A synthetic homing endonuclease-based gene drive system in the human malaria mosquito (original) (raw)
- Letter
- Published: 20 April 2011
- Miriam Menichelli1,
- Philippos Aris Papathanos1,
- Summer B. Thyme2,3,
- Hui Li4,
- Umut Y. Ulge4,5,
- Blake T. Hovde6,
- David Baker2,3,7,
- Raymond J. Monnat4,5,6,
- Austin Burt1,8 na1 &
- …
- Andrea Crisanti1,9 na1
Nature volume 473, pages 212–215 (2011)Cite this article
- 11k Accesses
- 335 Citations
- 136 Altmetric
- Metrics details
Subjects
Abstract
Genetic methods of manipulating or eradicating disease vector populations have long been discussed as an attractive alternative to existing control measures because of their potential advantages in terms of effectiveness and species specificity1,2,3. The development of genetically engineered malaria-resistant mosquitoes has shown, as a proof of principle, the possibility of targeting the mosquito’s ability to serve as a disease vector4,5,6,7. The translation of these achievements into control measures requires an effective technology to spread a genetic modification from laboratory mosquitoes to field populations8. We have suggested previously that homing endonuclease genes (HEGs), a class of simple selfish genetic elements, could be exploited for this purpose9. Here we demonstrate that a synthetic genetic element, consisting of mosquito regulatory regions10 and the homing endonuclease gene I-SceI11,12,13, can substantially increase its transmission to the progeny in transgenic mosquitoes of the human malaria vector Anopheles gambiae. We show that the I-SceI element is able to invade receptive mosquito cage populations rapidly, validating mathematical models for the transmission dynamics of HEGs. Molecular analyses confirm that expression of I-SceI in the male germline induces high rates of site-specific chromosomal cleavage and gene conversion, which results in the gain of the I-SceI gene, and underlies the observed genetic drive. These findings demonstrate a new mechanism by which genetic control measures can be implemented. Our results also show in principle how sequence-specific genetic drive elements like HEGs could be used to take the step from the genetic engineering of individuals to the genetic engineering of populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Analysis of HEG activity in transgenic mosquitoes.
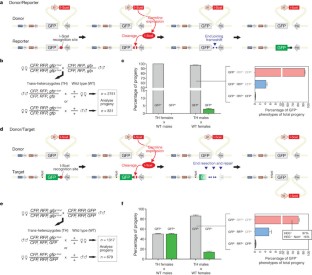
The alternative text for this image may have been generated using AI.
Figure 2: HEG invasion in mosquito cage populations.

The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
The plasmids pHome-T and pHome-D have been deposited to GenBank under the accession numbers HQ159398 and HQ159399.
References
- Curtis, C. F. Possible use of translocations to fix desirable genes in insect pest populations. Nature 218, 368–369 (1968)
Article ADS CAS Google Scholar - Hamilton, W. D. Extraordinary sex ratios. A sex-ratio theory for sex linkage and inbreeding has new implications in cytogenetics and entomology. Science 156, 477–488 (1967)
Article ADS CAS Google Scholar - Alphey, L. et al. Malaria control with genetically manipulated insect vectors. Science 298, 119–121 (2002)
Article ADS CAS Google Scholar - Corby-Harris, V. et al. Activation of Akt signaling reduces the prevalence and intensity of malaria parasite infection and lifespan in Anopheles stephensi mosquitoes. PLoS Pathog. 6, e1001003 (2010)
Article Google Scholar - Ito, J., Ghosh, A., Moreira, L. A., Wimmer, E. A. & Jacobs-Lorena, M. Transgenic anopheline mosquitoes impaired in transmission of a malaria parasite. Nature 417, 452–455 (2002)
Article ADS CAS Google Scholar - Moreira, L. A. et al. Bee venom phospholipase inhibits malaria parasite development in transgenic mosquitoes. J. Biol. Chem. 277, 40839–40843 (2002)
Article CAS Google Scholar - Li, F., Patra, K. P. & Vinetz, J. M. An anti-chitinase malaria transmission-blocking single-chain antibody as an effector molecule for creating a _Plasmodium falciparum_-refractory mosquito. J. Infect. Dis. 192, 878–887 (2005)
Article CAS Google Scholar - Sinkins, S. P. & Gould, F. Gene drive systems for insect disease vectors. Nature Rev. Genet. 7, 427–435 (2006)
Article CAS Google Scholar - Burt, A. Site-specific selfish genes as tools for the control and genetic engineering of natural populations. Proc. R. Soc. Lond. B 270, 921–928 (2003)
Article CAS Google Scholar - Catteruccia, F., Benton, J. P. & Crisanti, A. An Anopheles transgenic sexing strain for vector control. Nature Biotechnol. 23, 1414–1417 (2005)
Article CAS Google Scholar - Jacquier, A. & Dujon, B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell 41, 383–394 (1985)
Article CAS Google Scholar - Bellaiche, Y., Mogila, V. & Perrimon, N. I-SceI endonuclease, a new tool for studying DNA double-strand break repair mechanisms in Drosophila . Genetics 152, 1037–1044 (1999)
CAS PubMed PubMed Central Google Scholar - Windbichler, N. et al. Homing endonuclease mediated gene targeting in Anopheles gambiae cells and embryos. Nucleic Acids Res. 35, 5922–5933 (2007)
Article CAS Google Scholar - Stoddard, B. L. Homing endonuclease structure and function. Q. Rev. Biophys. 38, 49–95 (2005)
Article CAS Google Scholar - Goddard, M. R., Greig, D. & Burt, A. Outcrossed sex allows a selfish gene to invade yeast populations. Proc. R. Soc. Lond. B 268, 2537–2542 (2001)
Article CAS Google Scholar - Meredith, J. M. et al. Site-specific integration and expression of an anti-malarial gene in transgenic Anopheles gambiae significantly reduces Plasmodium infections. PLoS ONE 6, e14587 (2011)
Article ADS CAS Google Scholar - Burt, A. & Koufopanou, V. Homing endonuclease genes: the rise and fall and rise again of a selfish element. Curr. Opin. Genet. Dev. 14, 609–615 (2004)
Article CAS Google Scholar - Chen, C. H. et al. A synthetic maternal-effect selfish genetic element drives population replacement in Drosophila . Science 316, 597–600 (2007)
Article ADS CAS Google Scholar - McMeniman, C. J. et al. Stable introduction of a life-shortening Wolbachia infection into the mosquito Aedes aegypti . Science 323, 141–144 (2009)
Article ADS CAS Google Scholar - Ashworth, J. et al. Computational redesign of endonuclease DNA binding and cleavage specificity. Nature 441, 656–659 (2006)
Article ADS CAS Google Scholar - Jarjour, J. et al. High-resolution profiling of homing endonuclease binding and catalytic specificity using yeast surface display. Nucleic Acids Res. 37, 6871–6880 (2009)
Article CAS Google Scholar - Ashworth, J. et al. Computational reprogramming of homing endonuclease specificity at multiple adjacent base pairs. Nucleic Acids Res. 38, 5601–5608 (2010)
Article CAS Google Scholar - Thyme, S. B. et al. Exploitation of binding energy for catalysis and design. Nature 461, 1300–1304 (2009)
Article ADS CAS Google Scholar - Gao, H. et al. Heritable targeted mutagenesis in maize using a designed endonuclease. Plant J. 61, 176–187 (2010)
Article CAS Google Scholar - Grizot, S. et al. Efficient targeting of a SCID gene by an engineered single-chain homing endonuclease. Nucleic Acids Res. 37, 5405–5419 (2009)
Article CAS Google Scholar - Munoz, I. G. et al. Molecular basis of engineered meganuclease targeting of the endogenous human RAG1 locus. Nucleic Acids Res. 39, 729–743 (2010)
Article Google Scholar - Redondo, P. et al. Molecular basis of xeroderma pigmentosum group C DNA recognition by engineered meganucleases. Nature 456, 107–111 (2008)
Article ADS CAS Google Scholar - Arnould, S. et al. Engineered I-CreI derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells. J. Mol. Biol. 371, 49–65 (2007)
Article CAS Google Scholar - Rosen, L. E. et al. Homing endonuclease I-CreI derivatives with novel DNA target specificities. Nucleic Acids Res. 34, 4791–4800 (2006)
Article ADS CAS Google Scholar - Li, H., Pellenz, S., Ulge, U., Stoddard, B. L. & Monnat, R. J., Jr Generation of single-chain LAGLIDADG homing endonucleases from native homodimeric precursor proteins. Nucleic Acids Res. 37, 1650–1662 (2009)
Article CAS Google Scholar - Sheng, G., Thouvenot, E., Schmucker, D., Wilson, D. S. & Desplan, C. Direct regulation of rhodopsin 1 by Pax-6/eyeless in Drosophila: evidence for a conserved function in photoreceptors. Genes Dev. 11, 1122–1131 (1997)
Article CAS Google Scholar - Lobo, N. F., Clayton, J. R., Fraser, M. J., Kafatos, F. C. & Collins, F. H. High efficiency germ-line transformation of mosquitoes. Nature Protocols 1, 1312–1317 (2006)
Article CAS Google Scholar - Catteruccia, F., Godfray, H. C. & Crisanti, A. Impact of genetic manipulation on the fitness of Anopheles stephensi mosquitoes. Science 299, 1225–1227 (2003)
Article ADS CAS Google Scholar - Scalley-Kim, M., McConnell-Smith, A. & Stoddard, B. L. Coevolution of a homing endonuclease and its host target sequence. J. Mol. Biol. 372, 1305–1319 (2007)
Article CAS Google Scholar - Thyme, S. B. et al. Exploitation of binding energy for catalysis and design. Nature 461, 1300–1304 (2009)
Article ADS CAS Google Scholar - Doyon, J. B., Pattanayak, V., Meyer, C. B. & Liu, D. R. Directed evolution and substrate specificity profile of homing endonuclease I-SceI. J. Am. Chem. Soc. 128, 2477–2484 (2006)
Article CAS Google Scholar - Argast, G. M., Stephens, K. M., Emond, M. J. & Monnat, R. J., Jr I-_Ppo_I and I-_Cre_I homing site sequence degeneracy determined by random mutagenesis and sequential in vitro enrichment. J. Mol. Biol. 280, 345–353 (1998)
Article CAS Google Scholar - Ulge, U. Y., Baker, D. A. & Monnat, R. J. Comprehensive computational design of mCreI homing endonuclease cleavage specificity for genome engineering. Nucleic Acids Res. 10.1093/nar/gkr022 (1 February 2011)
- McConnell Smith, A. et al. Generation of a nicking enzyme that stimulates site-specific gene conversion from the I-AniI LAGLIDADG homing endonuclease. Proc. Natl Acad. Sci. USA 106, 5099–5104 (2009)
Article ADS CAS Google Scholar - Das, R. & Baker, D. Macromolecular modeling with Rosetta. Annu. Rev. Biochem. 77, 363–382 (2008)
Article CAS Google Scholar - Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005)
Article CAS Google Scholar
Acknowledgements
We thank M. Ashburner, S. Russell, D. Huen and S. Chan for comments, assistance and for plasmids. We thank M. P. Calos for providing the pET11phiC31polyA plasmid. We thank M. J. Fraser Jr for providing the pBSII-IFP2-orf plasmid. We thank J. Meredith and P. Eggleston for providing the docking strain. We thank A. Hall, T. Nolan, K. Magnusson, D. Rogers and S. Fuchs for assistance. We thank S. Arshiya Quadri and M. Szeto for experimental support and the members of the laboratories of D. Baker, R. Monnat, A. Scharenberg and B. Stoddard for their collective support of HEG engineering. A. F. M. Hackmann provided graphics support. Funded by a grant from the Foundation for the National Institutes of Health through the Vector-Based Control of Transmission: Discovery Research (VCTR) program of the Grand Challenges in Global Health initiative and by NIH RL1 awards GM084433 to D.B. and CA133831 to R.J.M.
Author information
Author notes
- Austin Burt and Andrea Crisanti: These authors contributed equally to this work.
Authors and Affiliations
- Department of Life Sciences, Imperial College London, South Kensington Campus, London, SW7 2AZ, UK
Nikolai Windbichler, Miriam Menichelli, Philippos Aris Papathanos, Austin Burt & Andrea Crisanti - Department of Biochemistry, University of Washington, Seattle, 98195, Washington, USA
Summer B. Thyme & David Baker - Graduate Program in Biomolecular Structure and Design, University of Washington, Seattle, 98195, Washington, USA
Summer B. Thyme & David Baker - Department of Pathology, University of Washington, Seattle, 98195, Washington, USA
Hui Li, Umut Y. Ulge & Raymond J. Monnat - Graduate Program in Molecular and Cellular Biology, University of Washington, Seattle, 98195, Washington, USA
Umut Y. Ulge & Raymond J. Monnat - Department of Genome Sciences, University of Washington, Seattle, 98195, Washington, USA
Blake T. Hovde & Raymond J. Monnat - Howard Hughes Medical Institute, University of Washington, Seattle, 98195, Washington, USA
David Baker - Department of Life Sciences, Imperial College London, Silwood Park Campus, Ascot, SL5 7PY, UK
Austin Burt - Department of Experimental Medicine, University of Perugia, Via Del Giochetto, 06122 Perugia, Italy
Andrea Crisanti
Authors
- Nikolai Windbichler
- Miriam Menichelli
- Philippos Aris Papathanos
- Summer B. Thyme
- Hui Li
- Umut Y. Ulge
- Blake T. Hovde
- David Baker
- Raymond J. Monnat
- Austin Burt
- Andrea Crisanti
Contributions
N.W. designed the experiments. N.W., M.M. and P.A.P. performed the experiments. N.W. and P.A.P. generated the transgenic lines. M.M. maintained mosquito populations. N.W. analysed the data. A.B. and N.W. generated the population dynamic models. A.C. and A.B. inspired the work and wrote the paper together with N.W. HEG redesign and target site cleavage analyses were performed by S.B.T., H.L., U.Y.U. (contributed equally) and B.T.H. with guidance from D.B. and R.J.M. All authors read and approved the final manuscript.
Corresponding author
Correspondence toAndrea Crisanti.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Windbichler, N., Menichelli, M., Papathanos, P. et al. A synthetic homing endonuclease-based gene drive system in the human malaria mosquito.Nature 473, 212–215 (2011). https://doi.org/10.1038/nature09937
- Received: 28 September 2010
- Accepted: 16 February 2011
- Published: 20 April 2011
- Issue date: 12 May 2011
- DOI: https://doi.org/10.1038/nature09937
This article is cited by
Comments
Commenting on this article is now closed.
- Nikolai Windbichler 8 July 2011, 06:06
Similar observations have been made in Drosophila by Sang Chan, Daniel Naujoks, David Huen and Steven Russell published in Genetics:
http://www.genetics.org/con...
Editorial Summary
Manipulating an insect vector
Genetic approaches to manipulating or eradicating disease vectors have been proposed as alternatives to malaria eradication. The success of this approach depends on efficient spread of a genetic modification in field populations. Windbichler et al. show that a synthetic genetic element consisting of mosquito regulatory elements and the homing endonuclease gene I-SceI can spread from a small number of individual Anopheles gambiae mosquitoes into large receptive populations in just a few generations. This is the first demonstration of a synthetic gene drive system in the main human malaria vector — and a similar approach should be applicable to many other pest species.