Real ribozymes suggest a relaxed error threshold (original) (raw)
- Letter
- Published: 28 August 2005
Nature Genetics volume 37, pages 1008–1011 (2005)Cite this article
- 1286 Accesses
- 137 Citations
- 6 Altmetric
- Metrics details
Abstract
The error threshold for replication, the critical copying fidelity below which the fittest genotype deterministically disappears, limits the length of the genome that can be maintained by selection. Primordial replication must have been error-prone, and so early replicators are thought to have been necessarily short1. The error threshold also depends on the fitness landscape. In an RNA world2, many neutral and compensatory mutations can raise the threshold, below which the functional phenotype3, rather than a particular sequence, is still present4,5. Here we show, on the basis of comparative analysis of two extensively mutagenized ribozymes, that with a copying fidelity of 0.999 per digit per replication the phenotypic error threshold rises well above 7,000 nucleotides, which permits the selective maintenance of a functionally rich riboorganism6 with a genome of more than 100 different genes, the size of a tRNA. This requires an order of magnitude of improvement in the accuracy of _in vitro_–generated polymerase ribozymes7,8. Incidentally, this genome size coincides with that estimated for a minimal cell achieved by top-down analysis9, omitting the genes dealing with translation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Mutagenized ribozymes.
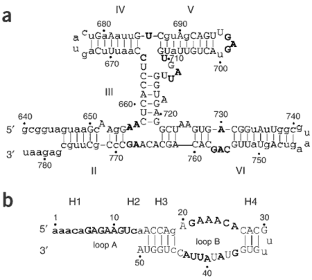
The alternative text for this image may have been generated using AI.
Figure 2: Time to extinction in generations as a function of the per digit effective mutation rate (μ*) in a population of constant size with N = 10,000 molecules for the Neurospora VS ribozyme (a,c,e,g) and the hairpin ribozyme (b,d,f,h).
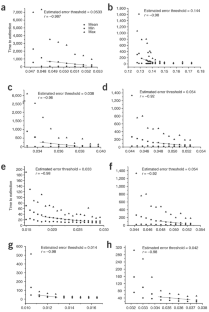
The alternative text for this image may have been generated using AI.
Figure 3: Fraction of mutants with full activity (filled circles) or any activity (open squares), as a function of the number of point mutations.
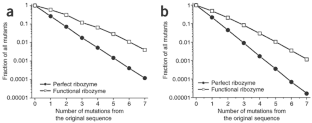
The alternative text for this image may have been generated using AI.
Figure 4: Relationship between the per digit replication accuracy (q) and the permissible genome size (L) estimated from equation 2 with λ = 0.22 and s = 351.
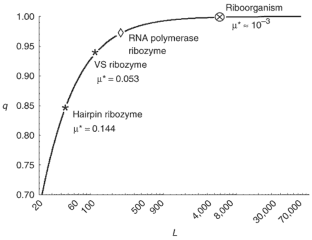
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Eigen, M. Self organization of matter and the evolution of biological macromolecules. Naturwissenschaften 10, 465–523 (1971).
Article Google Scholar - Gilbert, W. The RNA world. Nature 319, 618 (1986).
Article Google Scholar - Huynen, M.A., Stadler, P.F. & Fontana, W. Smoothness within ruggedness: the role of neutrality in adaptation. Proc. Natl. Acad. Sci. USA 93, 397–401 (1996).
Article CAS Google Scholar - Reidys, C., Forst, C.V. & Schuster, P. Replication and mutation on neutral networks. Bull. Math. Biol. 63, 57–94 (2001).
Article CAS Google Scholar - Takeuchi, N., Poorthuis, P.H. & Hogeweg, P. Phenotypic error threshold; additivity and epistasis in RNA evolution. BMC Evol. Biol. 5, 9 (2005).
Article Google Scholar - Jeffares, D.C., Poole, A.M. & Penny, D. Relics from the RNA world. J. Mol. Evol. 46, 18–36 (1998).
Article CAS Google Scholar - Johnston, W.K., Unrau, P.J., Lawrence, M.S., Glasen, M.E. & Bartel, D.P. RNA-catalyzed RNA polymerization: accurate and general RNA-templated primer extension. Science 292, 1319–1325 (2001).
Article CAS Google Scholar - Müller, U.F. & Bartel, D.P. Substrate 2′-hydroxyl groups required for ribozyme-catalyzed polymerization. Chem. Biol. 10, 799–806 (2003).
Article Google Scholar - Gil, R., Silva, F.J., Peretó, J. & Moya, A. Determination of the core of a minimal bacterial gene set. Microbiol. Mol. Biol. Rev. 68, 518–537 (2004).
Article CAS Google Scholar - Maynard Smith, J. Hypercycles and the origin of life. Nature 20, 445–446 (1979).
Article Google Scholar - Maynard Smith, J. Models of evolution. Proc. R. Soc. Lond. B 219, 315–325 (1983).
Article Google Scholar - Maynard Smith, J. & Szathmáry, E. The Major Transitions in Evolution (Oxford Univ. Press, Oxford, 1995).
Google Scholar - Benner, B., Ellington, A.D., Ge, L., Gasfeld, A. & Leanz, G.F. Natural selection, protein enzgineering, and the last riboorganism: rational model building in biochemistry. Cold Spring Harb. Symp. Quant. Biol. 52, 56–63 (1987).
Article Google Scholar - Fedor, M. Structure and function of the hairpin ribozyme. J. Mol. Biol. 297, 269–291 (2000).
Article CAS Google Scholar - Lafontaine, D.A., Norman, D.G. & Lilley, D.M.J. The structure and active site of the Varkund satellite ribozyme. Biochem. Soc. Trans. 30, 1170–1175 (2002).
Article CAS Google Scholar - Lehman, N. & Joyce, G.F. Evolution in vitro: analysis of a lineage of ribozymes. Curr. Biol. 3, 723–734 (1993).
Article CAS Google Scholar - Takeda, Y., Sarai, A. & Rivera, V.M. Analysis of the sequence-specific interactions between Cro repressor and operator DNA by systematic base substitution experiments. Proc. Natl. Acad. Sci. USA 86, 439–443 (1989).
Article CAS Google Scholar - Sandberg, W.S. & Terwilliger, T.C. Engineering multiple properties of a protein by combinatorial mutagenesis. Proc. Natl. Acad. Sci. USA 90, 8367–8371 (1993).
Article CAS Google Scholar - Skinner, M.M. & Terwilliger, T.C. Potential use of additivity of mutational effects in simplifying protein engineering. Proc. Natl. Acad. Sci. USA 93, 10753–10757 (1996).
Article CAS Google Scholar - Eigen, M., McCaskill, J.S. & Schuster, P. The molecular quasispecies. Adv. Chem. Phys. 75, 149–263 (1989).
CAS Google Scholar - Santos, M., Zintzaras, E. & Szathmáry, E. Recombination in primeval genomes: a step forward but still a long leap from maintaining a sizeable genome. J. Mol. Evol. 59, 507–519 (2004).
Article CAS Google Scholar - Domingo, E. & Holland, J.J. in The Evolutionary Biology of Viruses (ed. Morse, S.S.) 161–183 (Raven, New York, 1994).
Google Scholar - Hofacker, I.L. et al. Fast folding and comparison of RNA secondary structures. Monatsh. Chem. 125, 167–188 (1994).
Article CAS Google Scholar - Hofacker, I.L. Vienna RNA secondary structure server. Nucleic Acids Res. 31, 3429–3431 (2003).
Article CAS Google Scholar - Beattie, T.L., Olive, J.E. & Collins, R.A. A secondary-structure model for the self-cleaving region of the Neurospora VS RNA. Proc. Natl. Acad. Sci. USA 92, 4686–4690 (1995).
Article CAS Google Scholar - Butcher, S.E. & Burke, J.M. Structure-mapping of the hairpin ribozyme. Magnesium-dependent folding and evidence for tertiary interactions within the ribozyme-substrate complex. J. Mol. Biol. 244, 52–63 (1994).
Article CAS Google Scholar - Butcher, S.E. & Burke, J.M. A photo-cross-linkable tertiary structure motif found in functionally distinct RNA molecules is essential for catalytic function of the hairpin ribozyme. Biochemistry 33, 992–999 (1994).
Article CAS Google Scholar
Acknowledgements
We thank I. Hofacker for his help with the Vienna RNA package, P. Mezey for allowing us to use the computer facilities at MUN (Canada) and F. Kondrashov for comments on an earlier version of the manuscript. Computer facilities were provided by Microdigit. This work was supported by grant and postdoctoral fellowship from the Hungarian National Research Fund (OTKA) to Á.K. M.S. is partially supported by Fundación Ramón Areces (Spain). This work was also supported by the COST D27 action (Prebiotic chemistry and early evolution).
Author information
Authors and Affiliations
- Collegium Budapest, Institute for Advanced Study, Szentháromság u. 2., Budapest, H-1014, Hungary
Ádám Kun & Eörs Szathmáry - Department of Plant Taxonomy and Ecology, Eötvös University, Pázmány Péter sétány 1/C, Budapest, H-1117, Hungary
Ádám Kun & Eörs Szathmáry - Departament de Genètica i de Microbiologia, Grup de Biologia Evolutiva, Universitat Autònoma de Barcelona, 08193 Bellaterra, Barcelona, Spain
Mauro Santos - Research Group of Ecology and Theoretical Biology, Eötvös University, Hungarian Academy of Science, Pázmány Péter sétány 1/c, Budapest, H-1117, Hungary
Eörs Szathmáry
Authors
- Ádám Kun
- Mauro Santos
- Eörs Szathmáry
Corresponding author
Correspondence toEörs Szathmáry.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Kun, Á., Santos, M. & Szathmáry, E. Real ribozymes suggest a relaxed error threshold.Nat Genet 37, 1008–1011 (2005). https://doi.org/10.1038/ng1621
- Received: 03 May 2005
- Accepted: 07 July 2005
- Published: 28 August 2005
- Issue date: 01 September 2005
- DOI: https://doi.org/10.1038/ng1621
This article is cited by
Ribozyme Mutagenic Evolution: Mechanisms of Survival
- Carolina Diaz Arenas
- Aleksandra Ardaševa
- Yohei Yokobayashi
Origins of Life and Evolution of Biospheres (2021)
Negative Epistasis in Experimental RNA Fitness Landscapes
- Devin P. Bendixsen
- Bjørn Østman
- Eric J. Hayden
Journal of Molecular Evolution (2017)
Cooperation and selfishness both occur during molecular evolution
- David Penny
Biology Direct (2015)
- David Penny
Quasispecies in population of compositional assemblies
- Renan Gross
- Itzhak Fouxon
- Omer Markovitch
BMC Evolutionary Biology (2014)
The Case for an Early Biological Origin of DNA
- Anthony M. Poole
- Nobuyuki Horinouchi
- Jun Ogawa
Journal of Molecular Evolution (2014)