A regulatory circuit for piwi by the large Maf gene traffic jam in Drosophila (original) (raw)
- Letter
- Published: 07 October 2009
- Sachi Inagaki1,
- Toutai Mituyama2,
- Yoshinori Kawamura1,3,
- Yukiteru Ono4,
- Eri Sakota5,
- Hazuki Kotani1,
- Kiyoshi Asai2,6,
- Haruhiko Siomi1 &
- …
- Mikiko C. Siomi1,7
Nature volume 461, pages 1296–1299 (2009)Cite this article
- 7147 Accesses
- 426 Citations
- 17 Altmetric
- Metrics details
Abstract
PIWI-interacting RNAs (piRNAs) silence retrotransposons in Drosophila germ lines by associating with the PIWI proteins Argonaute 3 (AGO3), Aubergine (Aub) and Piwi1,2. piRNAs in Drosophila are produced from intergenic repetitive genes and piRNA clusters by two systems: the primary processing pathway and the amplification loop1,2,3,4,5,6,7. The amplification loop occurs in a Dicer-independent, PIWI-Slicer-dependent manner3,4,8. However, primary piRNA processing remains elusive. Here we analysed piRNA processing in a Drosophila ovarian somatic cell line where Piwi, but not Aub or AGO3, is expressed; thus, only the primary piRNAs exist. In addition to flamenco, a Piwi-specific piRNA cluster3, traffic jam (tj)9, a large Maf gene, was determined as a new piRNA cluster. piRNAs arising from tj correspond to the untranslated regions of tj messenger RNA and are sense-oriented. piRNA loading on to Piwi may occur in the cytoplasm. zucchini10, a gene encoding a putative cytoplasmic nuclease, is required for _tj_-derived piRNA production. In tj and piwi mutant ovaries, somatic cells fail to intermingle with germ cells and Fasciclin III is overexpressed. Loss of tj abolishes Piwi expression in gonadal somatic cells. Thus, in gonadal somatic cells, tj gives rise simultaneously to two different molecules: the TJ protein, which activates Piwi expression, and piRNAs, which define the Piwi targets for silencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Piwi in OSCs is associated with endogenous small RNAs.

The alternative text for this image may have been generated using AI.
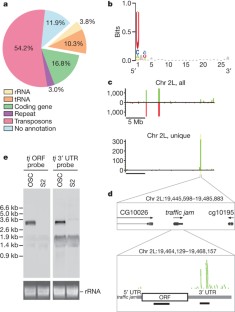
Figure 2: Piwi-associated piRNAs in OSCs.
The alternative text for this image may have been generated using AI.
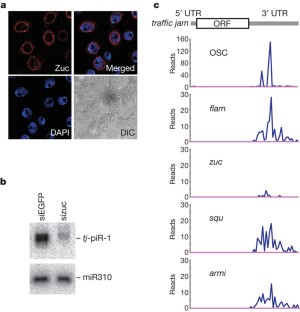
Figure 3: Involvement of zuc in the tj –piRNA production pathway.
The alternative text for this image may have been generated using AI.
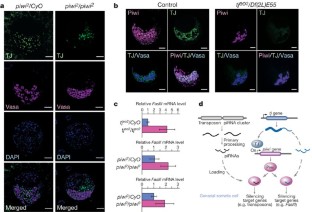
Figure 4: Phenotypes of tj and piwi mutant ovaries and testes.
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Small RNA sequences have been deposited at the GEO database under accession number GSE15137.
References
- Aravin, A. A., Hannon, G. J. & Brennecke, J. The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764 (2007)
Article CAS ADS Google Scholar - Siomi, H. & Siomi, M. C. On the road to reading the RNA-interference code. Nature 457, 396–404 (2009)
Article CAS ADS Google Scholar - Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007)
Article CAS Google Scholar - Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007)
Article CAS ADS Google Scholar - Li, C. et al. Collapse of germline piRNAs in the absent of Argonaute3 reveals somatic piRNAs in flies. Cell 137, 509–521 (2009)
Article CAS Google Scholar - Malone, C. et al. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell 137, 522–535 (2009)
Article CAS Google Scholar - Lau, N. et al. Abundant primary piRNAs, endo-siRNAs and microRNAs in a Drosophila ovary cell line. Genome Res. 10.1101/gr.094896.109 (14 July 2009)
- Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006)
Article CAS ADS Google Scholar - Li, M. A., Alls, J. D., Avancini, R. M., Koo, K. & Godt, D. The large Maf factor Traffic Jam controls gonad morphogenesis in Drosophila.. Nature Cell Biol. 5, 994–1000 (2003)
Article CAS Google Scholar - Pane, A., Wehr, K. & Schupbach, T. zucchini and squash encode two putative nucleases required for rasiRNA production in the Drosophila germilne. Dev. Cell 12, 851–862 (2007)
Article CAS Google Scholar - Cox, D. N. et al. A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal. Genes Dev. 12, 3715–3727 (1998)
Article CAS Google Scholar - Harris, A. N. & Macdonald, P. M. aubergine encodes a Drosophila polar granule component required for pole cell formation and related to eIF2C. Development 128, 2823–2832 (2001)
CAS PubMed Google Scholar - Niki, Y., Yamaguchi, T. & Mahowald, A. P. Establishment of stable cell lines of Drosophila germ-line stem cells. Proc. Natl Acad. Sci. USA 103, 16325–16330 (2006)
Article CAS ADS Google Scholar - Saito, K. et al. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes Dev. 20, 2214–2222 (2006)
Article CAS Google Scholar - Cox, D. N., Chao, A. & Lin, H. piwi encodes a nucleoplasmic factor whose activity modulates the number and division rate of germline stem cells. Genes Dev. 127, 503–514 (2000)
CAS Google Scholar - Nishida, M. K. et al. Gene silencing mechanisms mediated by Aubergine piRNA complexes in Drosophila male gonad. RNA 13, 1911–1922 (2007)
Article CAS Google Scholar - Saito, K. et al. Pimet, the Drosophila homolog of HEN1, mediates 2'-_O_-methylation of Piwi-interacting RNAs at their 3′ ends. Genes Dev. 21, 1603–1608 (2007)
Article CAS Google Scholar - Horwich, M. D. et al. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr. Biol. 17, 1265–1272 (2007)
Article CAS Google Scholar - Yin, H. & Lin, H. An epigenetic activation role of Piwi and a Piwi-associated piRNA in Drosophila melanogaster. Nature 450, 304–308 (2007)
Article CAS ADS Google Scholar - Czech, B. et al. An endogenous small interfering RNA pathway in Drosophila. Nature 453, 798–802 (2008)
Article CAS ADS Google Scholar - Arama, E., Dickman, D., Kimchie, Z., Shearn, A. & Lev, Z. Mutations in the beta-propeller domain of the Drosophila brain tumor (brat) protein induce neoplasm in the larval brain. Oncogene 19, 3706–3716 (2000)
Article CAS Google Scholar - Rogers, G. C. et al. Two mitotic kinesins cooperate to drive sister chromatid separation during anaphase. Nature 427, 364–370 (2004)
Article CAS ADS Google Scholar - Lin, H. & Spradling, A. C. A novel group of pumilio mutations affects the asymmetric division of germline stem cells in the Drosophila ovary. Development 124, 2463–2476 (1997)
CAS PubMed Google Scholar - Kataoka, K. Multiple mechanisms and functions of Maf transcriptional factors in the regulation of tissue-specific genes. J. Biochem. 141, 775–781 (2007)
Article CAS Google Scholar - Kawamura, Y. et al. Drosophila endogenous small RNAs bind to Argonaute 2 in somatic cells. Nature 453, 793–797 (2008)
Article CAS ADS Google Scholar - Kitadate, Y., Shigenobu, S., Arita, K. & Kobayashi, S. Boss/Sev signaling from germline to soma restricts germline-stem-cell-niche formation in the anterior region of Drosophila male gonads. Dev. Cell 13, 151–159 (2007)
Article CAS Google Scholar - Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M. C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005)
Article CAS Google Scholar - Wilson, R. J., Goodman, J. L., Strelets, V. B. & FlyBase Consortium FlyBase: integration and improvements to query tools. Nucleic Acids Res. 36, D588–D593 (2008)
Article CAS Google Scholar - Mituyama, T. et al. The Functional RNA Database 3.0: databases to support mining and annotation of functional RNAs. Nucleic Acids Res. 37, D89–D92 (2009)
Article CAS Google Scholar - Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo, a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Article CAS Google Scholar
Acknowledgements
We thank Y. Niki, D. Godt, H. Lin, E. Matunis, S. Kobayashi, Y. Kitadate, Y. Kageyama and H. Sano for providing reagents. We also thank Bloomington and Kyoto Drosophila Stock Center for the supply of Drosophila strains. We thank K. Yamada, E. Hattori, K. M. Nishida and T. N. Okada for technical assistance; S. Takahashi and K. Kataoka for discussions and suggestions; and other members of the Siomi laboratory for discussions and comments on the manuscript. We also thank K. Greer and D. McGowan for encouragement. This work was supported by MEXT grants to H.S. and NEDO (New Energy and Industrial Technology Development Organization) grants to M.C.S., T.M. and K.A. M.C.S. is supported by CREST from JST. M.C.S. is Associate Professor of Global COE for Human Metabolomics Systems Biology by MEXT.
Author Contributions K.S., S.I., Y.K. and H.K. conducted biochemical experiments. T.M., Y.O., E.S. and K.A. performed bioinformatics. K.S., T.M., H.S. and M.C.S designed experiments, interpreted data and prepared the manuscript.
Author information
Authors and Affiliations
- Keio University School of Medicine, Tokyo 160-8582, Japan
Kuniaki Saito, Sachi Inagaki, Yoshinori Kawamura, Hazuki Kotani, Haruhiko Siomi & Mikiko C. Siomi - Computational Biology Research Center (CBRC), National Institute of Advanced Industrial Science and Technology (AIST), Tokyo 135-0064, Japan
Toutai Mituyama & Kiyoshi Asai - Institute of Health Biosciences University of Tokushima, Tokushima 770-8503, Japan
Yoshinori Kawamura - Department of Life Sciences, Information and Mathematical Science Laboratory, Inc., Tokyo 112-0012, Japan
Yukiteru Ono - Japan Biological Informatics Consortium (JBIC), Tokyo 135-8073, Japan
Eri Sakota - Graduate School of Frontier Sciences, University of Tokyo, Chiba 277-8583, Japan
Kiyoshi Asai - JST, CREST, Saitama 332-0012, Japan
Mikiko C. Siomi
Authors
- Kuniaki Saito
- Sachi Inagaki
- Toutai Mituyama
- Yoshinori Kawamura
- Yukiteru Ono
- Eri Sakota
- Hazuki Kotani
- Kiyoshi Asai
- Haruhiko Siomi
- Mikiko C. Siomi
Corresponding authors
Correspondence toHaruhiko Siomi or Mikiko C. Siomi.
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Saito, K., Inagaki, S., Mituyama, T. et al. A regulatory circuit for piwi by the large Maf gene traffic jam in Drosophila.Nature 461, 1296–1299 (2009). https://doi.org/10.1038/nature08501
- Received: 03 August 2009
- Accepted: 16 September 2009
- Published: 07 October 2009
- Issue date: 29 October 2009
- DOI: https://doi.org/10.1038/nature08501
This article is cited by
Editorial Summary
A regulatory circuit for Piwi
A class of small RNAs, piRNAs, interact with PIWI domain-containing Argonaute proteins. Germline piRNAs derive from repetitive DNA and are produced by an amplification process. Recently piRNAs have also been found to be encoded by protein-coding genes in somatic cells. In this work, Siomi and colleagues show that the traffic jam (tj) locus, encoding the Maf transcription factor, encodes a piRNA cluster. These piRNAs are produced in the cytoplasm without amplification, and target FasIII for silencing. Maf itself activates Piwi expression. Thus, the tj locus encodes both an enhancer of Piwi and the piRNAs that interact with Piwi.