Adaptive prediction of environmental changes by microorganisms (original) (raw)
- Article
- Published: 17 June 2009
- Gal H. Romano2,
- Bella Groisman1,
- Avihu Yona1,
- Erez Dekel3,
- Martin Kupiec2,
- Orna Dahan1 na1 &
- …
- Yitzhak Pilpel1,4 na1
Nature volume 460, pages 220–224 (2009)Cite this article
- 12k Accesses
- 575 Citations
- 66 Altmetric
- Metrics details
Abstract
Natural habitats of some microorganisms may fluctuate erratically, whereas others, which are more predictable, offer the opportunity to prepare in advance for the next environmental change. In analogy to classical Pavlovian conditioning, microorganisms may have evolved to anticipate environmental stimuli by adapting to their temporal order of appearance. Here we present evidence for environmental change anticipation in two model microorganisms, Escherichia coli and Saccharomyces cerevisiae. We show that anticipation is an adaptive trait, because pre-exposure to the stimulus that typically appears early in the ecology improves the organism’s fitness when encountered with a second stimulus. Additionally, we observe loss of the conditioned response in E. coli strains that were repeatedly exposed in a laboratory evolution experiment only to the first stimulus. Focusing on the molecular level reveals that the natural temporal order of stimuli is embedded in the wiring of the regulatory network—early stimuli pre-induce genes that would be needed for later ones, yet later stimuli only induce genes needed to cope with them. Our work indicates that environmental anticipation is an adaptive trait that was repeatedly selected for during evolution and thus may be ubiquitous in biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Four possible regulation strategies in response to environmental stimuli.
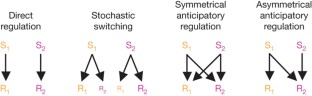
The alternative text for this image may have been generated using AI.
Figure 2: Conditioned response in E. coli sugar metabolism.
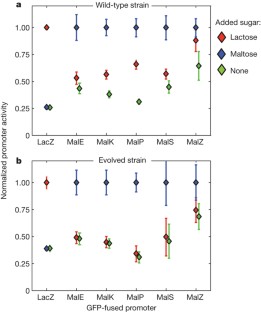
The alternative text for this image may have been generated using AI.
Figure 3: Fitness in an alternating sugar environment.
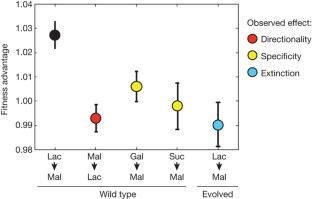
The alternative text for this image may have been generated using AI.
Figure 4: Cross-protection in the context of the diauxic shift.
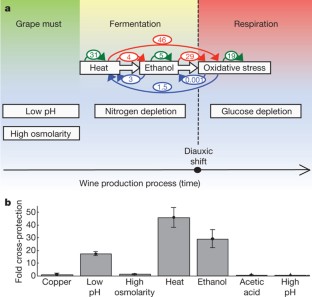
The alternative text for this image may have been generated using AI.
Figure 5: Candidate genes underlying the asymmetrical protection between heat and oxidative stresses.
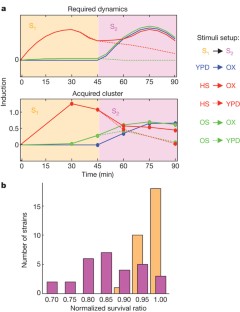
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Causton, H. C. et al. Remodeling of yeast genome expression in response to environmental changes. Mol. Biol. Cell 12, 323–337 (2001)
Article CAS Google Scholar - Gasch, A. P. et al. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 11, 4241–4257 (2000)
Article CAS Google Scholar - Cheung, K. J. et al. A microarray-based antibiotic screen identifies a regulatory role for supercoiling in the osmotic stress response of Escherichia coli. Genome Res. 13, 206–215 (2003)
Article CAS Google Scholar - Tirosh, I., Weinberger, A., Carmi, M. & Barkai, N. A genetic signature of interspecies variations in gene expression. Nature Genet. 38, 830–834 (2006)
Article CAS Google Scholar - Kussell, E. & Leibler, S. Phenotypic diversity, population growth, and information in fluctuating environments. Science 309, 2075–2078 (2005)
Article ADS CAS Google Scholar - Acar, M., Mettetal, J. T. & van Oudenaarden, A. Stochastic switching as a survival strategy in fluctuating environments. Nature Genet. 40, 471–475 (2008)
Article CAS Google Scholar - Balaban, N. Q. et al. Bacterial persistence as a phenotypic switch. Science 305, 1622–1625 (2004)
Article ADS CAS Google Scholar - Pavlov, I. P. Conditioned Reflexes (Oxford Univ. Press, 1927)
Google Scholar - Tagkopoulos, I., Liu, Y. C. & Tavazoie, S. Predictive behavior within microbial genetic networks. Science 320, 1313–1317 (2008)
Article ADS CAS Google Scholar - Savageau, M. A. Escherichia coli habitats, cell-types, and molecular mechanisms of gene control. Am. Nat. 122, 732–744 (1983)
Article CAS Google Scholar - Savageau, M. A. Demand theory of gene regulation. II. Quantitative application to the lactose and maltose operons of Escherichia coli. Genetics 149, 1677–1691 (1998)
CAS PubMed PubMed Central Google Scholar - Zaslaver, A. et al. A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nature Methods 3, 623–628 (2006)
Article CAS Google Scholar - Soupene, E. et al. Physiological studies of Escherichia coli strain MG1655: Growth defects and apparent cross-regulation of gene expression. J. Bacteriol. 185, 5611–5626 (2003)
Article CAS Google Scholar - Oxman, E., Alon, U. & Dekel, E. Defined order of evolutionary adaptations: experimental evidence. Evolution 62, 1547–1554 (2008)
Article Google Scholar - Mortimer, R. K. Evolution and variation of the yeast (Saccharomyces) genome. Genome Res. 10, 403–409 (2000)
Article CAS Google Scholar - Sablayrolles, J. M. & Barre, P. Kinetics of alcoholic fermentation under anisothermal enological conditions. Am. J. Enol. Vitic. 44, 127–133 (1993)
CAS Google Scholar - Johnston, M. & Carlson, M. in The Molecular Biology of the Yeast Saccharomyces: Gene Expression (eds Jones, E. W., Pringle, J. R. & Broach, J. R.) 193–282 (Cold Spring Harbor Laboratory, 1992)
Google Scholar - Bauer, F. F. & Pretorius, I. S. Yeast stress response and fermentation efficiency: How to survive the making of wine - a review. SAJEV 21, 27–51 (2000)
CAS Google Scholar - Pretorius, I. S. Tailoring wine yeast for the new millennium: novel approaches to the ancient art of winemaking. Yeast 16, 675–729 (2000)
Article CAS Google Scholar - Maris, A. F. et al. Diauxic shift-induced stress resistance against hydroperoxides in Saccharomyces cerevisiae is not an adaptive stress response and does not depend on functional mitochondria. Curr. Genet. 39, 137–149 (2001)
Article CAS Google Scholar - Sanchez, Y., Taulien, J., Borkovich, K. A. & Lindquist, S. Hsp104 is required for tolerance to many forms of stress. EMBO J. 11, 2357–2364 (1992)
Article CAS Google Scholar - Berry, D. B. & Gasch, A. P. Stress-activated genomic expression changes serve a preparative role for impending stress in yeast. Mol. Biol. Cell 19, 4580–4587 (2008)
Article CAS Google Scholar - Harris, M. A. et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32 (Database issue). D258–D261 (2004)
Article ADS CAS Google Scholar - Grant, C. M., MacIver, F. H. & Dawes, I. W. Mitochondrial function is required for resistance to oxidative stress in the yeast Saccharomyces cerevisiae. FEBS Lett. 410, 219–222 (1997)
Article CAS Google Scholar - Zacharioudakis, I., Gligoris, T. & Tzamarias, D. A yeast catabolic enzyme controls transcriptional memory. Curr. Biol. 17, 2041–2046 (2007)
Article CAS Google Scholar - Braun, E. & Brenner, N. Transient responses and adaptation to steady state in a eukaryotic gene regulation system. Phys. Biol. 1, 67–76 (2004)
Article ADS CAS Google Scholar - Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999)
Article CAS Google Scholar - Kaplan, S. et al. Diverse two-dimensional input functions control bacterial sugar genes. Mol. Cell 29, 786–792 (2008)
Article CAS Google Scholar - Dekel, E. & Alon, U. Optimality and evolutionary tuning of the expression level of a protein. Nature 436, 588–592 (2005)
Article ADS CAS Google Scholar
Acknowledgements
We thank S. Trattner–Frenkel and Z. Bloom for their help and support in the yeast microarray experiments. We thank members of the Pilpel laboratory for many discussions. We thank E. Schneidman, E. Ben-Jacob, M. Springer, A. Tanay, U. Alon and D. Cavalieri for discussions and advice. We thank U. Alon for providing the promoter–GFP fused plasmids. We thank the Tauber Foundation, the Minerva Foundation, the Israel Science Foundation ‘Bikura program’, the European Research Council ‘Ideas Program’ and the Ben May Foundation for grant support. M.K. was supported from grants from the Israel Science Foundation and the Israeli Ministry of Science and Technology.
Author Contributions A.M. raised the original idea and performed all the experiments; G.R., B.G. and A.Y. participated in experiments; E.D. evolved the E. coli strain; A.M., O.D. and Y.P. designed the experiments; A.M., M.K., O.D. and Y.P. analysed the data; O.D. and Y.P. supervised the project; A.M., O.D. and Y.P. interpreted the results and wrote the manuscript.
Author information
Author notes
- Orna Dahan and Yitzhak Pilpel: These authors contributed equally to this work.
Authors and Affiliations
- Department of Molecular Genetics, Weizmann Institute of Science Rehovot 76100, Israel
Amir Mitchell, Bella Groisman, Avihu Yona, Orna Dahan & Yitzhak Pilpel - Department of Molecular Microbiology and Biotechnology, Tel Aviv University, Tel Aviv 69978, Israel,
Gal H. Romano & Martin Kupiec - Department of Molecular Cell Biology, Weizmann Institute of Science Rehovot 76100, Israel
Erez Dekel - Department of Systems Biology, Harvard Medical School, Boston, Massachusetts 02115, USA,
Yitzhak Pilpel
Authors
- Amir Mitchell
- Gal H. Romano
- Bella Groisman
- Avihu Yona
- Erez Dekel
- Martin Kupiec
- Orna Dahan
- Yitzhak Pilpel
Corresponding author
Correspondence toYitzhak Pilpel.
Supplementary information
Supplementary information (download PDF )
This file contains Supplementary Methods, Supplementary Tables 1-5, Supplementary Equation1, Supplementary Figures 1-8 with Legends, Supplementary Notes and Supplementary References. (PDF 324 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Mitchell, A., Romano, G., Groisman, B. et al. Adaptive prediction of environmental changes by microorganisms.Nature 460, 220–224 (2009). https://doi.org/10.1038/nature08112
- Received: 25 March 2009
- Accepted: 07 May 2009
- Published: 17 June 2009
- Issue date: 09 July 2009
- DOI: https://doi.org/10.1038/nature08112
This article is cited by
Editorial Summary
Microorganism behaviour: be prepared
Microorganisms, at first glance, look to be at the mercy of their environment and any changes that might take place within it. Reacting to events as they happen would seem to be their lot. So the finding that both bacteria and yeast, in environments where a sequence of changes follows a repeated pattern, can associate a stimulus with an appropriate response to a future environment comes as something of a surprise. In a process that resembles Pavlovian conditioning in some ways — but depends on regulatory networks and natural selection rather than cognition — Escherichia coli passing through the gut and yeast through the various stages of fermentation 'anticipate' their next experience and assemble the metabolic pathways to cope with it. E. coli later exposed repeatedly to only the first of the series of environments even 'forget' their training and lose the conditioned response.