Quantitative analysis of blood cells from microscopic images using convolutional neural network (original) (raw)
Abstract
Blood cell count provides relevant clinical information about different kinds of disorders. Any deviation in the number of blood cells implies the presence of infection, inflammation, edema, bleeding, and other blood-related issues. Current microscopic methods used for blood cell counting are very tedious and are highly prone to different sources of errors. Besides, these techniques do not provide full information related to blood cells like shape and size, which play important roles in the clinical investigation of serious blood-related diseases. In this paper, deep learning-based automatic classification and quantitative analysis of blood cells are proposed using the YOLOv2 model. The model was trained on 1560 images and 2703-labeled blood cells with different hyper-parameters. It was tested on 26 images containing 1454 red blood cells, 159 platelets, 3 basophils, 12 eosinophils, 24 lymphocytes, 13 monocytes, and 28 neutrophils. The network achieved detection and segmentation of blood cells with an average accuracy of 80.6% and a precision of 88.4%. Quantitative analysis of cells was done following classification, and mean accuracy of 92.96%, 91.96%, 88.736%, and 92.7% has been achieved in the measurement of area, aspect ratio, diameter, and counting of cells respectively.
Graphical abstract
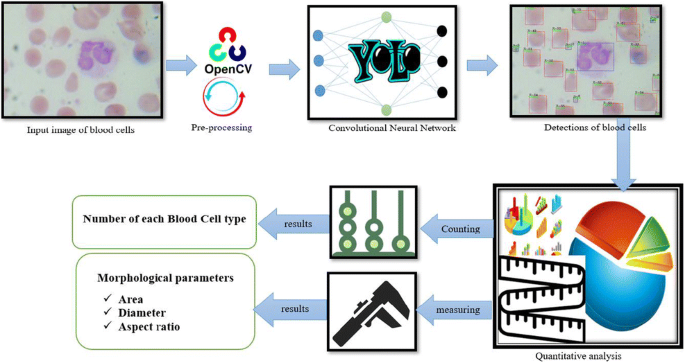
Graphical abstract where the first picture shows the input image of blood cells seen under a compound light microscope. The second image shows the tools used like OpenCV to pre-process the image. The third image shows the convolutional neural network used to train and perform object detection. The 4th image shows the output of the network in the detection of blood cells. The last images indicate post-processing applied on the output image such as counting of each blood cells using the class label of each detection and quantification of morphological parameters like area, aspect ratio, and diameter of blood cells so that the final result provides the number of each blood cell types (seven) and morphological information providing valuable clinical information.
Access this article
Subscribe and save
- Starting from 10 chapters or articles per month
- Access and download chapters and articles from more than 300k books and 2,500 journals
- Cancel anytime View plans
Buy Now
Price excludes VAT (USA)
Tax calculation will be finalised during checkout.
Instant access to the full article PDF.
Similar content being viewed by others
Data availability
The image data used to support the finding of this study are available from the corresponding author upon request.
References
- Khan A (2017) An accurate and cost effective approach to blood cell count, no. January, pp. 17–24
- Moravapalle UP, Deshpande A, Kapoor A, Ramjee R, Ravi P (2017) Blood count on a smartphone microscope. Work Mob Comput Syst Appl:37–42
- Khan S, Khan A, Saleh Khattak F, Naseem A (2012) An accurate and cost effective approach to blood cell count. Int J Comput Appl 50(1):18–24
Google Scholar - Yi F, Moon I, Javidi B (2017) Automated red blood cells extraction from holographic images using fully convolutional neural networks. Biomed Opt Express 8(10):4466
Article Google Scholar - Elmazi D, Cuka M, Ikeda M, Matsuo K, Barolli L (2019) Application of fuzzy logic for selection of actor nodes in wsans —implementation of two fuzzy-based systems and a testbed. Sensors (Switzerland) 19(24)
- Bayrakdar ME (2019) Priority based health data monitoring with IEEE 802.11af technology in wireless medical sensor networks. Med Biol Eng Comput 57(12):2757–2769
Article Google Scholar - Hao Y, Peng L, Lu H, Hassan MM, Alamri A (2017) Energy harvesting based body area networks for smart health. Sensors (Switzerland) 17(7):1–10
Article Google Scholar - Thejashwini M, Padma MC (2015) Counting of RBC’s and WBC’s using image processing technique. Int J Recent Innov Trends Comput Commun 3(5):2948–2953
Google Scholar - Azwai S, Abdouslam O (2007) Morphological characteristics of blood cells in clinically normal adult llamas (Lama glama). Vet 77(1):69–79
Google Scholar - Othman MZ, Mohammed TS, Ali AB (2017) Neural network classification of white blood cell using microscopic images, IJACSA. Int J Adv Comput Sci Appl 8(5)
- Shahin AI, Guo Y, Amin KM, Sharawi AA (2017) White blood cells identification system based on convolutional deep neural learning networks. Comput Methods Prog Biomed
- Paxton S, Peckham M, Adele K, Paxton S, Adele K, and Peckham M (2003) The leeds histology guide
- Ramesh N, Dangott B, Salama ME, Tasdizen T (2012) Isolation and two-step classification of normal white blood cells in peripheral blood smears. J Pathol Inform 3(1):13
Article Google Scholar - Veluchamy M, Perumal K, Ponuchamy T (2012) Feature extraction and classification of blood cells using artificial neural network. Am J Appl Sci
- Madabhushi A, Lee G (2016) Image analysis and machine learning in digital pathology: challenges and opportunities. Med Image Anal 33:170–175
Article Google Scholar - Aswathy MA and Jagannath M (2016) Detection of breast cancer on digital histopathology images: present status and future possibilities, Informatics Med. Unlocked, vol. 8, no. November 2016, pp. 74–79
- Parthasarathy D (2017) A brief history of CNNs in image segmentation: from R-CNN to mask R-CNN, Medium. [Online]. Available: https://blog.athelas.com/a-brief-history-of-cnns-in-image-segmentation-from-r-cnn-to-mask-r-cnn-34ea83205de4. [Accessed: 05-Dec-2018]
- Delforouzi A, Pamarthi B, Grzegorzek M (2018) Training-based methods for comparison of object detection methods for visual object tracking. Sensors (Basel) 18(11)
- Hui J (2018) Object detection: speed and accuracy comparison (Faster R-CNN, R-FCN, SSD, FPN, RetinaNet and…. [Online]. Available: https://medium.com/@jonathan_hui/object-detection-speed-and-accuracy-comparison-faster-r-cnn-r-fcn-ssd-and-yolo-5425656ae359. [Accessed: 11-Dec-2018]
- Jordan Jeremy (2018) An overview of object detection: one-stage methods. [Online]. Available: https://www.jeremyjordan.me/object-detection-one-stage/. [Accessed: 19-Aug-2019]
- Redmon J and Farhadi A (2018) YOLO: real-time object detection, 2018. [Online]. Available: https://pjreddie.com/darknet/yolo/. [Accessed: 05-Dec-2018]
- Gautam A and Bhadauria H (2014) Classification of white blood cells based on morphological features, in 2014 International Conference on Advances in Computing, Communications and Informatics (ICACCI), pp. 2363–2368
- Rejula MA, Kumar MKJ (2016) An efficient approach of segmentation on white blood cells using morphological operators and 2D-discrete wavelet transform with anfis classification. Int J Emerg Res Manag Technol 5(58):2278–9359
Google Scholar - Segui S, Pujol O, and Vitria J (2015) Learning to count with deep object features. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. Work. vol. 2015-Octob, pp. 90–96
- Cohen JP, Boucher G, Glastonbury CA, Lo HZ, Bengio Y (2017) Count-ception: counting by fully convolutional redundant counting. Comput Vis Pattern Recog
- Xie Y, Xing F, Kong X, Su H, and Yang L Beyond classification: structured regression for robust cell detection using convolutional neural network, in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. MICCAI 2015. Lecture Notes in Computer Science, vol 9351, 2015, pp. 358–365
- Xie W, Noble JA, Zisserman A (2018) Microscopy cell counting and detection with fully convolutional regression networks. Comput Methods Biomech Biomed Eng Imaging Vis 6(3):283–292
Article Google Scholar - Hernández CX, Sultan MM, Pande VS (2018) Using deep learning for segmentation and counting within microscopy data. Comput Vis Pattern Recog
- Cai F and Verbeek FJ (2015) Dam-based rolling ball with fuzzy-rough constraints, a new background subtraction algorithm for image analysis in microscopy, 5th Int. Conf. Image Process. Theory, Tools Appl. 2015, IPTA 2015, pp. 298–303
- Herrera-Navarro AM, Jiménez Hernández H, Peregrina-Barreto H, Manríquez-Guerrero F, and Terol-Villalobos IR (2013) A new measure of circularity based on distribution of the radius, vol. 17, no. 4, pp. 515–526
- Redmon J, Farhadi A (2016) YOLO9000: better, faster, stronger. Comput Vis Pattern Recog
- Palmerini T, Mehran R, Dangas G, Nikolsky E, Witzenbichler B, Guagliumi G, Dudek D, Genereux P, Caixeta A, Rabbani LR, Weisz G, Parise H, Fahy M, Xu K, Brodie B, Lansky A, Stone GW (2011) Impact of leukocyte count on mortality and bleeding in patients with myocardial infarction undergoing primary percutaneous coronary interventions: analysis from the harmonizing outcome with revascularization and stent in acute myocardial infarction trial. Circulation 123(24):2829–2837
Article Google Scholar - Alomari YM, Sheikh Abdullah SNH, Zaharatul Azma R, Omar K (2014) Automatic detection and quantification of WBCs and RBCs using iterative structured circle detection algorithm. Comput Math Methods Med 2014:979302
Article Google Scholar - Nah EH, Kim S, Cho S, Cho HI (2018) Complete blood count reference intervals and patterns of changes across pediatric, adult, and geriatric ages in Korea. Ann Lab Med 38(6):503–511
Article Google Scholar - Van Der Meer W, Van Gelder W, De Keijzer R, Willems H (2007) The divergent morphological classification of variant lymphocytes in blood smears. J Clin Pathol 60:838–839
Article Google Scholar
Author information
Authors and Affiliations
- Jimma Institute of Technology, School of Biomedical Engineering, Jimma University, P.O. Box 378, Jimma, Ethiopia
Abel Worku Tessema, Mohammed Aliy Mohammed, Gizeaddis Lamesgin Simegn & Timothy Chung Kwa - Department of Biomedical Engineering, School of Life Science, Ulsan National Institute of Science and Technology, Ulsan, Republic of Korea
Abel Worku Tessema - UCT Medical Imaging Unit, Division of Biomedical Engineering, Faculty of Health Sciences, University of Cape Town, Cape Town, South Africa
Gizeaddis Lamesgin Simegn
Authors
- Abel Worku Tessema
- Mohammed Aliy Mohammed
- Gizeaddis Lamesgin Simegn
- Timothy Chung Kwa
Corresponding author
Correspondence toAbel Worku Tessema.
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tessema, A.W., Mohammed, M.A., Simegn, G.L. et al. Quantitative analysis of blood cells from microscopic images using convolutional neural network.Med Biol Eng Comput 59, 143–152 (2021). https://doi.org/10.1007/s11517-020-02291-w
- Received: 21 April 2020
- Accepted: 15 November 2020
- Published: 01 January 2021
- Version of record: 01 January 2021
- Issue date: January 2021
- DOI: https://doi.org/10.1007/s11517-020-02291-w