mTORC1 couples immune signals and metabolic programming to establish Treg-cell function (original) (raw)
- Letter
- Published: 30 June 2013
Nature volume 499, pages 485–490 (2013)Cite this article
- 42k Accesses
- 759 Citations
- 48 Altmetric
- Metrics details
Subjects
Abstract
The mechanistic target of rapamycin (mTOR) pathway integrates diverse environmental inputs, including immune signals and metabolic cues, to direct T-cell fate decisions1. The activation of mTOR, which is the catalytic subunit of the mTORC1 and mTORC2 complexes, delivers an obligatory signal for the proper activation and differentiation of effector CD4+ T cells2,3, whereas in the regulatory T-cell (Treg) compartment, the Akt–mTOR axis is widely acknowledged as a crucial negative regulator of Treg-cell de novo differentiation4,5,6,7,8 and population expansion9. However, whether mTOR signalling affects the homeostasis and function of Treg cells remains largely unexplored. Here we show that mTORC1 signalling is a pivotal positive determinant of Treg-cell function in mice. Treg cells have elevated steady-state mTORC1 activity compared to naive T cells. Signals through the T-cell antigen receptor (TCR) and interleukin-2 (IL-2) provide major inputs for mTORC1 activation, which in turn programs the suppressive function of Treg cells. Disruption of mTORC1 through Treg-specific deletion of the essential component raptor leads to a profound loss of Treg-cell suppressive activity in vivo and the development of a fatal early onset inflammatory disorder. Mechanistically, raptor/mTORC1 signalling in Treg cells promotes cholesterol and lipid metabolism, with the mevalonate pathway particularly important for coordinating Treg-cell proliferation and upregulation of the suppressive molecules CTLA4 and ICOS to establish Treg-cell functional competency. By contrast, mTORC1 does not directly affect the expression of Foxp3 or anti- and pro-inflammatory cytokines in Treg cells, suggesting a non-conventional mechanism for Treg-cell functional regulation. Finally, we provide evidence that mTORC1 maintains Treg-cell function partly through inhibiting the mTORC2 pathway. Our results demonstrate that mTORC1 acts as a fundamental ‘rheostat’ in Treg cells to link immunological signals from TCR and IL-2 to lipogenic pathways and functional fitness, and highlight a central role of metabolic programming of Treg-cell suppressive activity in immune homeostasis and tolerance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: mTORC1 signalling is constitutively active in Treg cells and its disruption results in a fatal early onset inflammatory disorder.
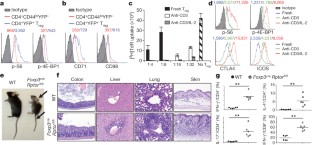
The alternative text for this image may have been generated using AI.
Figure 2: Raptor deletion abrogates Treg-cell suppressive activity.
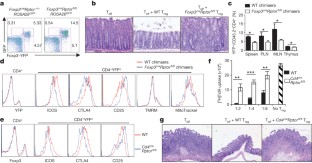
The alternative text for this image may have been generated using AI.
Figure 3: Raptor coordinates Treg-cell proliferation and effector molecule expression by orchestrating cholesterol/lipid biosynthetic metabolism, especially the mevalonate pathway.
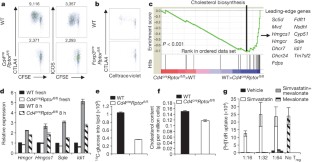
The alternative text for this image may have been generated using AI.
Figure 4: Deletion of Rictor does not alter Treg-cell function but partially rescues inflammation in Foxp3 cre _Rptor_fl/fl mice.
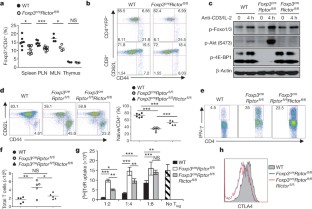
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Accessions
Gene Expression Omnibus
Data deposits
The microarray data have been deposited in the Gene Expression Omnibus under accession GSE46693.
References
- Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nature Rev. Immunol. 12, 325–338 (2012)
Article CAS Google Scholar - Delgoffe, G. M. et al. The kinase mTOR regulates the differentiation of helper T cells through the selective activation of signaling by mTORC1 and mTORC2. Nature Immunol. 12, 295–303 (2011)
Article CAS Google Scholar - Lee, K. et al. Mammalian target of rapamycin protein complex 2 regulates differentiation of Th1 and Th2 cell subsets via distinct signaling pathways. Immunity 32, 743–753 (2010)
Article CAS Google Scholar - Delgoffe, G. M. et al. The mTOR kinase differentially regulates effector and regulatory T cell lineage commitment. Immunity 30, 832–844 (2009)
Article CAS Google Scholar - Haxhinasto, S., Mathis, D. & Benoist, C. The AKT–mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J. Exp. Med. 205, 565–574 (2008)
Article CAS Google Scholar - Sauer, S. et al. T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc. Natl Acad. Sci. USA 105, 7797–7802 (2008)
Article ADS CAS Google Scholar - Liu, G. et al. The receptor S1P1 overrides regulatory T cell-mediated immune suppression through Akt-mTOR. Nature Immunol. 10, 769–777 (2009)
Article CAS Google Scholar - Liu, G., Yang, K., Burns, S., Shrestha, S. & Chi, H. The S1P1-mTOR axis directs the reciprocal differentiation of TH1 and Treg cells. Nature Immunol. 11, 1047–1056 (2010)
Article CAS Google Scholar - Battaglia, M., Stabilini, A. & Roncarolo, M. G. Rapamycin selectively expands CD4+CD25+FoxP3+ regulatory T cells. Blood 105, 4743–4748 (2005)
Article CAS Google Scholar - Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012)
Article CAS Google Scholar - Procaccini, C. et al. An oscillatory switch in mTOR kinase activity sets regulatory T cell responsiveness. Immunity 33, 929–941 (2010)
Article CAS Google Scholar - Kelly, A. P. et al. Notch-induced T cell development requires phosphoinositide-dependent kinase 1. EMBO J. 26, 3441–3450 (2007)
Article CAS Google Scholar - Cunningham, J. T. et al. mTOR controls mitochondrial oxidative function through a YY1–PGC-1α transcriptional complex. Nature 450, 736–740 (2007)
Article ADS CAS Google Scholar - van der Windt, G. J. et al. Mitochondrial respiratory capacity is a critical regulator of CD8+ T cell memory development. Immunity 36, 68–78 (2012)
Article CAS Google Scholar - Gurumurthy, S. et al. The Lkb1 metabolic sensor maintains haematopoietic stem cell survival. Nature 468, 659–663 (2010)
Article ADS CAS Google Scholar - Takahashi, T. et al. Immunologic self-tolerance maintained by CD25+CD4+ naturally anergic and suppressive T cells: induction of autoimmune disease by breaking their anergic/suppressive state. Int. Immunol. 10, 1969–1980 (1998)
Article CAS Google Scholar - Fisson, S. et al. Continuous activation of autoreactive CD4+ CD25+ regulatory T cells in the steady state. J. Exp. Med. 198, 737–746 (2003)
Article CAS Google Scholar - Gavin, M. A., Clarke, S. R., Negrou, E., Gallegos, A. & Rudensky, A. Homeostasis and anergy of CD4+CD25+ suppressor T cells in vivo. Nature Immunol. 3, 33–41 (2002)
Article CAS Google Scholar - Thornton, A. M., Donovan, E. E., Piccirillo, C. A. & Shevach, E. M. Cutting edge: IL-2 is critically required for the in vitro activation of CD4+CD25+ T cell suppressor function. J. Immunol. 172, 6519–6523 (2004)
Article CAS Google Scholar - Herman, A. E., Freeman, G. J., Mathis, D. & Benoist, C. CD4+CD25+ T regulatory cells dependent on ICOS promote regulation of effector cells in the prediabetic lesion. J. Exp. Med. 199, 1479–1489 (2004)
Article CAS Google Scholar - Wing, K. et al. CTLA-4 control over Foxp3+ regulatory T cell function. Science 322, 271–275 (2008)
Article ADS CAS Google Scholar - Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008)
Article CAS Google Scholar - Kanangat, S. et al. Disease in the scurfy (sf) mouse is associated with overexpression of cytokine genes. Eur. J. Immunol. 26, 161–165 (1996)
Article CAS Google Scholar - Chaudhry, A. et al. Interleukin-10 signaling in regulatory T cells is required for suppression of Th17 cell-mediated inflammation. Immunity 34, 566–578 (2011)
Article CAS Google Scholar - Josefowicz, S. Z., Lu, L. F. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012)
Article CAS Google Scholar - Ouyang, W. et al. Novel Foxo1-dependent transcriptional programs control Treg cell function. Nature 491, 554–559 (2012)
Article ADS CAS Google Scholar - Wang, R. & Green, D. R. Metabolic checkpoints in activated T cells. Nature Immunol. 13, 907–915 (2012)
Article CAS Google Scholar - Tai, X. et al. Basis of CTLA-4 function in regulatory and conventional CD4+ T cells. Blood 119, 5155–5163 (2012)
Article CAS Google Scholar - Corse, E. & Allison, J. P. Cutting edge: CTLA-4 on effector T cells inhibits in trans. J. Immunol. 189, 1123–1127 (2012)
Article CAS Google Scholar - Sadlack, B. et al. Ulcerative colitis-like disease in mice with a disrupted interleukin-2 gene. Cell 75, 253–261 (1993)
Article CAS Google Scholar - Yang, K., Neale, G., Green, D. R., He, W. & Chi, H. The tumor suppressor Tsc1 enforces quiescence of naive T cells to promote immune homeostasis and function. Nature Immunol. 12, 888–897 (2011)
Article CAS Google Scholar - Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Article ADS CAS Google Scholar
Acknowledgements
We thank A. Rudensky for _Foxp3_YFP-Cre mice, D. Green for help with metabolic assays, N. Brydon for animal colony management, Y. Wang for editing of the manuscript, and the St Jude Immunology FACS core facility for cell sorting. This work was supported by the US National Institutes of Health (K01 AR053573, R21 AI094089, R01 AI101407 and R01 NS064599), the Lupus Research Institute, and the American Lebanese Syrian Associated Charities (all to H.C.).
Author information
Authors and Affiliations
- Department of Immunology, St Jude Children’s Research Hospital, Memphis, 38105, Tennessee, USA
Hu Zeng, Kai Yang, Caryn Cloer & Hongbo Chi - Hartwell Center for Bioinformatics and Biotechnology, St Jude Children’s Research Hospital, Memphis, 38105, Tennessee, USA
Geoffrey Neale - Department of Pathology, St Jude Children’s Research Hospital, Memphis, 38105, Tennessee, USA
Peter Vogel
Authors
- Hu Zeng
- Kai Yang
- Caryn Cloer
- Geoffrey Neale
- Peter Vogel
- Hongbo Chi
Contributions
H.Z. designed and performed experiments, and wrote the manuscript; K.Y. contributed to cellular experiments; C.C. contributed to survival curves and technical support; G.N. performed bioinformatic analyses; P.V. performed histological analysis; H.C. designed experiments, contributed to writing the manuscript, and provided overall direction.
Corresponding author
Correspondence toHongbo Chi.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Zeng, H., Yang, K., Cloer, C. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function.Nature 499, 485–490 (2013). https://doi.org/10.1038/nature12297
- Received: 23 December 2012
- Accepted: 13 May 2013
- Published: 30 June 2013
- Issue date: 25 July 2013
- DOI: https://doi.org/10.1038/nature12297
This article is cited by
Editorial Summary
Treg cells linked to metabolic fitness
Foxp3-expressing regulatory T cells (Treg) have a central role in immune tolerance and homeostasis. Here Hongbo Chi and colleagues show that mTORC1-dependent cholesterol biosynthesis is important for Treg function, acting in part by upregulating the Treg effector molecules CTLA4 and ICOS. This finding suggests a linkage between immune signals and the metabolic state of a cell.