cells – NIH Director's Blog (original) (raw)
Millions of Single-Cell Analyses Yield Most Comprehensive Human Cell Atlas Yet
Posted on May 24th, 2022 by Lawrence Tabak, D.D.S., Ph.D.

There are 37 trillion or so cells in our bodies that work together to give us life. But it may surprise you that we still haven’t put a good number on how many distinct cell types there are within those trillions of cells.
That’s why in 2016, a team of researchers from around the globe launched a historic project called the Human Cell Atlas (HCA) consortium to identify and define the hundreds of presumed distinct cell types in our bodies. Knowing where each cell type resides in the body, and which genes each one turns on or off to create its own unique molecular identity, will revolutionize our studies of human biology and medicine across the board.
Since its launch, the HCA has progressed rapidly. In fact, it has already reached an important milestone with the recent publication in the journal Science of four studies that, together, comprise the first multi-tissue drafts of the human cell atlas. This draft, based on analyses of millions of cells, defines more than 500 different cell types in more than 30 human tissues. A second draft, with even finer definition, is already in the works.
Making the HCA possible are recent technological advances in RNA sequencing. RNA sequencing is a topic that’s been mentioned frequently on this blog in a range of research areas, from neuroscience to skin rashes. Researchers use it to detect and analyze all the messenger RNA (mRNA) molecules in a biological sample, in this case individual human cells from a wide range of tissues, organs, and individuals who voluntarily donated their tissues.
By quantifying these RNA messages, researchers can capture the thousands of genes that any given cell actively expresses at any one time. These precise gene expression profiles can be used to catalogue cells from throughout the body and understand the important similarities and differences among them.
In one of the published studies, funded in part by the NIH, a team co-led by Aviv Regev, a founding co-chair of the consortium at the Broad Institute of MIT and Harvard, Cambridge, MA, established a framework for multi-tissue human cell atlases [1]. (Regev is now on leave from the Broad Institute and MIT and has recently moved to Genentech Research and Early Development, South San Francisco, CA.)
Among its many advances, Regev’s team optimized single-cell RNA sequencing for use on cell nuclei isolated from frozen tissue. This technological advance paved the way for single-cell analyses of the vast numbers of samples that are stored in research collections and freezers all around the world.
Using their new pipeline, Regev and team built an atlas including more than 200,000 single-cell RNA sequence profiles from eight tissue types collected from 16 individuals. These samples were archived earlier by NIH’s Genotype-Tissue Expression (GTEx) project. The team’s data revealed unexpected differences among cell types but surprising similarities, too.
For example, they found that genetic profiles seen in muscle cells were also present in connective tissue cells in the lungs. Using novel machine learning approaches to help make sense of their data, they’ve linked the cells in their atlases with thousands of genetic diseases and traits to identify cell types and genetic profiles that may contribute to a wide range of human conditions.
By cross-referencing 6,000 genes previously implicated in causing specific genetic disorders with their single-cell genetic profiles, they identified new cell types that may play unexpected roles. For instance, they found some non-muscle cells that may play a role in muscular dystrophy, a group of conditions in which muscles progressively weaken. More research will be needed to make sense of these fascinating, but vital, discoveries.
The team also compared genes that are more active in specific cell types to genes with previously identified links to more complex conditions. Again, their data surprised them. They identified new cell types that may play a role in conditions such as heart disease and inflammatory bowel disease.
Two of the other papers, one of which was funded in part by NIH, explored the immune system, especially the similarities and differences among immune cells that reside in specific tissues, such as scavenging macrophages [2,3] This is a critical area of study. Most of our understanding of the immune system comes from immune cells that circulate in the bloodstream, not these resident macrophages and other immune cells.
These immune cell atlases, which are still first drafts, already provide an invaluable resource toward designing new treatments to bolster immune responses, such as vaccines and anti-cancer treatments. They also may have implications for understanding what goes wrong in various autoimmune conditions.
Scientists have been working for more than 150 years to characterize the trillions of cells in our bodies. Thanks to this timely effort and its advances in describing and cataloguing cell types, we now have a much better foundation for understanding these fundamental units of the human body.
But the latest data are just the tip of the iceberg, with vast flows of biological information from throughout the human body surely to be released in the years ahead. And while consortium members continue making history, their hard work to date is freely available to the scientific community to explore critical biological questions with far-reaching implications for human health and disease.
References:
[1] Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Eraslan G, Drokhlyansky E, Anand S, Fiskin E, Subramanian A, Segrè AV, Aguet F, Rozenblatt-Rosen O, Ardlie KG, Regev A, et al. Science. 2022 May 13;376(6594):eabl4290.
[2] Cross-tissue immune cell analysis reveals tissue-specific features in humans. Domínguez Conde C, Xu C, Jarvis LB, Rainbow DB, Farber DL, Saeb-Parsy K, Jones JL,Teichmann SA, et al. Science. 2022 May 13;376(6594):eabl5197.
[3] Mapping the developing human immune system across organs. Suo C, Dann E, Goh I, Jardine L, Marioni JC, Clatworthy MR, Haniffa M, Teichmann SA, et al. Science. 2022 May 12:eabo0510.
Links:
Ribonucleic acid (RNA) (National Human Genome Research Institute/NIH)
Studying Cells (National Institute of General Medical Sciences/NIH)
Regev Lab (Broad Institute of MIT and Harvard, Cambridge, MA)
NIH Support: Common Fund; National Cancer Institute; National Human Genome Research Institute; National Heart, Lung, and Blood Institute; National Institute on Drug Abuse; National Institute of Mental Health; National Institute on Aging; National Institute of Allergy and Infectious Diseases; National Institute of Neurological Disorders and Stroke; National Eye Institute
Tags: autoimmune disorders, cell biology, cell reference maps, cells, connective tissue, genotype-tissue expression, GTEx, HCA, heart disease, Human Cell Atlas, immunology, inflammatory bowel disease, lungs, machine learning, macrophage, messenger RNA, mRNA, muscle, muscular dystrophy, RNA, RNA sequencing, single-cell analysis, single-cell RNA sequencing
New Microscope Technique Provides Real-Time 3D Views
Posted on September 30th, 2021 by Dr. Francis Collins
Most of the “cool” videos shared on my blog are borne of countless hours behind a microscope. Researchers must move a biological sample through a microscope’s focus, slowly acquiring hundreds of high-res 2D snapshots, one painstaking snap at a time. Afterwards, sophisticated computer software takes this ordered “stack” of images, calculates how the object would look from different perspectives, and later displays them as 3D views of life that can be streamed as short videos.
But this video is different. It was created by what’s called a multi-angle projection imaging system. This new optical device requires just a few camera snapshots and two mirrors to image a biological sample from multiple angles at once. Because the device eliminates the time-consuming process of acquiring individual image slices, it’s up to 100 times faster than current technologies and doesn’t require computer software to construct the movie. The kicker is that the video can be displayed in real time, which isn’t possible with existing image-stacking methods.
The video here shows two human melanoma cells, rotating several times between overhead and side views. You can see large amounts of the protein PI3K (brighter orange hues indicate higher concentrations), which helps some cancer cells divide and move around. Near the cell’s perimeter are small, dynamic surface protrusions. PI3K in these “blebs” is thought to help tumor cells navigate and survive in foreign tissues as the tumor spreads to other organs, a process known as metastasis.
The new multi-angle projection imaging system optical device was described in a paper published recently in the journal Nature Methods [1]. It was created by Reto Fiolka and Kevin Dean at the University of Texas Southwestern Medical Center, Dallas.
Like most technology, this device is complicated. Rather than the microscope and camera doing all the work, as is customary, two mirrors within the microscope play a starring role. During a camera exposure, these mirrors rotate ever so slightly and warp the acquired image in such a way that successive, unique perspectives of the sample magically come into view. By changing the amount of warp, the sample appears to rotate in real-time. As such, each view shown in the video requires only one camera snapshot, instead of acquiring hundreds of slices in a conventional scheme.
The concept traces to computer science and an algorithm called the shear warp transform method. It’s used to observe 3D objects from different perspectives on a 2D computer monitor. Fiolka, Dean, and team found they could implement a similar algorithm optically for use with a microscope. What’s more, their multi-angle projection imaging system is easy-to-use, inexpensive, and can be converted for use on any camera-based microscope.
The researchers have used the device to view samples spanning a range of sizes: from mitochondria and other tiny organelles inside cells to the beating heart of a young zebrafish. And, as the video shows, it has been applied to study cancer and other human diseases.
In a neat, but also scientifically valuable twist, the new optical method can generate a virtual reality view of a sample. Any microscope user wearing the appropriately colored 3D glasses immediately sees the objects.
While virtual reality viewing of cellular life might sound like a gimmick, Fiolka and Dean believe that it will help researchers use their current microscopes to see any sample in 3D—offering the chance to find rare and potentially important biological events much faster than is possible with even the most advanced microscopes today.
Fiolka, Dean, and team are still just getting started. Because the method analyzes tissue very quickly within a single image frame, they say it will enable scientists to observe the fastest events in biology, such as the movement of calcium throughout a neuron—or even a whole bundle of neurons at once. For neuroscientists trying to understand the brain, that’s a movie they will really want to see.
Reference:
[1] Real-time multi-angle projection imaging of biological dynamics. Chang BJ, Manton JD, Sapoznik E, Pohlkamp T, Terrones TS, Welf ES, Murali VS, Roudot P, Hake K, Whitehead L, York AG, Dean KM, Fiolka R. Nat Methods. 2021 Jul;18(7):829-834.
Links:
Metastatic Cancer: When Cancer Spreads (National Cancer Institute)
Fiolka Lab (University of Texas Southwestern Medical Center, Dallas)
Dean Lab (University of Texas Southwestern)
Microscopy Innovation Lab (University of Texas Southwestern)
NIH Support: National Cancer Institute; National Institute of General Medical Sciences
Posted In: Cool Videos
Tags: 3D imaging, basic research, brain, cancer, cancer biology, cancer metastasis, cell biology, cells, imaging, melanoma, microscopy, multi-angle projection imaging system optical device, neurons, PI3K, shear warp transform method
Defining Neurons in Technicolor
Posted on November 7th, 2019 by Dr. Francis Collins

Credit: Allen Institute for Brain Science, Seattle
Can you identify a familiar pattern in this image’s square grid? Yes, it’s the outline of the periodic table! But instead of organizing chemical elements, this periodic table sorts 46 different types of neurons present in the visual cortex of a mouse brain.
Scientists, led by Hongkui Zeng at the Allen Institute for Brain Science, Seattle, constructed this periodic table by assigning colors to their neuronal discoveries based upon their main cell functions [1]. Cells in pinks, violets, reds, and oranges have inhibitory electrical activity, while those in greens and blues have excitatory electrical activity.
For any given cell, the darker colors indicate dendrites, which receive signals from other neurons. The lighter colors indicate axons, which transmit signals. Examples of electrical properties—the number and intensity of their “spikes”—appear along the edges of the table near the bottom.
To create this visually arresting image, Zeng’s NIH-supported team injected dye-containing probes into neurons. The probes are engineered to carry genes that make certain types of neurons glow bright colors under the microscope.
This allowed the researchers to examine a tiny slice of brain tissue and view each colored neuron’s shape, as well as measure its electrical response. They followed up with computational tools to combine these two characteristics and classify cell types based on their shape and electrical activity. Zeng’s team could then sort the cells into clusters using a computer algorithm to avoid potential human bias from visually interpreting the data.
Why compile such a detailed atlas of neuronal subtypes? Although scientists have been surveying cells since the invention of the microscope centuries ago, there is still no consensus on what a “cell type” is. Large, rich datasets like this atlas contain massive amounts of information to characterize individual cells well beyond their appearance under a microscope, helping to explain factors that make cells similar or dissimilar. Those differences may not be apparent to the naked eye.
Just last year, Allen Institute researchers conducted similar work by categorizing nearly 24,000 cells from the brain’s visual and motor cortex into different types based upon their gene activity [2]. The latest research lines up well with the cell subclasses and types categorized in the previous gene-activity work. As a result, the scientists have more evidence that each of the 46 cell types is actually distinct from the others and likely drives a particular function within the visual cortex.
Publicly available resources, like this database of cell types, fuel much more discovery. Scientists all over the world can look at this table (and soon, more atlases from other parts of the brain) to see where a cell type fits into a region of interest and how it might behave in a range of brain conditions.
References:
[1] Classification of electrophysiological and morphological neuron types in the mouse visual cortex. N Gouwens NW, et al. Neurosci. 2019 Jul;22(7):1182-1195.
[2] Shared and distinct transcriptomic cell types across neocortical areas. Tasic B, et al. Nature. 2018 Nov;563(7729):72-78.
Links:
Brain Basics: The Life and Death of a Neuron (National Institute of Neurological Disorders and Stroke/NIH)
Cell Types: Overview of the Data (Allen Brain Atlas/Allen Institute for Brain Science, Seattle)
Hongkui Zeng (Allen Institute)
NIH Support: National Institute of Mental Health; Eunice Kennedy Shriver National Institute of Child Health & Human Development
Posted In: Snapshots of Life
Tags: Allen Mouse Brain Atlas, axons, brain, brain atlas, cell biology, cells, dendrites, electrophysiology, motor cortex, mouse, neurology, neuronal subclasses, neuronal subtypes, neurons, neuroscience, periodic table, visual cortex
Teaching Computers to “See” the Invisible in Living Cells
Posted on April 24th, 2018 by Dr. Francis Collins
Caption: While analyzing brain cells, a computer program “thinks” about which cellular structure to identify.
Credit: Steven Finkbeiner, University of California, San Francisco and the Gladstone Institutes
For centuries, scientists have trained themselves to look through microscopes and carefully study their structural and molecular features. But those long hours bent over a microscope poring over microscopic images could be less necessary in the years ahead. The job of analyzing cellular features could one day belong to specially trained computers.
In a new study published in the journal Cell, researchers trained computers by feeding them paired sets of fluorescently labeled and unlabeled images of brain tissue millions of times in a row [1]. This allowed the computers to discern patterns in the images, form rules, and apply them to viewing future images. Using this so-called deep learning approach, the researchers demonstrated that the computers not only learned to recognize individual cells, they also developed an almost superhuman ability to identify the cell type and whether a cell was alive or dead. Even more remarkable, the trained computers made all those calls without any need for harsh chemical labels, including fluorescent dyes or stains, which researchers normally require to study cells. In other words, the computers learned to “see” the invisible!
Posted In: News
Tags: Alzheimer’s disease, brain, Brain Bot, cell biology, cells, computer learning, computers, deep learning, Google, machine learning, microscopy, neurology, neuroscience, Parkinson's disease, schizophrenia
Cool Videos: Making Multicolored Waves in Cell Biology
Posted on March 16th, 2017 by Dr. Francis Collins
Bacteria are single-cell organisms that reproduce by dividing in half. Proteins within these cells organize themselves in a number of fascinating ways during this process, including a recently discovered mechanism that makes the mesmerizing pattern of waves, or oscillations, you see in this video. Produced when the protein MinE chases the protein MinD from one end of the cell to the other, such oscillations are thought to center the cell’s division machinery so that its two new “daughter cells” will be the same size.
To study these dynamic patterns in greater detail, Anthony Vecchiarelli purified MinD and MinE proteins from the bacterium Esc herichia coli . Vecchiarelli, who at the time was a postdoc in Kiyoshi Mizuuchi’s intramural lab at NIH’s National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), labeled the proteins with fluorescent markers and placed them on a synthetic membrane, where their movements were then visualized by total internal reflection fluorescence microscopy. The proteins self-organized and generated dynamic spirals of waves: MinD (blue, left); MinE (red, right); and both MinD and MinE (purple, center) [1].
Tags: art, bacteria, cell biology, cell division, cell migration, cell-free biology, cell-free systems, cells, chemotaxis, E. coli, endocytosis, Escherichia coli, FASEB Bioart 2016, MinD, MinE, mitosis, oscillation, protein pattern self-organization, protein self-organization, reaction-diffusion model, Science, spatial organization, subcellular organization, total internal reflection fluorescence microscopy, Turing patterns
Snapshots of Life: A Flare for the Dramatic
Posted on September 29th, 2016 by Dr. Francis Collins

Credit: Valentin Romanov, University of Utah, Salt Lake City
Oil and water may not mix, but under the right conditions—like those in the photo above—it can sure produce some interesting science that resembles art. You’re looking at a water droplet suspended in an emulsion of olive oil (black and purple) and lipids, molecules that serve as the building blocks of cell membranes. Each lipid has been tagged with a red fluorescent marker, and what look like red and yellow flames are the markers reacting to a beam of UV light. Their glow shows the lipids sticking to the surface of the water droplet, which will soon engulf the droplet to form a single lipid bilayer, which can later be transformed into a lipid bilayer that closely resembles a cell membrane. Scientists use these bubbles, called liposomes, as artificial cells for a variety of research purposes.
In this case, the purpose is structural biology studies. Valentin Romanov, the graduate student at the University of Utah, Salt Lake City, who snapped the image, creates liposomes to study proteins that help cells multiply. By encapsulating and letting the proteins interact with lipids in the artificial cell membrane, Romanov and his colleagues in the NIH-supported labs of Bruce Gale at the University of Utah and Adam Frost at the University of California, San Francisco, can freeze and capture their changing 3D structures at various points in the cell division process with high-resolution imaging techniques. These snapshots will help the researchers to understand in finer detail how the proteins work and perhaps to design drugs to manipulate their functions.
Tags: BioArt, cell biology, cell membrane, cells, imaging, lipids, liposomes, membrane biology, microfluidics, Science as Art, structural biology, University of Utah, water droplet
Snapshots of Life: Fish Awash in Color
Posted on March 31st, 2016 by Dr. Francis Collins
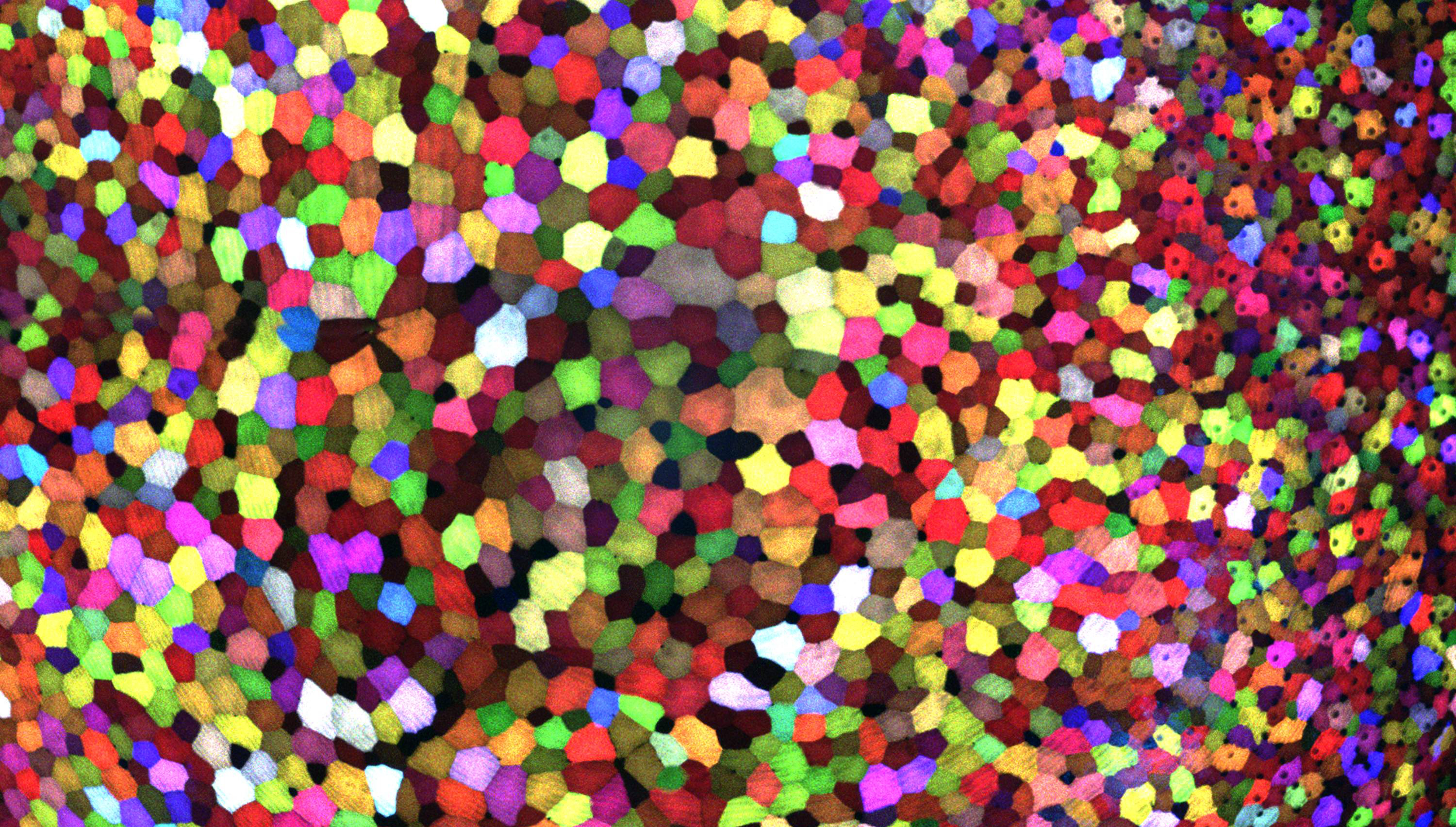

Credit: Chen-Hui Chen, Duke University
If this image makes you think of a modern art, you’re not alone. But what you’re actually seeing are hundreds of live cells from a tiny bit (0.0003348 square inches) of skin on the tail fin of a genetically engineered adult zebrafish. Zebrafish are normally found in tropical freshwater and are a favorite research model to study vertebrate development and tissue regeneration. The cells have been labeled with a cool, new fluorescent imaging tool called Skinbow. It uniquely color codes cells by getting them to express genes encoding red, green, and blue fluorescent proteins at levels that are randomly determined. The different ratios of these colorful proteins mix to give each cell a distinctive hue when imaged under a microscope. Here, you can see more than 70 detectable Skinbow colors that make individual cells as visually distinct from one another as jellybeans in a jar.
Skinbow is the creation of NIH-supported scientists Chen-Hui Chen and Kenneth Poss at Duke University, Durham, NC, with imaging computational help from collaborators Stefano Di Talia and Alberto Puliafito. As reported recently in the journal Developmental Cell [1], Skinbow’s distinctive spectrum of color occurs primarily in the outermost part of the skin in a layer of non-dividing epithelial cells. Using Skinbow, Poss and colleagues tracked these epithelial cells, individually and as a group, over their entire 2 to 3 week lifespans in the zebrafish. This gave them an unprecedented opportunity to track the cellular dynamics of wound healing or the regeneration of lost tissue over time. While Skinbow only works in zebrafish for now, in theory, it could be adapted to mice and maybe even humans to study skin and possibly other organs.
Posted In: Science
Tags: Brainbow, cell biology, cells, dermatology, fish, gene expression, genes, model organism, skin, Skinbow, tissue regeneration, wound healing, zebra fish, zebrafish
Snapshots of Life: Cell Skeleton on the Move
Posted on April 2nd, 2015 by Dr. Francis Collins
Credit: Torsten Wittmann, University of California, San Francisco
Cells are constantly on the move. They shift, grow, and migrate to new locations—for example, to heal a wound or to intercept an infectious agent as part of an immune response. But how do cells actually move?
In this image, Torsten Wittmann, an NIH-funded cell biologist at the University of California, San Francisco, reveals the usually-invisible cytoskeleton of a normal human skin cell that lends the cell its mobility. The cytoskeleton is made from protein structures called microtubules—the wispy threads surrounding the purple DNA-containing nucleus—and filaments of a protein called actin, seen here as the fine blue meshwork in the cell periphery. Both actin and microtubules are critical for growth and movement.
Cool Videos: Metabolomics
Posted on August 14th, 2014 by Dr. Francis Collins
Today’s feature in my Cool Video series is a scientific film noir from the University of Florida in Gainesville. Channeling Humphrey Bogart’s hard-boiled approach to detective work, the protagonist of this video is tracking down metabolites—molecules involved in biological mysteries with more twists and turns than “The Maltese Falcon.”
If you’d like a few more details before or after watching the video, here’s how the scientists themselves describe their project: “Inside our cells, chemical heroes, victims, and villains leave behind clues about our health. Meet Dr. Art Edison, one of many metabolomics PIs who are on the case. Their quest? To tail and fingerprint small molecules, called metabolites, which result from the chemical processes that fuel and sustain life. Metabolites can shed light on the state of health, nutrition, or disease in a living thing—whether human, animal, or plant. Funded by National Institutes of Health grant U24DK097209, the University of Florida Southeast Center for Integrated Metabolomics is sleuthing through these cellular secrets.”
Snapshots of Life: Wild Outcome from Knocking Out Mobility Proteins
Posted on July 31st, 2014 by Dr. Francis Collins
Credit: Praveen Suraneni and Rong Li, Stowers Institute for Medical Research
When biologists disabled proteins critical for cell movement, the result was dramatic. The membrane, normally a smooth surface enveloping the cell, erupted in spiky projections. This image, which is part of the Life: Magnified exhibit, resembles a supernova. Although it looks like it exploded, the cell pictured is still alive.
To create the image, Rong Li and Praveen Suraneni, NIH-funded cell biologists at the Stowers Institute for Medical Research in Kansas City, Missouri, disrupted two proteins essential to movement in fibroblasts—connective tissue cells that are also important for healing wounds. The first, called ARPC3, is a protein in the Arp2/3 complex. Without it, the cell moves more slowly and randomly [1]. Inhibiting the second protein gave this cell its spiky appearance. Called myosin IIA (green in the image), it’s like the cell’s muscle, and it’s critical for movement. The blue color is DNA; the red represents a protein called F-actin.
Posted In: Science
Tags: Arp2/3, ARPC3, cancer cells, cell movement, cells, DNA, F-actin, fibroblasts, Life: Magnified, metastasis, myosin IIA, proteins, stem cells, wound healing


