assay Details for non-CPTAC-2650 Collapse assay details
Data source: Panorama
Official Gene Symbol
F12
Peptide Sequence
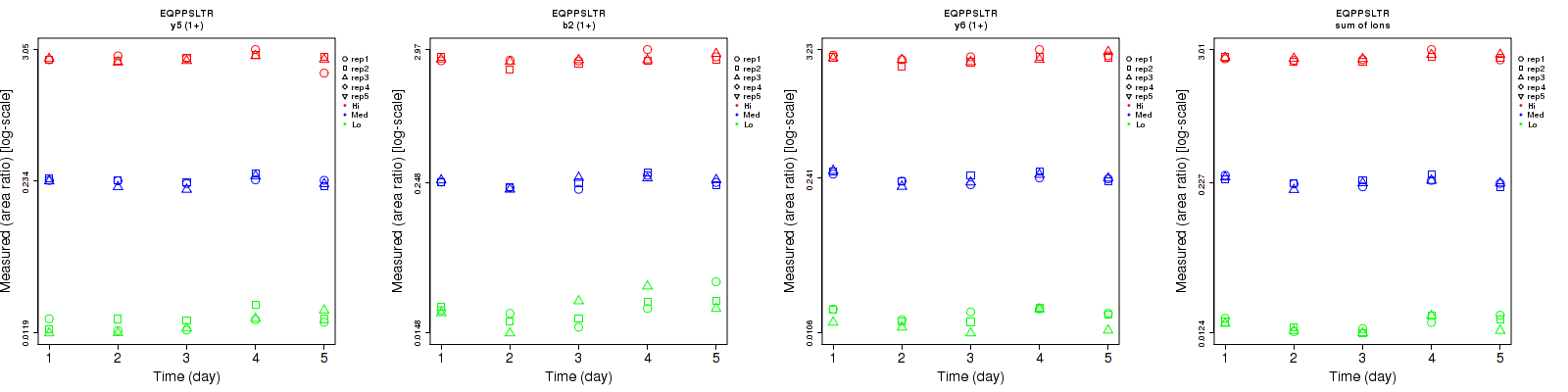
EQPPSLTR
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
346
Peptide End
353
CPTAC ID
non-CPTAC-2650
Peptide Molecular Mass
926.4821
Species
Homo sapiens (Human)
Assay Type
Direct MRM
Matrix
plasma
Submitting Laboratory
UVic-Genome BC Proteomics Centre
Submitting Lab PI
Christoph Borchers
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
6490 Triple Quad (Agilent)
Internal Standard
synthetic peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C and 15N at C-terminus R
LC
1290 LC (Agilent)
Column Packing
Zorbax Eclipse Plus C18, 1.8 µm
Column Dimensions
2.1 x 150 mm
Flow Rate
400 µL/min
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV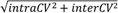 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y6 (2+) | 10.1 | 7 | 4.7 | 15 | 9.7 | 6.2 | 18.1 | 12 | 7.8 | 15 | 15 | 15 |
| b2 (1+) | 19.6 | 5.1 | 6.8 | 27.5 | 10.1 | 8 | 33.8 | 11.3 | 10.5 | 15 | 15 | 15 |
| y5 (2+) | 25.2 | 9.7 | 8.2 | 41 | 11.6 | 10.4 | 48.1 | 15.1 | 13.2 | 15 | 15 | 15 |
| y6 (1+) | 12.5 | 5.8 | 6.9 | 14 | 10.1 | 8.1 | 18.8 | 11.6 | 10.6 | 15 | 15 | 15 |
| y5 (1+) | 13.8 | 5.7 | 6.8 | 16.5 | 7.3 | 8.3 | 21.5 | 9.3 | 10.7 | 15 | 15 | 15 |
| sum | 7.4 | 5.4 | 3.9 | 13.5 | 8.8 | 5.9 | 15.4 | 10.3 | 7.1 | 15 | 15 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y6 (2+) | 10.1 | 7 | 4.7 | 15 | 9.7 | 6.2 | 18.1 | 12 | 7.8 | 15 | 15 | 15 |
| b2 (1+) | 19.6 | 5.1 | 6.8 | 27.5 | 10.1 | 8 | 33.8 | 11.3 | 10.5 | 15 | 15 | 15 |
| y5 (2+) | 25.2 | 9.7 | 8.2 | 41 | 11.6 | 10.4 | 48.1 | 15.1 | 13.2 | 15 | 15 | 15 |
| y6 (1+) | 12.5 | 5.8 | 6.9 | 14 | 10.1 | 8.1 | 18.8 | 11.6 | 10.6 | 15 | 15 | 15 |
| y5 (1+) | 13.8 | 5.7 | 6.8 | 16.5 | 7.3 | 8.3 | 21.5 | 9.3 | 10.7 | 15 | 15 | 15 |
| sum | 7.4 | 5.4 | 3.9 | 13.5 | 8.8 | 5.9 | 15.4 | 10.3 | 7.1 | 15 | 15 | 15 |
Additional Resources and Comments
Assay Details for non-CPTAC-3707 Collapse assay details
Data source: Panorama
Official Gene Symbol
F12
Peptide Sequence
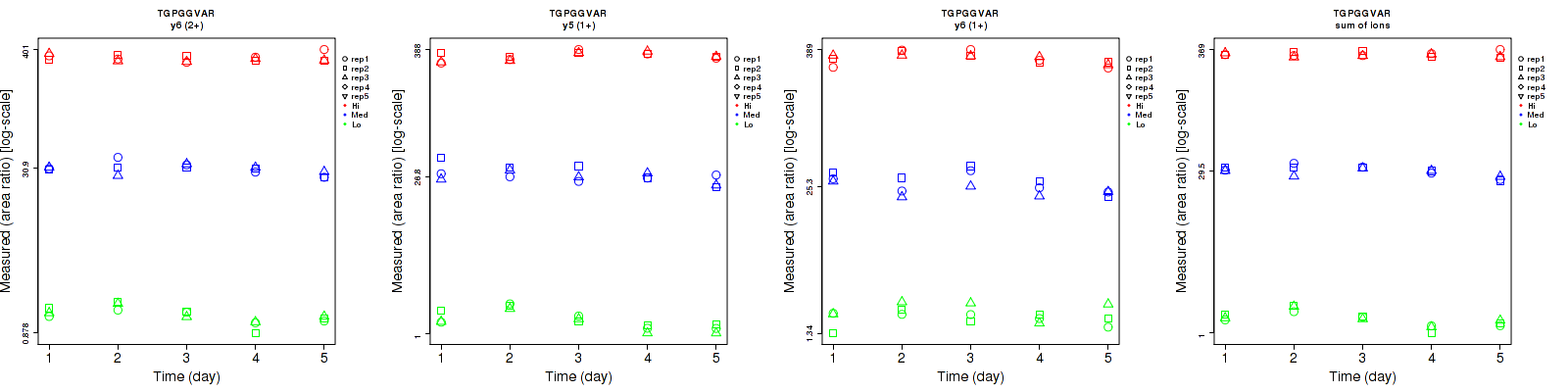
TGPGGVAR
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
153
Peptide End
160
CPTAC ID
non-CPTAC-3707
Peptide Molecular Mass
713.3820
Species
Mus musculus (Mouse)
Assay Type
Direct MRM
Matrix
Plasma
Submitting Laboratory
UVic-Genome BC Proteomics Centre
Submitting Lab PI
Christoph Borchers
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
Agilent 6490/6495 QQQ
Internal Standard
synthetic peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C and 15N at C-terminus R
LC
1290 LC (Agilent)
Column Packing
Zorbax Eclipse Plus C18, 1.8 µm
Column Dimensions
2.1 x 150 mm
Flow Rate
400 µL/min
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV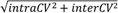 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y6 (2+) | 8.4 | 7.9 | 8.1 | 17 | 11.6 | 8.1 | 19 | 14 | 11.5 | 15 | 15 | 15 |
| y4 (1+) | 31.3 | 26.8 | 13.6 | 36.5 | 35.3 | 19.8 | 48.1 | 44.3 | 24 | 15 | 15 | 15 |
| y5 (1+) | 8.6 | 14.3 | 5.2 | 21.4 | 13.8 | 8.5 | 23.1 | 19.9 | 10 | 15 | 15 | 15 |
| y6 (1+) | 16.7 | 14 | 7.4 | 14.9 | 18 | 11.9 | 22.4 | 22.8 | 14 | 15 | 15 | 15 |
| y7 (1+) | 25.7 | 57.4 | 10.1 | 24.4 | 46.1 | 20.3 | 35.4 | 73.6 | 22.7 | 15 | 15 | 15 |
| sum | 5.5 | 5 | 5.4 | 16.5 | 10.5 | 5.1 | 17.4 | 11.6 | 7.4 | 15 | 15 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y6 (2+) | 8.4 | 7.9 | 8.1 | 17 | 11.6 | 8.1 | 19 | 14 | 11.5 | 15 | 15 | 15 |
| y4 (1+) | 31.3 | 26.8 | 13.6 | 36.5 | 35.3 | 19.8 | 48.1 | 44.3 | 24 | 15 | 15 | 15 |
| y5 (1+) | 8.6 | 14.3 | 5.2 | 21.4 | 13.8 | 8.5 | 23.1 | 19.9 | 10 | 15 | 15 | 15 |
| y6 (1+) | 16.7 | 14 | 7.4 | 14.9 | 18 | 11.9 | 22.4 | 22.8 | 14 | 15 | 15 | 15 |
| y7 (1+) | 25.7 | 57.4 | 10.1 | 24.4 | 46.1 | 20.3 | 35.4 | 73.6 | 22.7 | 15 | 15 | 15 |
| sum | 5.5 | 5 | 5.4 | 16.5 | 10.5 | 5.1 | 17.4 | 11.6 | 7.4 | 15 | 15 | 15 |
Additional Resources and Comments
Assay Details for non-CPTAC-1100 Collapse assay details
Data source: Panorama
Official Gene Symbol
F12
Peptide Sequence
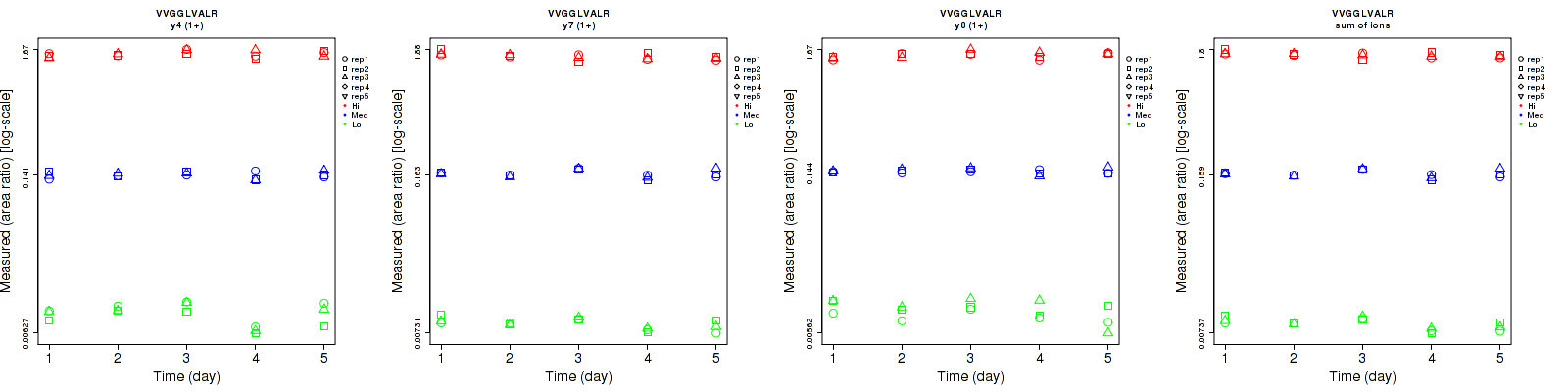
VVGGLVALR
Modification Type
unmodified
Protein - Site of Modification
N/A
Peptide - Site of Modification
N/A
Peptide Start
373
Peptide End
381
CPTAC ID
non-CPTAC-1100
Peptide Molecular Mass
882.5651
Species
Homo sapiens (Human)
Assay Type
Direct MRM
Matrix
plasma
Submitting Laboratory
UVic-Genome BC Proteomics Centre
Submitting Lab PI
Christoph Borchers
Assay Parameters Collapse assay parameters
Data source: Panorama
Instrument
6490 Triple Quad (Agilent)
Internal Standard
synthetic peptide
Peptide Standard Purity
>95%
Peptide Standard Label Type
13C and 15N at C-terminus R
LC
1290 LC (Agilent)
Column Packing
Zorbax Eclipse Plus C18, 1.8 µm
Column Dimensions
2.1 x 150 mm
Flow Rate
400 µL/min
Chromatograms
Data source: Panorama
Response Curves
Data source: Panorama
Retrieving Data

Repeatability
Data source: Panorama
| | Average intra-assay CV(within day CV) | Average inter-assay CV(between day CV) | Total CV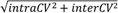 | n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y4 (1+) | 10.9 | 6.2 | 4.9 | 18.5 | 6.6 | 5.9 | 21.5 | 9.1 | 7.7 | 15 | 15 | 15 |
| y7 (1+) | 5.8 | 3.6 | 5.1 | 10.8 | 7.6 | 6 | 12.3 | 8.4 | 7.9 | 15 | 15 | 15 |
| y8 (1+) | 17.4 | 4.5 | 4.3 | 15.3 | 4.4 | 5.7 | 23.2 | 6.3 | 7.1 | 15 | 15 | 15 |
| sum | 5 | 3.4 | 4.3 | 10.8 | 6.9 | 4.7 | 11.9 | 7.7 | 6.4 | 15 | 15 | 15 |
| n= | | | | | | | | | |
| ---------------------------------------- | -------------------------------------- | --------------------------------------------------------------------------------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- | ------- | ------- | -------- |
| Fragment ion / Transition | Low | Med | High | Low | Med | High | Low | Med | High | Low | Med | High |
| y4 (1+) | 10.9 | 6.2 | 4.9 | 18.5 | 6.6 | 5.9 | 21.5 | 9.1 | 7.7 | 15 | 15 | 15 |
| y7 (1+) | 5.8 | 3.6 | 5.1 | 10.8 | 7.6 | 6 | 12.3 | 8.4 | 7.9 | 15 | 15 | 15 |
| y8 (1+) | 17.4 | 4.5 | 4.3 | 15.3 | 4.4 | 5.7 | 23.2 | 6.3 | 7.1 | 15 | 15 | 15 |
| sum | 5 | 3.4 | 4.3 | 10.8 | 6.9 | 4.7 | 11.9 | 7.7 | 6.4 | 15 | 15 | 15 |