Microarrays: lost in a storm of data? (original) (raw)
- Opinion
- Published: 01 June 2001
Nature Reviews Neuroscience volume 2, pages 441–443 (2001)Cite this article
- 210 Accesses
- 28 Citations
- Metrics details
Abstract
Microarray expression profiling is instrumental to our understanding of the function of the genome. Resolution of functionally relevant expression patterns will require the analysis of large data sets compiled from multiple investigators. For this and other reasons, I argue that it is crucial for array data to be publicly shared in a format as close to the 'raw data' as possible. Issues such as protection of intellectual property, ensuring quality of the data, and the format and timing for sharing array data are also discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Linearity in microarray results.

The alternative text for this image may have been generated using AI.
Figure 2: Nonlinearity in microarray results.
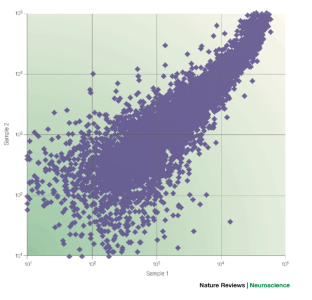
The alternative text for this image may have been generated using AI.
Figure 3: Position effects in microarray analysis.

The alternative text for this image may have been generated using AI.
Similar content being viewed by others
A comparison of genotyping arrays
Article Open access 18 June 2021


References
- Iyer, V. R. et al. The transcriptional program in the response of human fibroblasts to serum. Science 283, 83–87 (1999).
Article CAS Google Scholar - Hughes, T. R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Article CAS Google Scholar - Thibault, C. et al. Expression profiling of neural cells reveals specific patterns of ethanol-responsive gene expression. Mol. Pharmacol. 58, 1593–1600 (2000).
Article CAS Google Scholar - Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Article CAS Google Scholar - Lewohl, J. M. et al. Gene expression in human alcoholism: microarray analysis of frontal cortex. Alcohol Clin. Exp. Res. 24, 1873–1882 (2000).
Article CAS Google Scholar - Perou, C. M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Article CAS Google Scholar - Raychaudhuri, S., Sutphin, P. D., Chang, J. T. & Altman, R. B. Basic microarray analysis: grouping and feature reduction. Trends Biotechnol. 19, 189–193 (2001).
Article CAS Google Scholar - Brown, M. P. et al. Knowledge-based analysis of microarray gene expression data by using support vector machines. Proc. Natl Acad. Sci. USA 97, 262–267 (2000).
Article CAS Google Scholar - Dudoit, S., Yang, Y. H., Callow, M. J. & Speed, T. J. Statistical Methods for Identifying Differentially Expressed Genes in Replicated cDNA Microarray Experiments. Technical Report 578, Stanford Univ. School of Medicine (2000).
Google Scholar - Li, C. & Wong, W. H. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Natl Acad. Sci. USA 98, 31–36 (2001).
Article CAS Google Scholar - Yue, H. et al. An evaluation of the performance of cDNA microarrays for detecting changes in global mRNA expression. Nucleic Acids Res. 29, E41 (2001).
Article CAS Google Scholar - Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl Acad. Sci. USA 98, 5116–5121 (2001).
Article CAS Google Scholar - Lee, M. L., Kuo, F. C., Whitmore, G. A. & Sklar, J. Importance of replication in microarray gene expression studies: statistical methods and evidence from repetitive cDNA hybridizations. Proc. Natl Acad. Sci. USA 97, 9834–9839 (2000).
Article CAS Google Scholar - Toronen, P., Kolehmainen, M., Wong, G. & Castren, E. Analysis of gene expression data using self-organizing maps. FEBS Lett. 451, 142–146 (1999).
Article CAS Google Scholar - Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Article CAS Google Scholar - Sherlock, G. Analysis of large-scale gene expression data. Curr. Opin. Immunol. 12, 201–205 (2000).
Article CAS Google Scholar - Jenssen, T. K., Laegreid, A., Komorowski, J. & Hovig, E. A literature network of human genes for high-throughput analysis of gene expression. Nature Genet. 28, 21–28 (2001).
CAS PubMed Google Scholar
Acknowledgements
I would like to thank all the members of the Miles laboratory for their enthusiasm, dedication and many helpful discussions. In particular, I thank Li Zhang for stimulating discussions regarding the analysis of microarray data. I also would like to acknowledge support from the National Institute on Alcohol Abuse and Alcoholism, the National Institute on Drug Abuse and the State of California, through a grant to the University of California for research on alcoholism and drug abuse.
Author information
Authors and Affiliations
- the Ernest Gallo Clinic and Research Center, 5858 Horton Street, Suite 200, Emeryville, 94608, California, USA
Michael F. Miles
Related links
Rights and permissions
About this article
Cite this article
Miles, M. Microarrays: lost in a storm of data?.Nat Rev Neurosci 2, 441–443 (2001). https://doi.org/10.1038/35077582
- Issue date: 01 June 2001
- DOI: https://doi.org/10.1038/35077582