In silico simulations reveal that replicators with limited dispersal evolve towards higher efficiency and fidelity (original) (raw)
- Letter
- Published: 21 November 2002
Nature volume 420, pages 340–343 (2002)Cite this article
- 1220 Accesses
- 149 Citations
- 3 Altmetric
- Metrics details
Abstract
The emergence of functional replicases, acting quickly and with high accuracy, was crucial to the origin of life1. Although where the first RNA molecules came from is still unknown, it is nevertheless assumed that catalytic RNA enzymes (ribozymes) with replicase function emerged at some early stage of evolution1. The fidelity of copying is especially important because the mutation load limits the length of replicating templates that can be maintained by natural selection2. An increase in template length is disadvantageous for a fixed digit copying fidelity, however, longer molecules are expected to be better replicases. An iteration for longer molecules with better replicase function has been suggested3,4 and analysed mathematically5. Here we show that more efficient replicases can spread, provided they are adsorbed to a prebiotic mineral surface. A cellular automaton6 simulation reveals that copying fidelity, replicase speed and template efficiency all increase with evolution, despite the presence of molecular parasites, essentially because of reciprocal atruism7 (‘within-species mutualism’) on the surface8, thus making a gradual improvement of replicase function more plausible.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Long-term evolution of replicators (population-wide averages).
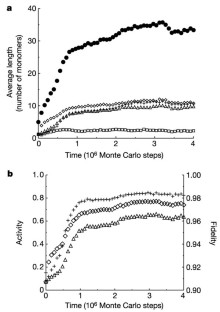
The alternative text for this image may have been generated using AI.
Figure 2: Distribution of replicators at the stationary state.
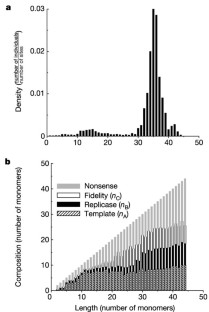
The alternative text for this image may have been generated using AI.
Figure 3: Replicator model with diffusion (population-wide averages).
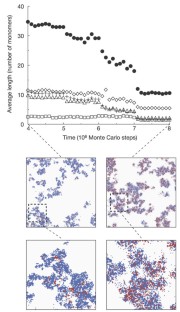
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Joyce, G. F. & Orgel, L. E. in The RNA World, 2nd edn (eds Gesteland, R. F., Cech, T. R. & Atkins, J. F.) 49–77 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1999)
Google Scholar - Eigen, M. Self-organization of matter and the evolution of biological macromolecules. Naturwissenschatten 58, 465–523 (1971)
Article ADS CAS Google Scholar - Poole, A., Jeffares, D. & Penny, D. Early evolution: the new kids on the block. BioEssays 21, 880–889 (1999)
Article CAS Google Scholar - James, K. D. & Ellington, A. D. The fidelity of template-directed oligonucleotide ligation and the inevitability of polymerase function. Orig. Life Evol. Biosphere 29, 375–390 (1998)
Article ADS Google Scholar - Scheuring, I. Avoiding catch-22 of early evolution by stepwise increase in copying fidelity. Selection 1, 135–145 (2000)
Article Google Scholar - Wolfram, S. A New Kind of Science (Wolfram Media, Champaign, Illinois, 2002)
MATH Google Scholar - Trivers, R. L. The evolution of reciprocal altruism. Q. Rev. Biol. 46, 35–57 (1971)
Article Google Scholar - Nowak, M. & Sigmund, K. Tit for tat in heterogeneous populations. Nature 355, 250–253 (1992)
Article ADS Google Scholar - Inoue, T. & Orgel, L. E. A nonenzymatic RNA polymerase model. Science 219, 859–862 (1983)
Article ADS CAS Google Scholar - Florentz, C. & Giegé, R. tRNA: Structure, Biosynthesis, Function (eds Söll, D. & RajBhandary, U. L.) 141–164 (American Society of Microbiology Press, Washington, 1995)
Book Google Scholar - Galas, D. J. & Branscomb, E. W. Enzymatic determinants of DNA polymerase accuracy. Theory of coliphage T4 polymerase mechanisms. J. Molec. Biol. 124, 653–687 (1978)
Article CAS Google Scholar - Goodman, M. F. & Fygenson, D. K. DNA polymerase fidelity: From genetics toward a biochemical understanding. Genetics 148, 1475–1482 (1998)
CAS PubMed PubMed Central Google Scholar - Wang, J. et al. Crystal structure of a pol α family replication DNA polymerase from bacteriophage RB69. Cell 89, 1087–1099 (1997)
Article CAS Google Scholar - Doublié, S. et al. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 391, 251–258 (1998)
Article ADS Google Scholar - Kiefer, J. R. et al. Visualizing DNA replication in a catalytically active Bacillus DNA polymerase crystal. Nature 391, 304–307 (1998)
Article ADS CAS Google Scholar - Steitz, T. A. Structural biology. A mechanism for all polymerases. Nature 391, 231–232 (1998)
Article ADS CAS Google Scholar - Toffoli, T. & Margolus, N. Cellular Automata Machines: A New Environment for Modeling 155–167 (MIT Press, Cambridge, Massachusetts, 1987)
MATH Google Scholar - Boerlijst, M. C. & Hogeweg, P. Spiral wave structure in prebiotic evolution: Hypercycles stable against parasites. Physica D. 48, 17–28 (1991)
Article ADS Google Scholar - Czárán, T. & Szathmáry, E. The Geometry of Ecological Interactions: Simplifying Spatial Complexity 116–134 (Cambridge Univ. Press, Cambridge, 2000)
Book Google Scholar - Szathmáry, E. & Demeter, L. Group selection of early replicators and the origin of life. J. Theoret. Biol. 128, 463–486 (1987)
Article Google Scholar - Szathmáry, E. & Maynard Smith, J. The origin of chromosomes II. Molecular mechanisms. J. Theoret. Biol. 164, 447–454 (1993)
Article Google Scholar - Ferris, J. P. et al. Synthesis of long prebiotic oligomers on mineral surfaces. Nature 381, 59–61 (1996)
Article ADS CAS Google Scholar - Orgel, L. E. Polymerisation on the rocks: Theoretical introduction. Orig. Life Evol. Biosphere 28, 227–234 (1998)
Article ADS CAS Google Scholar - von Kiedrowski, G. Primordial soup or crêpes? Nature 381, 20–21 (1996)
Article ADS CAS Google Scholar - von Kiedrowski, G. Surface-promoted replication and exponential amplification of DNA analogues. Nature 396, 245–248 (1998)
Article ADS Google Scholar - von Kiedrowski, G. & Szathmáry, E. Selection versus coexistence of parabolic replicators spreading on surfaces. Selection 1, 173–179 (2000)
Article Google Scholar - Johnston, W. K. et al. RNA-catalyzed RNA polymerization: accurate and general RNA-templated primer extension. Science 292, 1319–1325 (2001)
Article ADS MathSciNet CAS Google Scholar
Acknowledgements
Financial support by the National Research Fund (OTKA) is gratefully acknowledged.
Author information
Authors and Affiliations
- Department of Plant Taxonomy and Ecology, Research Group of Ecology and Theoretical Biology, Eötvös University, 1/c Pázmány P. sétány, H-1117, Budapest
Péter Szabó, István Scheuring, Tamás Czárán & Eörs Szathmáry - Adaptive Dynamics Network, International Institute for Applied Systems Analysis, A-2361, Laxenburg, Austria
Péter Szabó - Collegium Budapest (Institute for Advanced Study) 2 Szentháromság u., H-1014, Budapest, Hungary
Eörs Szathmáry
Authors
- Péter Szabó
- István Scheuring
- Tamás Czárán
- Eörs Szathmáry
Corresponding author
Correspondence toEörs Szathmáry.
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Szabó, P., Scheuring, I., Czárán, T. et al. In silico simulations reveal that replicators with limited dispersal evolve towards higher efficiency and fidelity.Nature 420, 340–343 (2002). https://doi.org/10.1038/nature01187
- Received: 08 May 2002
- Accepted: 13 September 2002
- Issue date: 21 November 2002
- DOI: https://doi.org/10.1038/nature01187