DeNovoGear: de novo indel and point mutation discovery and phasing (original) (raw)
- Brief Communication
- Published: 25 August 2013
- Michiel J Noordam1 na1,
- Rachel S Schwartz2,
- Arthur Wuster3,
- Matthew E Hurles3,
- Reed A Cartwright2,4 &
- …
- Donald F Conrad1,5
Nature Methods volume 10, pages 985–987 (2013) Cite this article
- 9464 Accesses
- 176 Citations
- 83 Altmetric
- Metrics details
Subjects
Abstract
We present DeNovoGear software for analyzing de novo mutations from familial and somatic tissue sequencing data. DeNovoGear uses likelihood-based error modeling to reduce the false positive rate of mutation discovery in exome analysis and fragment information to identify the parental origin of germ-line mutations. We used DeNovoGear on human whole-genome sequencing data to produce a set of predicted de novo insertion and/or deletion (indel) mutations with a 95% validation rate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Using beta-binomial likelihoods to model exome data.
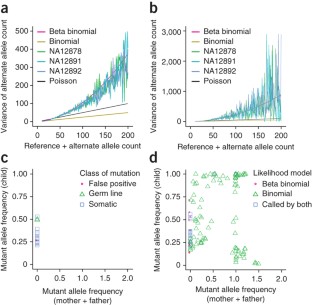
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Accessions
Sequence Read Archive
References
- Conrad, D.F. et al. Nat. Genet. 43, 712–714 (2011).
Article CAS Google Scholar - Roach, J.C. et al. Science 328, 636–639 (2010).
Article CAS Google Scholar - Kong, A. et al. Nature 488, 471–475 (2012).
Article CAS Google Scholar - Cartwright, R.A., Hussin, J., Keebler, J.E., Stone, E.A. & Awadalla, P. Stat. Appl. Genet. Mol. Biol. 11, pii (2012).
Article Google Scholar - Abecasis, G.R. et al. Nature 491, 56–65 (2012).
Article Google Scholar - Heinrich, V. et al. Nucleic Acids Res. 40, 2426–2431 (2012).
Article CAS Google Scholar - DePristo, M.A. et al. Nat. Genet. 43, 491–498 (2011).
Article CAS Google Scholar - Li, H. Bioinformatics 27, 2987–2993 (2011).
Article CAS Google Scholar - Li, B. et al. PLoS Genet. 8, e1002944 (2012).
Article CAS Google Scholar - Albers, C.A. et al. Genome Res. 21, 961–973 (2011).
Article CAS Google Scholar - Lynch, M. Proc. Natl. Acad. Sci. USA 107, 961–968 (2010).
Article CAS Google Scholar - Lunter, G. Bioinformatics 23, i289–i296 (2007).
Article CAS Google Scholar - Lynch, M. et al. Proc. Natl. Acad. Sci. USA 105, 9272–9277 (2008).
Article CAS Google Scholar - Kvikstad, E.M., Tyekucheva, S., Chiaromonte, F. & Makova, K.D. PLoS Comput. Biol. 3, 1772–1782 (2007).
Article CAS Google Scholar - Benson, G. Nucleic Acids Res. 27, 573–580 (1999).
Article CAS Google Scholar - Smith, D.M. Appl. Stat. 32, 196–204 (1983).
Article Google Scholar - Watterson, G.A. Theor. Popul. Biol. 7, 256–276 (1975).
Article CAS Google Scholar - Conrad, D. et al. Nature 464, 704–712 (2010).
Article CAS Google Scholar
Acknowledgements
We thank R. Hardwick for assistance with primer design, V. Plagnol and H. Li for helpful discussion, and members of the 1000 Genomes community for generating software, data and resources that we used as part of this project. This research was supported in part by Wellcome Trust grant WT098051.
Author information
Author notes
- Avinash Ramu and Michiel J Noordam: These authors contributed equally to this work.
Authors and Affiliations
- Department of Genetics, Washington University School of Medicine, St. Louis, Missouri, USA
Avinash Ramu, Michiel J Noordam & Donald F Conrad - Center for Evolutionary Medicine and Informatics, The Biodesign Institute, Arizona State University, Tempe, Arizona, USA
Rachel S Schwartz & Reed A Cartwright - Genome Mutation and Genetic Disease Group, Wellcome Trust Sanger Institute, Cambridge, UK
Arthur Wuster & Matthew E Hurles - School of Life Sciences, Arizona State University, Tempe, Arizona, USA
Reed A Cartwright - Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA
Donald F Conrad
Authors
- Avinash Ramu
- Michiel J Noordam
- Rachel S Schwartz
- Arthur Wuster
- Matthew E Hurles
- Reed A Cartwright
- Donald F Conrad
Contributions
A.R. implemented methods, analyzed data and wrote the paper; M.J.N. performed validation experiments, analyzed data and wrote the paper; R.S.S. performed simulations; A.W. provided code and performed early analysis demonstrating the utility of beta-binomials; M.E.H. and R.A.C. gave conceptual advice, supervised the project and wrote the paper; D.F.C. designed and supervised the project, implemented methods, analyzed data and wrote the paper.
Corresponding author
Correspondence toDonald F Conrad.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Source data
Rights and permissions
About this article
Cite this article
Ramu, A., Noordam, M., Schwartz, R. et al. DeNovoGear: de novo indel and point mutation discovery and phasing.Nat Methods 10, 985–987 (2013). https://doi.org/10.1038/nmeth.2611
- Received: 15 February 2013
- Accepted: 24 July 2013
- Published: 25 August 2013
- Issue date: October 2013
- DOI: https://doi.org/10.1038/nmeth.2611