Addicting diverse bacteria to a noncanonical amino acid (original) (raw)
- Brief Communication
- Published: 18 January 2016
- Jared W Ellefson1,
- Ross Thyer1,
- Bo Wang1,
- Jimmy Gollihar1,
- Matthew T Forster1 &
- …
- Andrew D Ellington1
Nature Chemical Biology volume 12, pages 138–140 (2016)Cite this article
- 6757 Accesses
- 67 Citations
- 29 Altmetric
- Metrics details
Subjects
Abstract
Engineered orthogonal translation systems have greatly enabled the expansion of the genetic code using noncanonical amino acids (NCAAs). However, the impact of NCAAs on organismal evolution remains unclear, in part because it is difficult to force the adoption of new genetic codes in organisms. By reengineering TEM-1 β-lactamase to be dependent on a NCAA, we maintained bacterial NCAA dependence for hundreds of generations without escape.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Characterization of NCAA dependent β-lactamase variants.
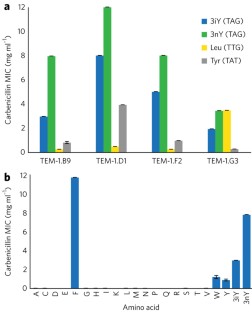
The alternative text for this image may have been generated using AI.
Figure 2: β-lactamase variant TEM-1.B9 maintained NCAA dependence in different bacterial species during serial culture.

The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Primary accessions
NCBI Reference Sequence
Protein Data Bank
References
- Wong, J.T. Proc. Natl. Acad. Sci. USA 80, 6303–6306 (1983).
Article CAS Google Scholar - Rovner, A.J. et al. Nature 518, 89–93 (2015).
Article CAS Google Scholar - Mandell, D.J. et al. Nature 518, 55–60 (2015).
Article CAS Google Scholar - Lemeignan, B., Sonigo, P. & Marlière, P. J. Mol. Biol. 231, 161–166 (1993).
Article CAS Google Scholar - Chin, J.W. et al. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Article CAS Google Scholar - Wang, J., Xie, J. & Schultz, P.G. J. Am. Chem. Soc. 128, 8738–8739 (2006).
Article CAS Google Scholar - Xie, J., Supekova, L. & Schultz, P.G. ACS Chem. Biol. 2, 474–478 (2007).
Article CAS Google Scholar - Liu, C.C. & Schultz, P.G. Annu. Rev. Biochem. 79, 413–444 (2010).
Article CAS Google Scholar - Hammerling, M.J. et al. Nat. Chem. Biol. 10, 178–180 (2014).
Article CAS Google Scholar - Bacher, J.M., Bull, J.J. & Ellington, A.D. BMC Evol. Biol. 3, 24 (2003).
Article Google Scholar - Bacher, J.M. & Ellington, A.D. J. Bacteriol. 183, 5414–5425 (2001).
Article CAS Google Scholar - Wang, Q. et al. ChemBioChem 15, 1744–1749 (2014).
Article CAS Google Scholar - Kato, Y. PeerJ 3, e1247 (2015).
Article Google Scholar - Cooley, R.B. et al. Biochemistry 53, 1916–1924 (2014).
Article CAS Google Scholar - Sakamoto, K. et al. Structure 17, 335–344 (2009).
Article CAS Google Scholar - Ohtake, K. et al. Sci. Rep. 5, 9762 (2015).
Article CAS Google Scholar - Bradford, P.A. Clin. Microbiol. Rev. 14, 933–951 (2001).
Article CAS Google Scholar - Fonzé, E. et al. Acta Crystallogr. D Biol. Crystallogr. 51, 682–694 (1995).
Article Google Scholar - Baba, T. et al. Mol. Syst. Biol. 2, 2006.0008 (2006).
Article Google Scholar - Lajoie, M.J. et al. Science 342, 357–360 (2013).
Article CAS Google Scholar - Thyer, R., Robotham, S.A., Brodbelt, J.S. & Ellington, A.D. J. Am. Chem. Soc. 137, 46–49 (2015).
Article CAS Google Scholar - Gibson, D.G. et al. Nat. Methods 6, 343–345 (2009).
Article CAS Google Scholar - Hashimoto-Gotoh, T. et al. Gene 241, 185–191 (2000).
Article CAS Google Scholar - Chung, C.T., Niemela, S.L. & Miller, R.H. Proc. Natl. Acad. Sci. USA 86, 2172–2175 (1989).
Article CAS Google Scholar - Stec, B., Holtz, K.M., Wojciechowski, C.L. & Kantrowitz, E.R. Acta Crystallogr. D Biol. Crystallogr. 61, 1072–1079 (2005).
Article Google Scholar - Labute, P. Proteins 75, 187–205 (2009).
Article CAS Google Scholar - Labute, P. J. Chem. Inf. Model. 50, 792–800 (2010).
Article CAS Google Scholar - Clark, A.M. & Labute, P. J. Chem. Inf. Model. 47, 1933–1944 (2007).
Article CAS Google Scholar
Acknowledgements
This work was supported by the following grants: National Security Science and Engineering Faculty Fellowship grant FA9550-10-1-0169 to A.D.E. Welch Foundation grant F-1654. Defense Advanced Research Projects Agency N66001-14-2-4051 to A.D.E. Air Force Office of Scientific Research grant FA9550-14-1-0089 to A.D.E. Defense Advanced Research Projects Agency HR0011-15-C0095 to A.D.E.
Author information
Authors and Affiliations
- Center for Systems and Synthetic Biology, University of Texas at Austin, Austin, Texas, USA
Drew S Tack, Jared W Ellefson, Ross Thyer, Bo Wang, Jimmy Gollihar, Matthew T Forster & Andrew D Ellington
Authors
- Drew S Tack
- Jared W Ellefson
- Ross Thyer
- Bo Wang
- Jimmy Gollihar
- Matthew T Forster
- Andrew D Ellington
Contributions
D.S.T. designed and performed experiments and wrote the manuscript. J.W.E. designed experiments and wrote the manuscript. R.T. designed experiments and wrote the manuscript. B.W. screened TEM-1 variants. M.T.F. screened TEM-1 variants. J.G. performed computational analysis using the Molecular Operating Environment. A.D.E. directed experimental work and wrote the manuscript.
Corresponding author
Correspondence toAndrew D Ellington.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Source data
Rights and permissions
About this article
Cite this article
Tack, D., Ellefson, J., Thyer, R. et al. Addicting diverse bacteria to a noncanonical amino acid.Nat Chem Biol 12, 138–140 (2016). https://doi.org/10.1038/nchembio.2002
- Received: 12 June 2015
- Accepted: 13 November 2015
- Published: 18 January 2016
- Issue date: March 2016
- DOI: https://doi.org/10.1038/nchembio.2002