Multilocus half-tetrad analysis and centromere mapping in citrus: evidence of SDR mechanism for 2n megagametophyte production and partial chiasma interference in mandarin cv ‘Fortune’ (original) (raw)
Introduction
Sexual polyploidisation has been widely exploited in several plant-breeding programmes (Ramanna and Jacobsen, 2003). Diploidy is the general rule in Citrus and related genera, with a basic chromosome number x_=9. However, sexual polyploidisation is currently a central approach used in triploid citrus-breeding programmes, aiming to develop new seedless ‘mandarin-type’ cultivars (Ollitrault et al., 2008); very large triploid progenies have been obtained from 2_x × 2_x_ crosses and several cultivars patented (Aleza et al., 2010). The recovery of triploid citrus hybrids arising from 2_n_ megagametophytes produced by diploid plants was described in the 1970s (Esen and Soost, 1971, 1973). Cytogenetic studies (Esen and Soost, 1971) showed that triploid embryos are associated with pentaploid endosperm, indicating that triploid hybrids result from the fertilisation of unreduced ovules by normal haploid pollen. According to the genotype, the frequency of duplication in the female gametes can range from below 1% to over 20%. Esen et al., 1979 proposed that 2_n_ eggs result from the abortion of the second meiotic division in the megaspore (SDR) in citrus. This hypothesis has been corroborated for Clementine (Citrus clementina Hort. ex Tan.) by molecular marker analysis (Luro et al., 2000). However, Chen et al. (2008) proposed that 2_n_ eggs of sweet orange (C. sinensis (L.) Osb.) resulted from first meiotic division restitution (FDR). The genetic configuration of 2_n_ gametes depends on the mechanism of their formation (Figure 1), and the rate of parental heterozygosity restitution is directly linked with the rate of effective chiasma between the centromere and the considered locus (Mendiburu and Peloquin, 1976; Park et al., 2007). It is, total and null until the first chiasma for FDR and SDR, respectively. It is, thus, essential to gain a better understanding of the mechanisms underlying 2_n_ gamete formation to optimise sexual polyploidy breeding schemes and to carry out trait-association studies of breeding populations. The objective of the present work was to shed light on the mechanism underlying 2_n_ gamete formation in ‘Fortune’ mandarin (C. clementina Hort. ex Tan. x C. tangerina Hort. ex Tan.) by simple sequence repeat (SSR) marker analysis. ‘Fortune’ mandarin is widely used in triploid breeding because of its fruit qualities, late maturing period and relatively high percentage of 2_n_ eggs. This diploid cultivar is highly fertile, producing an average of 18.5 seeds per fruit, including 6.5% of seeds arising from 2_n_ gametes (our unpublished data). Without the previous knowledge of centromere position, and to avoid the risk of misinterpreting data due to an insufficient or biased set of markers, we selected 35 SSR markers according to their position on the Clementine (C. clementina Hort. ex Tan.) map (Ollitrault et al., 2011). A general approach is proposed to estimate the centromere position by best-fit value between observed data and theoretical functions of heterozygosity restitution for no interference and partial interference models. For this purpose, we used the functions developed by Zhao and Speed (1998a) for ordered tetrads, based on the random spindle–centromere attachment hypothesis (Griffiths et al., 1996), and extended by the same authors to half-tetrad analysis (HTA; Zhao and Speed, 1998b). The results obtained were compared with the method proposed by Tavoletti et al. (1996) using the multilocus structure of half-tetrads. Finally, we discuss the implications of the 2_n_ gamete restitution mechanisms for citrus triploid breeding.
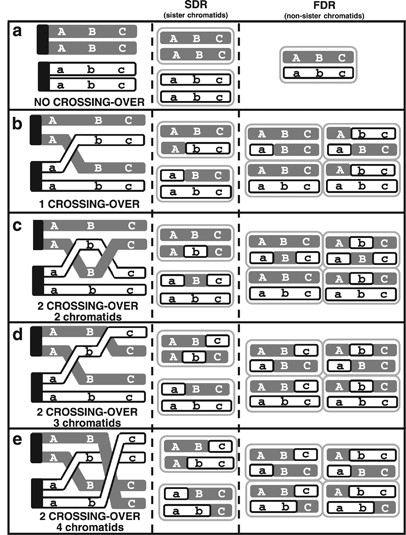
Figure 1
SDR and FDR 2n gametes resulting from: (a) no crossover; (b) one crossover; (c) two crossovers involving two chromatids; (d) two crossovers involving three chromatids and (e) two crossovers involving four chromatids.
Materials and methods
Plant material
The mechanism underlying 2_n_ gamete formation in ‘Fortune’ mandarin was investigated in 171 triploid hybrids, derived from four crosses between ‘Fortune’ mandarin (C. clementina Hort. ex Tan. × C. tangerina Hort. ex Tan.) as female diploid genitor and ‘Ellendale’ (C. reticulata Blanco × C. sinensis (L.) Osb.), ‘Common Mandarin’ (C. deliciosa Ten.), ‘Minneola’ (C. paradisi Macf. × C. tangerina Hort. ex Tan.) or ‘Murcott’ (C. reticulata Blanco × C. sinensis (L.) Osb.) as male diploid genitors. Parental accessions and hybrids were grown at the ‘Instituto Valenciano de Investigaciones Agrarias’ orchards in Moncada, Valencia, Spain. Practical details on how the triploid populations were established from diploid × diploid crosses by embryo rescue and triploid selection by flow cytometry, can be found in Aleza et al. (2010). Genomic DNA of triploid hybrids and their parents was isolated using the Plant DNAeasy kit from Qiagen Inc. (Valencia, CA, USA), following the manufacturer’s protocol.
Triploid progeny genotyping
SSR markers that proved heterozygous for ‘Fortune’ were selected to genotype each triploid progeny, depending on the polymorphism existing between ‘Fortune’ and the male genitor. Thirty-five microsatellite loci were used to analyse the triploid progenies: CAC15, TAA41 (Kijas et al., 1997), CX6F03, CX6F23 (Chen et al., 2007), mest121 (Luro et al., 2008), mest56, mest192, mest123 (Aleza et al., 2011), mest104, mest110, mest247, mest488 (François Luro, personal communication; mail to luro@corse.inra.fr for further information), mCrCIR07F11 (Kamiri et al., 2011), Ci01C07, Ci02B07, Ci08C05, mCrCIR01E02, mCrCIR01F04a, mCrCIR06A12, mCrCIR06B05, mCrCIR06B07, mCrCIR07E12 (Froelicher et al., 2008) and thirteen new designed primers (mCrCIR01C06, mCrCIR02A09, mCrCIR02D09, mCrCIR02F12, mCrCIR02G01, mCrCIR02G02, mCrCIR03B07, mCrCIR03C08, mCrCIR03G05, mCrCIR04H06, mCrCIR05A05, mCrCIR07D05, mCrCIR07D06; Table 1). The polymerase chain reactions (PCRs) were performed with wellRED oligonucleotides (Sigma-Aldrich, St Louis, MO, USA) with the following protocol: Mastercycler epgradient S (Eppendorf Scientific Inc., Westbury, NY, USA); reaction volume 15 μl: 0.8 U Taq polymerase (Fermentas, Burlington, VT, USA), reaction buffer 750 mM Tris-HCl (pH 9), 50 mM KCl, 200 mM (NH4)2SO4, 0.001% bovine serum albumin, 0.1 mM of each dNTP, 5 mM MgCl2, 3 μM of each primer, 30 ng DNA; PCR programme: 94 °C for 5 min; 40 cycles of 30 s at 94 °C, 1 min at 50–55 °C and 30 s at 72 °C; final elongation 10 min at 72 °C). Separation was carried out by Capillary Gel Electrophoresis (CEQ 8000 Genetic Analysis System; Beckman Coulter Inc., Fullerton, CA, USA). Data collection and analysis were carried out with GenomeLab GeXP (Beckman Coulter Inc.) version 10.0 software.
Table 1 New primers designed to amplify the markers used in this study
2_n_ gamete allelic structure inference from triploid hybrid genotypes
For a locus bearing totally different parental allelic configurations (A1A2 × A3A4), the genotype of the 2_n_ gamete was directly read from the triploid hybrid structure. When the male and female genitor shared one allele (A1A2 × A2A2 and A1A2 × A2A3), the inference of the 2_n_ female gamete structure was carried out from the measured allele dosage by the microsatellite DNA allele counting-peak ratio method (MAC-PR; Esselink et al., 2004), for the triploid hybrids that have inherited the common allele from the male genitor. The validation of the MAC-PR method for the analysed loci and populations is given as Supplementary Information.
Identification of 2_n_ gamete origin by single-locus analysis
Once female gamete structures were inferred, the percentages of heterozygosity were calculated for each locus in the whole population and for each genotype over all analysed loci. Without previous knowledge about relative markers/centromere position, the observation of heterozygosity restitution over 50% for a single locus is not informative because it could have come from either FDR or SDR; however, theoretical heterozygosity restitutions lower than 50% are only found for SDR (Park et al., 2007). When such low values of heterozygosity restitution were observed for a marker, we compared the highest probability of such a population structure under the SDR and FDR hypothesis. Under SDR, the highest probability for such an observation is obtained for a centromere position, leading to a theoretical proportion of h heterozygous gametes, whereas the best theoretical proportion of heterozygous gametes to fit with such observed data is 0.5 in the case of FDR. Thus, logarithm of the odds ratios (LODs) were estimated as:

with h being the heterozygosity transmission observed for the marker and n, the number of genotypes analysed with this marker.
Comparison of observed heterozygosity restitution within a linkage group with theoretical functions to infer the 2_n_ gamete formation mechanism and centromere position
To integrate the information on all loci of the same linkage group coming from different populations, we propose a method based on the comparison between observed heterozygosity restitution among a linkage group with theoretical restitution functions under different models. We have tested FDR and SDR mechanisms, for no-interference and partial chiasma interference on a chromosome arm (and no between-arm interference), assuming several centromere positions (interval of 0.5 cM between two theoretical adjacent positions of the centromere). Discrepancies between the different models and the observed data were estimated by the sum of the squared differences between observed and theoretical values at the marker map positions. Under one model of restitution (FDR or SDR), let Fit(c) be the value of the sum of the squared distance for each position of the centromere; the best theoretical centromere position under this model is deduced by searching c, which minimises Fit(c). The confidence interval (95%) for the centromere position was estimated by bootstrap on the loci (500 bootstraps). For this analysis, we have used the marker position of the Clementine genetic map (Ollitrault et al., 2011), which should be very similar to the ‘Fortune’ map, because ‘Fortune’ is a hybrid between Clementine and ‘Dancy’ mandarin.
No interference model
According to Zhao and Speed (1998a), assuming that the number of chiasmata in an interval follows a Poisson process (no interference model corresponding to Haldane's map function), the probabilities of a tetrad displaying a first-division segregation (FDS; Griffiths et al., 1996) pattern and a second-division segregation (SDS; Griffiths et al., 1996) pattern are as follows: p(FDS)=(1/3)(1+2_e_−3_d_) and p(SDS)=(2/3)(1−e_−3_d), where d is the genetic distance in Morgan units (Haldane’s map function) between a given locus and the centromere. Under the FDR mechanism for 2_n_ gamete formation, the FDS tetrads and half of the SDS tetrads transfer the parental heterozygosity, whereas under the SDR mechanism, the restitution of parental heterozygosity will result from SDS tetrads (Zhao and Speed, 1998b). We can, thus, derive the rates of heterozygosity transmission (H) as function of the distance to the centromere (d):
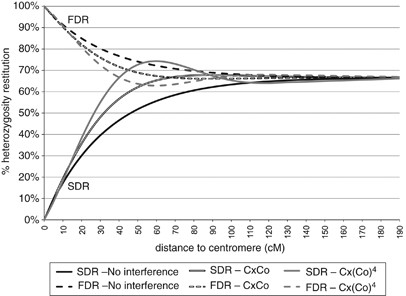
H(d)=(1/3)(1+2_e_−3_d_)+(1/3)(1−e_−3_d)=(1/3)(2+e_−3_d) for FDR and H(d)=(2/3)(1−e_−3_d) for SDR. According to this model, from the centromere to the telomere, H varies between 1 and 2/3 for FDR and from 0 to 2/3 for SDR (Figure 2).
Figure 2
Theoretical curves of heterozygosity restitution as function of genetic distance to the centromere for FDR and SDR without chiasma interference and two _χ_2-(Cx(Co)m) models of partial interference.
Let p and c, respectively, be the positions of a locus and of the centromere in a linkage group. As Haldane's map function is additive, the distance between the considered locus and the centromere is _d=_∣_p_−_c_∣. The heterozygosity restitution H(d) as a function of the distance to the centromere (d) can thus be applied to any locus position (p) on the Clementine's genetic map after transposition according to each theoretical position of the centromere (c) on the linkage group:
H(p)=(1/3)(2+_e_−3∣(p_−_c)∣) for FDR and H(p)=(2/3)(1−_e_−3∣(p_−_c)∣) for SDR. Theoretical curves of H(p) are presented in Figure 2 for FDR and SDR models.
Partial interference models
There are several proposals in the literature to incorporate chiasma interference in relating the map distance and SDS-ordered tetrad proportion. Zhao and Speed (1998a) developed functions for ordered tetrad frequencies derived from the _χ_2-chiasma interference models, which provide good fits to data from different organisms (Zhao et al, 1995). Moreover, most map functions could be approximated by one of the _χ_2-models (Zhao and Speed, 1996). The model is represented in the form Cx(Co)m, where m is a parameter positively related to the interference level. For _m_>0, the SDS proportion rises above 2/3 and as m increases, the maximal frequency of SDS increases (Zhao and Speed, 1998a). Likewise for the no interference model, the restitution of heterozygosity under FDR and SDR can be directly derived from the SDS-tetrad frequency for different _χ_2-models; as examples, Figure 2 shows the pattern of heterozygosity restitution along a linkage group for the Cx(Co) and Cx(Co)4 models. The heterozygosity restitution at any position (p) on the Clementine's genetic map can, therefore, be inferred for each theoretical position of the centromere (c) on the linkage group and thus, Fit(c) evaluated. The genetic map is established according to the map function corresponding to each considered _χ_2-model (Foss et al., 1993).
Identification of the mode of 2_n_ gamete formation and localisation of the centromere by half-tetrad multilocus structure analysis
Multilocus analyses were performed on 87 genotypes and five loci mapped in the linkage group whose order was obtained from the current map of Clementine (Ollitrault et al., 2011). The analysis was made according to Tavoletti et al. (1996), assuming that multiple crossovers do not occur between contiguous markers. Under this hypothesis, each crossover between two markers, i and _i+_1, leads to a change from homozygosity to heterozygosity in the case of SDR and a half change from heterozygosity to homozygosity in the case of FDR. Thus, the distance between two adjacent markers (_d_M_i_M_i+1) can be estimated by the proportion of 2_n gametes with changes (homozygosity versus heterozygosity; _C_M_i_M_i+_1) between the two markers. For FDR, _d_M_i_M_i+1_=_C_M_i_M_i+_1 and for SDR, _d_M_i_M_i+1=½_C_M_i_M_i+1.
The probability of the observed multilocus progeny under the different models (FDR or SDR with different centromere positions) was calculated according to Tavoletti et al. (1996). Detailed formulas can be found in Supplementary Information. Both under FDR and SDR models, the centromere position was virtually moved from before the first considered locus along the linkage group to after the last considered locus (intervals of 0.5 cM), and the relative probability was estimated for each position. The LOD between best position under SDR and FDR was calculated in order to determine the mode of 2_n_ gamete restitution, and the position of the centromere was considered as the one producing the highest relative probability under the identified mode of restitution. The confidence interval was estimated following the LOD drop-off method (Lander and Botstein, 1989). After determining the centromere position, chiasma interference can be estimated for each chromosome arm with three-point linkage mapping as follows:
Let r M1M2 denote the observed recombination rate (heterozygous to homozygous and vice versa) between the locus 1 and 2; r M1M2 the observed one between locus 2 and 3; and rd the observed rate of double recombination between the three loci. Thus, chromosome interference (I) is:

Results
Genotyping of triploid progenies
Thirty-five heterozygous SSR markers in ‘Fortune’ were selected and used for genotyping the different triploid families, according to their polymorphism between ‘Fortune’ and the male genitors. Overall, 22, 18, 21 and 26 of these markers were polymorphic between the two parents for the families with ‘Minneola’, ‘Common Mandarin’, ‘Ellendale’ and ‘Murcott’, respectively, (Table 2) as male parent. The unambiguous differentiation of allele dosage in heterozygous triploids has been confirmed by the very clear bimodal distribution of the peaks area ratio of the different triploid hybrids for all markers (Supplementary Information). The loci with total allelic differentiation between female and male genitors enabled (based on heterozygosity transmission or allele dosage estimation) the genitor producing the 2_n_ gametes for each triploid hybrid to be identified unequivocally. Maternal origin of the 2_n_ gamete has been observed for all the analysed triploids; therefore, on the basis of the maternal origin of the 2_n_ gamete and allele dosage, it was possible to infer the 2_n_ gamete structure from the triploid hybrid genotypes. Potential allelic segregation distortion in the 2_n_ gamete population was tested for each marker by _χ_12-analysis (Table 3); only one of them (Ci02B07) showed significant distortion.
Table 2 Polymorphism observed among ‘Fortune’ mandarin and the male genitors for the 35 heterozygous loci in ‘Fortune’
Table 3 Heterozygosity restitution (% HR) for each analysed locus, _χ_2-test for allelic segregation, and LOD SDR/FDR for markers with <50% of heterozygosity restitution, linkage group (LG), number of individuals (NI) and populations (NP) analysed
Maternal heterozygosity restitution in the 2_n_ eggs
Restitution of maternal heterozygosity in each 2_n_ gamete (based on all analysed loci) varied between 15.38 and 100%, with 54.98% as mean value.
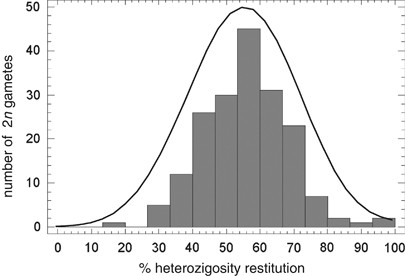
The unimodal distribution of heterozygosity restitution in the 2_n_ megaspores among the analysed genotypes suggests that all these 2_n_ gametes arise from the same mechanism (Figure 3).
Figure 3
Distribution and density trace of the heterozygosity restitution rates estimated for each 2_n_ megaspore (estimation of restitution rates based on all markers).
Global heterozygosity restitution for each marker
Rate of maternal heterozygosity restitution was calculated for 35 loci, covering eight out of nine linkage groups from the current map for Clementine (Table 3); the average rate was 55.23% and values varied from 10.34% (mCrCIR02G02 marker) up to 82.46% (mCrCIR02D09 marker). Twelve of the analysed markers displayed <50% maternal heterozygosity restitution. For these markers, LODs of SDR/FDR probabilities were calculated and varied between 0.77 and 19.05. These observations could only fulfil the SDR hypothesis, with markers that are close to the centromere, and rule out the FDR scenario.
Pattern of heterozygosity restitution within linkage group II and centromere mapping
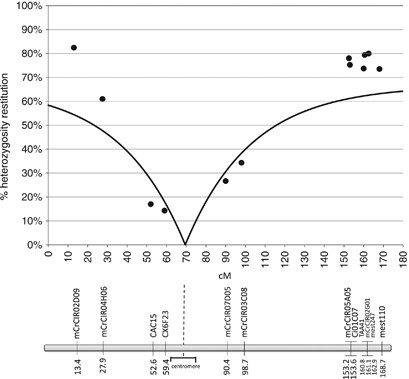
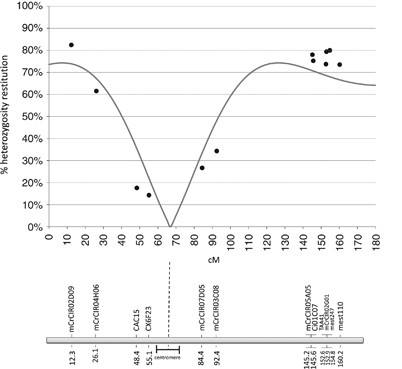
Twelve of the analysed markers in linkage group II of the Clementine genetic map (Ollitrault et al., 2011) were used to perform this analysis. Taking into account marker order and the distances between them (Haldane’s map function), the pattern of heterozygosity is represented in Figure 4 (dots). From one side of the linkage group to the other, the heterozygosity decreases from 82.46% for mCrCIR02D09 marker to 14.29% for CX6F23 marker and increases again up to 80% as the position rises on the map. Such a pattern within a linkage group is totally incompatible with the FDR model, in which the opposite variation is expected and the lowest theoretical restitution value would be 50%. Therefore, the search for the centromere position producing the best fit between the theoretical function of heterozygosity restitution and the observed data was only conducted under the SDR hypothesis. Under the SDR hypothesis with no interference, the best estimation of centromere position is 10.1±6.4 cM from CX6F23 marker (Figure 4). Without chiasma interference, the maximum theoretical heterozygosity restitution is 2/3, whereas we have observed values over 75% for five telomeric markers (two of them with very low _P-_value: mCrCIR02D09, _P_=10−5 and mCrCIR05A05, _P_=0.006), suggesting the existence of chiasma interference. On the other hand, the maximum restitution found over all analysed markers was 82.5%, whereas 100% should be expected for telomeric markers in case of total interference. Therefore, among the χ _2_-models proposed by Zhao and Speed (1998a) for partial interference, we have tested the Cx(Co)4 model, allowing a maximum restitution of heterozygosity close to 75%. Under this model of chiasma interference and SDR hypothesis, the best estimation of centromere position is 11.8±7.1 cM from CX6F23 marker (Figure 5). This model fits better with the observed data than the no interference model (Fit(c)=0.08 and Fit(c)=0.24, respectively).
Figure 4
Observed heterozygosity restitution values for markers in linkage group II (dots) and theoretical heterozygosity restitutions (line) for the best-fitting centromere position under SDR model without chiasma interference (markers in x axis are positioned according to Clementine genetic map using Haldane’s map function).
Figure 5
Observed heterozygosity restitution values for markers in linkage group II (dots) and theoretical heterozygosity restitutions (line) for the best-fitting centromere position under SDR and the Cx(Co)4 model of partial interference (markers in x axis are positioned according to Clementine genetic map using the corresponding Cx(Co)4 map function).
Multilocus half-tetrad structure analysis in linkage group II
Considering homozygosity and heterozygosity at each locus, 15 different multilocus profiles have been observed. These profiles and the number of corresponding 2_n_ gametes are given in Table 4. Probabilities of the observed 2_n_ gamete population under the FDR and SDR hypotheses for moving centromere positions have been calculated from this table. The LOD value of the SDR highest probability/FDR highest probability was 6.8, confirming that SDR is the most probable model. Under the SDR hypothesis, the probability variation as a function of the centromere position suggests that the most probable position is between 2.25 and 7.00 cM (Morgan’s map function; which is between 2.30 and 7.54 cM with Haldane’s map function) close to the CX6F23 marker, between the former and the mCrCIR05A05 marker. This confidence interval overlaps the ones of the estimations carried out by the best-fit method. If up to two crossovers per chromosome occur, it is possible to observe phase-changing between two markers when complementary crossovers take place (that is, two crossovers involving all four chromatids) via the SDR restitution mechanism (Figure 1). The HTA analysis detected 16 complementary crossovers, by revealing allelic phase changes between markers in homozygosity, as shown in Table 5; only two double crossovers affecting a chromatid pair have been identified.
Table 4 Heterozygous (HE) and homozygous (HO) profiles for 87 genotypes and five simple-sequence repeat markers within linkage group II
Table 5 Number of observed crossing over events on each arm in chromosome II for 87 genotypes and five markers
Interference analysis
Considering centromere position between CX6F23 and mCrCIR07D05 markers, the interference coefficient was estimated for each arm of chromosome II with three markers per side. For arm 1, analysing 87 genotypes for mCrCIR02D09, mCrCIR04H06 and CX6F23 markers, the interference coefficient was found to be 0.73. For the other arm, the interference estimation was 0.53, analysing 66 2_n_ gametes for mCrCIR03C08, mCrCIR05A05 and TAA41 markers.
Discussion
MAC-PR is an efficient method to determine allelic configurations in triploid citrus-segregating progenies
In this work, the HTA is based on gamete allelic configuration inferred from triploid progeny genotyping. MAC-PR has been proposed to deal with differential amplification intensities among alleles in polyploid plant species (Esselink et al., 2004) for allele dosage estimation. Under the PCR conditions used for citrus SSR analysis, we have successfully verified the correlation between allele dosage and PCR product ratio. Finally, the very clear bimodal distribution of estimated doses for the 35 analysed SSR markers among triploid hybrids ruled out the occurrence of random PCR drift in our amplifications and validated the MAC-PR approaches for triploid citrus progenies genotyping. A basic assumption of the MAC-PR method is the repeatability of relative allelic amplification intensities among individuals and, thus, the total homology of primer sites for DNA fragments producing the same allele (same PCR product size). Homoplasy of SSR markers was found at interspecific levels in citrus (Barkley et al., 2009), and could limit general application of MAC-PR. ‘Fortune’ mandarin and most of the male genitors used in our study are closely related, which should explain why we have not encountered this difficulty because of the by-descent homology of alleles.
Origin of 2_n_ gamete-producing triploids in citrus
We observed that all the 171 analysed triploid hybrids show maternal heterozygosity restitution for at least one marker. This confirms that all triploid hybrids found in the progenies of 2_x_ × 2_x_ crosses with ‘Fortune’ mandarin as female genitor arose from 2_n_ megaspores. This result is in agreement with the cytogenetic observations of Esen and Soost (1971), and with previous molecular observations (Luro et al., 2000; Chen et al., 2008; Ferrante et al., 2010). In the present work, the restitution heterozygosity rates significantly lower than 50% for several markers (distributed in six of the eight represented linkage groups) and the pattern of heterozygosity inside linkage group II, definitively ruled out the FDR hypothesis. Multilocus 2_n_ gamete allelic configuration in the same linkage group also revealed that SDR was much more likely to be the mechanism underlying the unreduced gamete formation than FDR (LOD=6.8). Moreover, this analysis enabled us to detect four-strand (complementary) double crossovers by phase changes of several homozygous markers. Such phase changes between homozygous positions are only possible under the SDR hypothesis if up to two crossovers per arm are considered. This conclusion for SDR is in agreement with that proposed by Luro et al., 2000, who observed low heterozygosity restitution in C. clementina 2_n_ megagametophyte. The conclusion of FDR given for sweet orange (Chen et al., 2008) is questionable because of the low number of analysed markers. Indeed, the unambiguous identification of FDR without previous location of the centromere must be based on a large set of markers with good genome coverage. In the same way, the results of Ferrante et al. (2010), based on a very low number of individuals and markers for each parental genotype are not sufficient to prove the authors’s conclusions of SDR for ‘Fortune’ and ‘Wilking’ mandarin and FDR for lemon. Systematic analysis of 2_n_ gamete allelic configuration with the same set of loci, close and far from the centromere, will shed light on whether SDR is the only mechanism underlying 2_n_ egg formation in citrus, or whether there is a different mechanism depending on genotype or environmental conditions.
Analysis of the pattern of heterozygosity restitution within a linkage group is an alternative way to map the centromere compared with half-tetrad multilocus allelic configuration analysis
Centromere mapping has been carried out in several crops (Douches and Quiros, 1988; Okagaki et al., 2008) and animals (Kauffman et al., 1995; Lindner et al., 2000), using HTA. In the present work, HTA has been carried out in two ways: by multilocus allelic conformation analysis, as described in Tavoletti et al. (1996) and by comparison of observed and theoretical pattern of heterozygosity restitution rate within the linkage group under several models. Both methods estimated the centromere position to be between CX6F23 and mCrCIR07D05 markers. Confidence intervals of the positions obtained with the two methods overlapped, validating the best-fit method. The fitting curve adjustment has the advantage of potential application to a set of loci analysed in different progenies (between a same parent producing the 2_n_ eggs but different male parents), potentially enlarging the usable set of markers to all those heterozygous for the female 2_n_ gamete producer. It could potentially be used to compare a large range of interference model functions. Furthermore, this method should be easily applied to dominant markers by estimating the heterozygous restitution as 1−2_f_ (with f being the frequency of homozygous progeny for the recessive allele). However, it requires the use of an existing genetic map and assumes that crossover distribution is similar during normal meiosis and 2_n_ gamete formation. Multilocus allelic HTA is advantageous in that it can be used without a previous genetic map of the markers and can be applied with a predefined order of markers (Tavoletti et al., 1996) as we have performed, or without any previous information about marker position (Da et al., 1995). An excel template has been developed for easy identification of restitution model (FDR or SDR) and positioning of centromere within a linkage group, from heterozygosity restitution data for a set of mapped loci. It includes an estimation of confidence interval for the centromere position by bootstrap on the loci. It is available upon request to the authors.
Evidence for positive chiasma interference in citrus cv ‘Fortune’
Many models of HTA are based on the hypothesis of complete interference. In the present work, the analysis of multilocus configuration in linkage group II revealed the occurrence of up to four crossovers in the same chromosome arm and thus, incomplete interference. It is confirmed by the maximum restitution values between 75 and 82% observed for five of the markers, whereas for SDR, the maximum restitution should be 66.6% under the no interference hypothesis and should reach 100% for total interference. A better adjustment was found between observed data and theoretical curve with the Cx(Co)4 _χ_2-model for partial interference than the no interference one. Interference values were estimated by three-point analysis for each arm of linkage group II, with results suggesting that it was higher for one arm (0.73) than for the other (0.53). Such variation of interference level between different parts of the genome has also been observed in Arabidopsis (Drouaud et al., 2007), in human (Lian et al., 2008) and in mouse (Broman et al., 2002).
Implications for citrus breeding
Seedlessness is one of the most important characteristics for the citrus fresh-fruit market. An efficient way to achieve this aim is to obtain triploid mandarin varieties. Sexual triploidisation is a classical method to obtain triploid citrus hybrids (Ollitrault et al., 2008). Indeed, some genotypes such as ‘Temple’, ‘Wilking’ and ‘Fortune’ mandarin (Esen and Soost, 1973; Aleza et al., 2010) are well known for their production of diploid megagametophytes. The other classical method to create triploid citrus progenies is inter-ploidy hybridisation with doubled-diploid. Assuming an SDR origin of 2_n_ gametes in ‘Fortune mandarin’, sexual polyploidisation may lead to lower average of heterozygosity restitution than interploid hybridisation, whatever the segregation model considered for the doubled diploid (Marsden et al., 1987). As heterozygosity and epistatic interactions are maintained for a great number of individuals in the progeny from interploid crosses with doubled-diploid, this triploid breeding strategy should be more efficient than 2_x_ × 2_x_ hybridisation for developing new cultivars that are phenotypically close to ‘Fortune mandarin’ genitor. Conversely, 2_x_ × 2_x_ hybridisation should produce more polymorphic progenies, by creating larger number of new multilocus allelic combinations (David et al., 1995), providing the opportunity to select innovative products within the perspective of market segmentation as a commercial strategy.
References
- Aleza P, Froelicher Y, Schwarz S, Agustí M, Hernández M, Juárez J et al. (2011). Tetraploidization events by chromosome doubling of nucellar cells are frequent in apomictic citrus and are dependent on genotype and environment. Ann Bot (doi:10.1093/aob/mcr099).
- Aleza P, Juárez J, Cuenca J, Ollitrault P, Navarro L (2010). Recovery of citrus triploid hybrids by embryo rescue and flow cytometry from 2x × 2x sexual hybridisation and its application to extensive breeding programs. Plant Cell Rep 29: 1023–1034.
Google Scholar - Barkley NL, Krueger R, Federici CT, Roose ML (2009). What phylogeny and gene genealogy analyses reveal about homoplasy in citrus microsatellite alleles. Plant Syst Evol 282: 71–86.
Google Scholar - Broman KW, Rowe LB, Churchill GA, Paigen K (2002). Crossover interference in the mouse. Genetics 160: 1123–1131.
Google Scholar - Chen C, Bowman KD, Ya C, Dang PM, Rao MN, Huang S et al. (2007). EST-SSR genetic maps for Citrus sinensis and Poncirus trifoliate. Tree Genet Genomes 4: 1–10.
Google Scholar - Chen C, Lyon MT, O’Malley D, Federici CT, Gmitter J, Grosser JW et al. (2008). Origin and frequency of 2n gametes in Citrus sinensis × Poncirus trifoliata and their reciprocal crosses. Plant Sci 174: 1–8.
Google Scholar - Da Y, Jarrell VL, Wang T, Fernando RL, Wheeler MB, Lewi HA (1995). Multilocus analysis for gene-centromere mapping using first polar bodies and secondary oocytes. Genetics 139: 1091–1097.
Google Scholar - David JL, Boudec P, Gallais A (1995). Quantitative genetics of 4x-2x hybrid populations with first-division restitution and second-division restitution 2n gametes produced by diploid parents. Genetics 139: 1797–1803.
Google Scholar - Douches DS, Quiros CF (1988). Genetic strategies to determine the mode of 2n egg formation in diploid potatoes. Euphytica 38: 247–260.
Google Scholar - Drouaud J, Mercier R, Chelysheva L, Berard A, Falque M, Martin O et al. (2007). Sex-specific crossover distributions and variations in interference level along Arabidopsis thaliana chromosome 4. PLoS Genet 3: e106.
Google Scholar - Esen A, Soost RK (1971). Unexpected triploids in citrus: their origin, identification and possible use. J Hered 62: 329–333.
Google Scholar - Esen A, Soost RK (1973). Precocious development and germination of spontaneous triploid seed in citrus. J Hered 64: 147–154.
Google Scholar - Esen A, Soost RK, Geraci G (1979). Genetic evidence for the origin of diploid megagametophytes in citrus. J Hered 70: 5–8.
Google Scholar - Esselink GD, Nybom H, Vosman B (2004). Assignment of allelic configuration in polyploids using the MAC-PR (microsatellite DNA allele counting—peak ratios) method. Theor Appl Genet 109: 402–408.
Google Scholar - Ferrante SP, Lucretti S, Reale S, De Patrizio A, Abbate L, Tusa N et al. (2010). Assessment of the origin of new citrus tetraploid hybrids (2n=4x) by means of SSR markers and PCR based dosage effects. Euphytica 173: 223–233.
Google Scholar - Foss E, Lande R, Stahl FW, Steinberg CM (1993). Chiasma interference as a function of genetic distance. Genetics 133: 681–691.
Google Scholar - Froelicher Y, Dambier D, Costantino G, Lotfy S, Didout C, Beaumont V et al. (2008). Characterization of microsatellite markers in Citrus reticulata Blanco. Mol Ecol Resour 8: 119–122.
Google Scholar - Griffiths AJF, Miller JH, Suzuki DT, Lewontin RC, Gelbart WM (1996). An Introduction To Genetic Analysis, Ed 6 W H Freeman and Company, New York.
Google Scholar - Kamiri M, Stift M, Srairi I, Costantino G, Moussadik A, Hmyene A . et al. (2011). Evidence for non-disomic inheritance in a Citrus interspecific tetraploid somatic between C. reticulata and C. lemon hybrid using SSR markers and cytogenetic analysis. Plant Cell Rep (doi:10.1007/s00299-011-1050-x).
- Kauffman EJ, Gest EE, Kim DJ, Walker C, Hite JM, Yan G et al. (1995). Microsatellite–centromere mapping in the Zebrafish (Danio rerio). Genomics 30: 337–341.
Google Scholar - Kijas JMH, Thomas MR, Fowler JCS, Roose ML (1997). Integration of trinucleotide microsatellites into a linkage map of citrus. Theor Appl Genet 94: 701–706.
Google Scholar - Lander ES, Botstein D (1989). Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121: 185–199.
Google Scholar - Lian J, Yin Y, Oliver-Bonet M, Liehr T, Ko E, Turek P et al. (2008). Variation in crossover interference levels on individual chromosomes from human males. Hum Mol Genet 17: 2583–2594.
Google Scholar - Lindner KR, Seeb JE, Habicht C, Knudsen KL, Kretschmer E, Reedy DJ et al. (2000). Gene–centromere mapping of 312 loci in pink salmon by half-tetrad analysis. Genome 43: 538–549.
Google Scholar - Luro F, Costantino G, Terol J, Argout X, Allario T, Wincker P et al. (2008). Transferability of the EST-SSRs developed on Nules clementine (Citrus clementina Hort ex Tan) to other Citrus species and their effectiveness for genetic mapping. BMC Genomics 9: 287.
Google Scholar - Luro F, Rist D, Ollitrault P (2000). Evaluation of genetic relationships in Citrus genus by means of sequence tagged microsatellites. In: Doré C, Dosba F, Baril C (eds) Proceedings of the International Symposium on Molecular Markers for Characterizing Genotypes and Identifying Cultivars in Horticulture. March 6–8 2000. Volume 546 ISHS: Acta Horticulturae 2001: 537–542.
- Marsden JE, Schwager SJ, May B (1987). Single-locus inheritance in the tetraploid treefrog Hyla versicolor with an analysis of expected progeny ratios in tetraploid organisms. Genetics 116: 299–311.
Google Scholar - Mendiburu AO, Peloquin SJ (1976). Sexual polyploidization and depolyploidization: some terminology and definitions. Theor Appl Genet 48: 137–143.
Google Scholar - Okagaki RJ, Jacobs MS, Stec AO, Kynast RG, Buescher E, Rines HW et al. (2008). Maize centromere mapping: a comparison of physical and genetic strategies. J Hered 99: 85–93.
Google Scholar - Ollitrault P, Dambier D, Luro F, Froelicher Y (2008). Ploidy manipulation for breeding seedless triploid citrus. In: Plant Breed Rev 30: 323–352.
Google Scholar - Ollitrault P, Terol J, Chen C, Federici CT, Lotfy S, Hippolyte I et al. (2011). Reference SNPs, SSRs and InDels C. clementina linkage map. Plant Animal Genome. XIX Conference San Diego, California, USA, P477.
Google Scholar - Park TH, Kim JB, Hutten RCB, van Eck HJ, Jacobsen E, Visser RGE (2007). Genetic positioning of centromeres using half-tetrad analysis in a 4x-2x cross population of potato. Genetics 176: 85–94.
Google Scholar - Ramanna MS, Jacobsen E (2003). Relevance of sexual polyploidization for crop improvement–A review. Euphytica 133: 3–18.
Google Scholar - Tavoletti S, Bingham ET, Yandell B S, Veronesi F, Osborn TC (1996). Half tetrad analysis in alfalfa using multiple restriction fragment length polymorphism markers. Proc Natl Acad Sci USA 93: 10918–10922.
Google Scholar - Zhao H, Speed TP, McPeek MS (1995). Statistical analysis of crossover interference using the chi-square model. Genetics 139: 1045–1056.
Google Scholar - Zhao H, Speed TP (1996). On genetic map functions. Genetics 142: 1369–1377.
Google Scholar - Zhao H, Speed TP (1998a). Statistical analysis of ordered tetrads. Genetics 150: 459–472.
Google Scholar - Zhao H, Speed TP (1998b). Statistical analysis of half-tetrads. Genetics 150: 473–485.
Google Scholar
Acknowledgements
We thank Frédérique Ollitrault, Ana M Moreno and J Joaquín Saurí for their contribution to this work. We also thank François Luro from INRA (France) for providing some unpublished SSRs primers. This work is jointly financed by the AGL2008-00596-MCI and Prometeo 2008/121 Generalitat Valenciana projects.
Author information
Authors and Affiliations
- Instituto Valenciano de Investigaciones Agrarias, Centro de Protección Vegetal y Biotecnología, Ctra, Moncada, Valencia, Spain
J Cuenca, P Aleza, J Juárez, L Navarro & P Ollitrault - BIOS Department, Centre de Coopération Internationale en Recherche Agronomique pour le Développement (CIRAD), UPR amélioration génétique des espèces à multiplication végétative, Montpellier cedex, France
Y Froelicher & P Ollitrault
Authors
- J Cuenca
You can also search for this author inPubMed Google Scholar - Y Froelicher
You can also search for this author inPubMed Google Scholar - P Aleza
You can also search for this author inPubMed Google Scholar - J Juárez
You can also search for this author inPubMed Google Scholar - L Navarro
You can also search for this author inPubMed Google Scholar - P Ollitrault
You can also search for this author inPubMed Google Scholar
Corresponding author
Correspondence toP Ollitrault.
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Cuenca, J., Froelicher, Y., Aleza, P. et al. Multilocus half-tetrad analysis and centromere mapping in citrus: evidence of SDR mechanism for 2_n_ megagametophyte production and partial chiasma interference in mandarin cv ‘Fortune’.Heredity 107, 462–470 (2011). https://doi.org/10.1038/hdy.2011.33
- Received: 09 September 2010
- Revised: 25 March 2011
- Accepted: 31 March 2011
- Published: 18 May 2011
- Issue Date: November 2011
- DOI: https://doi.org/10.1038/hdy.2011.33