Synthetic analog computation in living cells (original) (raw)
- Letter
- Published: 15 May 2013
- Jacob R. Rubens2,3,4,
- Rahul Sarpeshkar1,3,4,5,6,7 na1 &
- …
- Timothy K. Lu2,3,4,5,6,8 na1
Nature volume 497, pages 619–623 (2013)Cite this article
- 36k Accesses
- 436 Citations
- 176 Altmetric
- Metrics details
Subjects
Abstract
A central goal of synthetic biology is to achieve multi-signal integration and processing in living cells for diagnostic, therapeutic and biotechnology applications1. Digital logic has been used to build small-scale circuits, but other frameworks may be needed for efficient computation in the resource-limited environments of cells2,3. Here we demonstrate that synthetic analog gene circuits can be engineered to execute sophisticated computational functions in living cells using just three transcription factors. Such synthetic analog gene circuits exploit feedback to implement logarithmically linear sensing, addition, ratiometric and power-law computations. The circuits exhibit Weber’s law behaviour as in natural biological systems4, operate over a wide dynamic range of up to four orders of magnitude and can be designed to have tunable transfer functions. Our circuits can be composed to implement higher-order functions that are well described by both intricate biochemical models and simple mathematical functions. By exploiting analog building-block functions that are already naturally present in cells3,5, this approach efficiently implements arithmetic operations and complex functions in the logarithmic domain. Such circuits may lead to new applications for synthetic biology and biotechnology that require complex computations with limited parts, need wide-dynamic-range biosensing or would benefit from the fine control of gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Positive-feedback linearization of gene circuits for wide-dynamic-range analog computation.
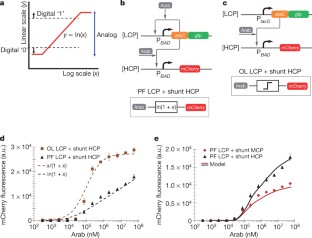
The alternative text for this image may have been generated using AI.
Figure 2: Analog behaviour versus digital.
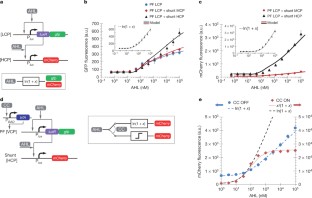
The alternative text for this image may have been generated using AI.
Figure 3: Log-domain analog computation.
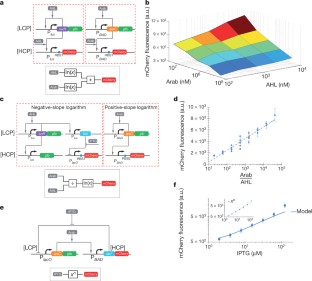
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Chen, Y. Y., Galloway, K. E. & Smolke, C. D. Synthetic biology: advancing biological frontiers by building synthetic systems. Genome Biol. 13, 240 (2012)
Article Google Scholar - Cardinale, S. & Arkin, A. P. Contextualizing context for synthetic biology: identifying causes of failure of synthetic biological systems. Biotechnol. J. 7, 856–866 (2012)
Article CAS Google Scholar - Sarpeshkar, R. Analog versus digital: extrapolating from electronics to neurobiology. Neural Comput. 10, 1601–1638 (1998)
Article CAS Google Scholar - Ferrell, J. E. Signaling motifs and Weber’s law. Mol. Cell 36, 724–727 (2009)
Article CAS Google Scholar - Sarpeshkar, R. Ultra Low Power Bioelectronics: Fundamentals, Biomedical Applications, and Bio-Inspired Systems 651–694, 753–786 (Cambridge Univ. Press, 2010)
Book Google Scholar - Sprinzak, D. et al. _Cis_-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010)
Article ADS CAS Google Scholar - Canton, B., Labno, A. & Endy, D. Refinement and standardization of synthetic biological parts and devices. Nature Biotechnol. 26, 787–793 (2008)
Article CAS Google Scholar - Giorgetti, L. et al. Noncooperative interactions between transcription factors and clustered DNA binding sites enable graded transcriptional responses to environmental inputs. Mol. Cell 37, 418–428 (2010)
Article CAS Google Scholar - Clark, B. & Hausser, M. Neural coding: hybrid analog and digital signalling in axons. Curr. Biol. 16, R585–R588 (2006)
Article CAS Google Scholar - Daniel, R., Woo, S. S., Turicchia, L. & Sarpeshkar, R. in Biomedical Circuits and Systems Conference (BioCAS 2011) 333–336 (IEEE, 2011)
Book Google Scholar - Tavakoli, M. & Sarpeshkar, R. A sinh resistor and its application to tanh linearization. IEEE J. Solid-State Circuits 40, 536–543 (2005)
Article ADS Google Scholar - Wild, J., Hradecna, Z. & Szybalski, W. Conditionally amplifiable BACs: switching from single-copy to high-copy vectors and genomic clones. Genome Res. 12, 1434–1444 (2002)
Article CAS Google Scholar - Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011)
Article ADS CAS Google Scholar - Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008)
Article ADS CAS Google Scholar - Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000)
ADS CAS PubMed Google Scholar - McMillen, D., Kopell, N., Hasty, J. & Collins, J. J. Synchronizing genetic relaxation oscillators by intercell signaling. Proc. Natl Acad. Sci. USA 99, 679–684 (2002)
Article ADS CAS Google Scholar - Madar, D., Dekel, E., Bren, A. & Alon, U. Negative auto-regulation increases the input dynamic-range of the arabinose system of Escherichia coli. BMC Syst. Biol. 5, 111 (2011)
Article Google Scholar - Nevozhay, D., Adams, R. M., Murphy, K. F., Josić, K. & Balázsi, G. Negative autoregulation linearizes the dose–response and suppresses the heterogeneity of gene expression. Proc. Natl Acad. Sci. USA 106, 5123–5128 (2009)
Article ADS CAS Google Scholar - Shen-Orr, S. S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nature Genet. 31, 64–68 (2002)
Article CAS Google Scholar - You, L., Cox, R. S., Weiss, R. & Arnold, F. H. Programmed population control by cell–cell communication and regulated killing. Nature 428, 868–871 (2004)
Article ADS CAS Google Scholar - Tabor, J. J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009)
Article Google Scholar - Tamsir, A., Tabor, J. J. & Voigt, C. A. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. Nature 469, 212–215 (2011)
Article ADS CAS Google Scholar - Auslander, S., Auslander, D., Muller, M., Wieland, M. & Fussenegger, M. Programmable single-cell mammalian biocomputers. Nature 487, 123–127 (2012)
Article ADS Google Scholar - Isaacs, F. J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nature Biotechnol. 22, 841–847 (2004)
Article CAS Google Scholar - Win, M. N. & Smolke, C. D. Higher-order cellular information processing with synthetic RNA devices. Science 322, 456–460 (2008)
Article ADS CAS Google Scholar - Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307–1311 (2011)
Article ADS CAS Google Scholar - Khalil, A. et al. A synthetic biology framework for programming eukaryotic transcription functions. Cell 150, 647–658 (2012)
Article CAS Google Scholar - Dueber, J. E., Yeh, B. J., Chak, K. & Lim, W. A. Reprogramming control of an allosteric signaling switch through modular recombination. Science 301, 1904–1908 (2003)
Article ADS CAS Google Scholar - Hahnloser, R. H. R., Sarpeshkar, R., Mahowald, M. A., Douglas, R. J. & Seung, H. S. Digital selection and analogue amplification coexist in a cortex-inspired silicon circuit. Nature 405, 947–951 (2000)
Article ADS CAS Google Scholar - Lu, T. K., Khalil, A. S. & Collins, J. J. Next-generation synthetic gene networks. Nature Biotechnol. 27, 1139–1150 (2009)
Article CAS Google Scholar
Acknowledgements
We would like to thank J. Nungesser for assistance with figures and members of the Lu and Sarpeshkar laboratories for discussions. This work was supported in part by a campus collaboration initiative from Lincoln Labs (R.D. and R.S.), the US National Science Foundation (R.D., J.R.R., R.S. and T.K.L.) under grant number 1124247, and the Office of Naval Research (J.R.R. and T.K.L.) under grant number N000141110725.
Author information
Author notes
- Rahul Sarpeshkar and Timothy K. Lu: These authors contributed equally to this work.
Authors and Affiliations
- Analog Circuits and Biological Systems Group, Research Lab of Electronics, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Ramiz Daniel & Rahul Sarpeshkar - Synthetic Biology Group, Research Lab of Electronics, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Ramiz Daniel, Jacob R. Rubens & Timothy K. Lu - Synthetic Biology Center, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Ramiz Daniel, Jacob R. Rubens, Rahul Sarpeshkar & Timothy K. Lu - MIT Microbiology Program, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Jacob R. Rubens, Rahul Sarpeshkar & Timothy K. Lu - Department of Electrical Engineering & Computer Science, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Rahul Sarpeshkar & Timothy K. Lu - MIT Computational and Systems Biology Program, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Rahul Sarpeshkar & Timothy K. Lu - MIT Biophysics Program, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Rahul Sarpeshkar - Department of Biological Engineering, Massachusetts Institute of Technology, Cambridge, 02139, Massachusetts, USA
Timothy K. Lu
Authors
- Ramiz Daniel
- Jacob R. Rubens
- Rahul Sarpeshkar
- Timothy K. Lu
Contributions
R.D., R.S. and T.K.L. designed the study. R.D. and J.R.R. performed experiments and collected data. R.D., J.R.R., R.S. and T.K.L. invented the analog circuit motifs. R.D., R.S. and T.K.L. developed the analog circuit motifs and associated models and simulations. All authors analysed the data, discussed results and wrote the manuscript.
Corresponding authors
Correspondence toRahul Sarpeshkar or Timothy K. Lu.
Ethics declarations
Competing interests
Massachusetts Institute of Technology, with which all the authors are affiliated, has filed a PCT patent application based on this work.
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary Text and Data, Supplementary Figures 1-53, Supplementary Tables 1-4, and Supplementary References (see Table of Contents for more details). (PDF 6125 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Daniel, R., Rubens, J., Sarpeshkar, R. et al. Synthetic analog computation in living cells.Nature 497, 619–623 (2013). https://doi.org/10.1038/nature12148
- Received: 10 October 2012
- Accepted: 03 April 2013
- Published: 15 May 2013
- Issue date: 30 May 2013
- DOI: https://doi.org/10.1038/nature12148
This article is cited by
Editorial Summary
Living-cell computation simplified
The design of novel genetic control systems for synthetic biology is dominated by digital logic. This is necessarily a complex arrangement. Now Timothy Lu and colleagues have harnessed analog building-blocks found in natural cells to perform arithmetic operations in the logarithmic domain. Such analog circuits — which could be integrated with digital — should make it possible to use fewer components to implement complex computations that require wide dynamic range in biosensing.