Mutations of the SLX4 gene in Fanconi anemia (original) (raw)
- Letter
- Published: 16 January 2011
- Francis P Lach1 na1,
- Rohini Desetty1,
- Helmut Hanenberg2,3,
- Arleen D Auerbach4 &
- …
- Agata Smogorzewska1
Nature Genetics volume 43, pages 142–146 (2011)Cite this article
- 6454 Accesses
- 340 Citations
- 3 Altmetric
- Metrics details
Subjects
Abstract
Fanconi anemia is a rare recessive disorder characterized by genome instability, congenital malformations, progressive bone marrow failure and predisposition to hematologic malignancies and solid tumors1. At the cellular level, hypersensitivity to DNA interstrand crosslinks is the defining feature in Fanconi anemia2. Mutations in thirteen distinct Fanconi anemia genes3 have been shown to interfere with the DNA-replication–dependent repair of lesions involving crosslinked DNA at stalled replication forks4. Depletion of SLX4, which interacts with multiple nucleases and has been recently identified as a Holliday junction resolvase5,6,7, results in increased sensitivity of the cells to DNA crosslinking agents. Here we report the identification of biallelic SLX4 mutations in two individuals with typical clinical features of Fanconi anemia and show that the cellular defects in these individuals' cells are complemented by wildtype SLX4, demonstrating that biallelic mutations in SLX4 (renamed here as FANCP) cause a new subtype of Fanconi anemia, Fanconi anemia-P.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Characterization of cell lines from individuals with Fanconi anemia with SLX4 mutations.
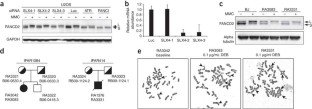
The alternative text for this image may have been generated using AI.
Figure 2: SLX4 is defective in two individuals with Fanconi anemia.
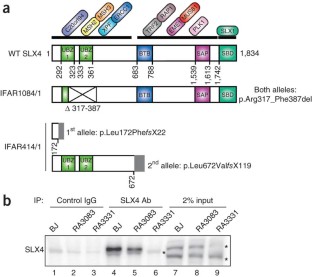
The alternative text for this image may have been generated using AI.
Figure 3: Complementation of RA3083 and RA3331 cells with the SLX4 cDNA.
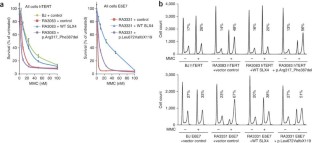

The alternative text for this image may have been generated using AI.
Figure 4: Interaction of mutant forms of SLX4 with its partners and with ubiquitin.
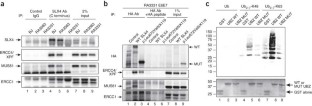
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Accessions
NCBI Reference Sequence
References
- Auerbach, A.D. Fanconi anemia and its diagnosis. Mutat. Res. 668, 4–10 (2009).
Article CAS Google Scholar - Auerbach, A.D. & Wolman, S.R. Susceptibility of Fanconi′s anaemia fibroblasts to chromosome damage by carcinogens. Nature 261, 494–496 (1976).
Article CAS Google Scholar - Moldovan, G.L. & D'Andrea, A.D. How the fanconi anemia pathway guards the genome. Annu. Rev. Genet. 43, 223–249 (2009).
Article CAS Google Scholar - Knipscheer, P. et al. The Fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science 326, 1698–1701 (2009).
Article CAS Google Scholar - Fekairi, S. et al. Human SLX4 is a Holliday junction resolvase subunit that binds multiple DNA repair/recombination endonucleases. Cell 138, 78–89 (2009).
Article CAS Google Scholar - Munoz, I.M. et al. Coordination of structure-specific nucleases by human SLX4/BTBD12 is required for DNA repair. Mol. Cell 35, 116–127 (2009).
Article CAS Google Scholar - Svendsen, J.M. et al. Mammalian BTBD12/SLX4 assembles a Holliday junction resolvase and is required for DNA repair. Cell 138, 63–77 (2009).
Article CAS Google Scholar - Ciccia, A., McDonald, N. & West, S.C. Structural and functional relationships of the XPF/MUS81 family of proteins. Annu. Rev. Biochem. 77, 259–287 (2008).
Article CAS Google Scholar - Garcia-Higuera, I. et al. Interaction of the Fanconi anemia proteins and BRCA1 in a common pathway. Mol. Cell 7, 249–262 (2001).
Article CAS Google Scholar - Smogorzewska, A. et al. Identification of the FANCI protein, a monoubiquitinated FANCD2 paralog required for DNA repair. Cell 129, 289–301 (2007).
Article CAS Google Scholar - Kratz, K. et al. Deficiency of FANCD2-associated nuclease KIAA1018/FAN1 sensitizes cells to interstrand crosslinking agents. Cell 142, 77–88 (2010).
Article CAS Google Scholar - Liu, T., Ghosal, G., Yuan, J., Chen, J. & Huang, J. FAN1 acts with FANCI-FANCD2 to promote DNA interstrand cross-link repair. Science 329, 693–696 (2010).
Article CAS Google Scholar - MacKay, C. et al. Identification of KIAA1018/FAN1, a DNA repair nuclease recruited to DNA damage by monoubiquitinated FANCD2. Cell 142, 65–76 (2010).
Article CAS Google Scholar - Smogorzewska, A. et al. A genetic screen identifies FAN1, a Fanconi anemia-associated nuclease necessary for DNA interstrand crosslink repair. Mol. Cell 39, 36–47 (2010).
Article CAS Google Scholar - Vaz, F. et al. Mutation of the RAD51C gene in a Fanconi anemia-like disorder. Nat. Genet. 42, 406–409 (2010).
Article CAS Google Scholar - Kutler, D.I. et al. A 20-year perspective on the International Fanconi Anemia Registry (IFAR). Blood 101, 1249–1256 (2003).
Article CAS Google Scholar - Hofmann, K. Ubiquitin-binding domains and their role in the DNA damage response. DNA Repair (Amst.) 8, 544–556 (2009).
Article CAS Google Scholar - Sobhian, B. et al. RAP80 targets BRCA1 to specific ubiquitin structures at DNA damage sites. Science 316, 1198–1202 (2007).
Article CAS Google Scholar - Howlett, N.G. et al. Biallelic inactivation of BRCA2 in Fanconi anemia. Science 297, 606–609 (2002).
Article CAS Google Scholar - Xia, B. et al. Fanconi anemia is associated with a defect in the BRCA2 partner PALB2. Nat. Genet. 39, 159–161 (2007).
Article CAS Google Scholar - Meindl, A. et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat. Genet. 42, 410–414 (2010).
Article CAS Google Scholar - Rahman, N. et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat. Genet. 39, 165–167 (2007).
Article CAS Google Scholar - Seal, S. et al. Truncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility alleles. Nat. Genet. 38, 1239–1241 (2006).
Article CAS Google Scholar - Wooster, R. et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 378, 789–792 (1995).
Article CAS Google Scholar - Lamesch, P. et al. hORFeome v3.1: a resource of human open reading frames representing over 10,000 human genes. Genomics 89, 307–315 (2007).
Article CAS Google Scholar - Dolma, S., Lessnick, S.L., Hahn, W.C. & Stockwell, B.R. Identification of genotype-selective antitumor agents using synthetic lethal chemical screening in engineered human tumor cells. Cancer Cell 3, 285–296 (2003).
Article CAS Google Scholar
Acknowledgements
We are grateful to the affected individuals and their families for their participation in this study. We thank the Harper Lab, Harvard Medical School, Boston, Massachusetts, USA for reagents, E. Foley for advice and J. de Winter for communicating unpublished results. H.H. is supported by the Deutsche Forschungsgemeinschaft SPP1230, the Bundesministerium für Bildung und Forschung network for Bone Marrow failure Syndrome, and FoneFA. A.S. is supported by the Burroughs Wellcome Fund Career Award for Medical Scientists and is a Rita Allen Foundation and an Irma T. Hirschl Scholar.
Author information
Author notes
- Yonghwan Kim and Francis P Lach: These authors contributed equally to this work.
Authors and Affiliations
- Laboratory of Genome Maintenance, The Rockefeller University, New York, New York, USA
Yonghwan Kim, Francis P Lach, Rohini Desetty & Agata Smogorzewska - Division of Pediatric Hematology/Oncology, Herman B. Wells Center for Pediatric Research, Riley Hospital for Children, Indiana University School of Medicine, Indianapolis, Indiana, USA
Helmut Hanenberg - Department of Otorhinolaryngology, Heinrich Heine University, Duesseldorf, Germany
Helmut Hanenberg - Human Genetics and Hematology, The Rockefeller University, New York, New York, USA
Arleen D Auerbach
Authors
- Yonghwan Kim
- Francis P Lach
- Rohini Desetty
- Helmut Hanenberg
- Arleen D Auerbach
- Agata Smogorzewska
Contributions
The study was designed by A.S., Y.K. and F.P.L. Subject recruitment and sample collection was done by A.D.A., F.P.L. and A.S. Characterization with respect to Fanconi anemia subgroups was performed by A.S., F.P.L., H.H. and A.D.A. Mutation analysis and functional studies were performed by A.S., Y.K., F.P.L. and R.D. The manuscript was written by A.S. with help from other authors.
Corresponding author
Correspondence toAgata Smogorzewska.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Kim, Y., Lach, F., Desetty, R. et al. Mutations of the SLX4 gene in Fanconi anemia.Nat Genet 43, 142–146 (2011). https://doi.org/10.1038/ng.750
- Received: 31 August 2010
- Accepted: 15 December 2010
- Published: 16 January 2011
- Issue date: February 2011
- DOI: https://doi.org/10.1038/ng.750