Detection of differentially culturable tubercle bacteria in sputum using mycobacterial culture filtrates (original) (raw)
Introduction
The global tuberculosis (TB) epidemic remains a leading cause of death with an estimated 10 million people afflicted with the disease and 1.5 million deaths annually1. Moreover, a large proportion of people harbour the infectious agent, Mycobacterium tuberculosis (Mtb), in lung lesions without any symptoms, providing a massive reservoir of the pathogen that can reactivate to cause active disease and drive further transmission. TB is curable however, the 6 month protracted treatment regimen is difficult to implement and challenges related to patient adherence result in the development of drug resistant forms of the pathogen that cannot be eradicated using the current standard first line drugs[2](/articles/s41598-021-86054-z#ref-CR2 "Dheda, K., Barry, C. E. 3rd. & Maartens, G. Tuberculosis. Lancet 387, 1211–1226. https://doi.org/10.1016/S0140-6736(15)00151-8
(2016)."). Although several clinical trials have investigated novel treatment shortening therapies, published trials have yet to show broad benefit, possibly due to limited understanding of mycobacterial growth states during active and asymptomatic or subclinical disease.Several studies recently have provided evidence that sputum from TB patients with active disease is characterised by a large fraction of non-replicating bacteria[3](#ref-CR3 "Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540. https://doi.org/10.1164/rccm.201604-0769OC
(2016)."),[4](#ref-CR4 "Dartois, V., Saito, K., Warrier, T. & Nathan, C. New evidence for the complexity of the population structure of Mycobacterium tuberculosis increases the diagnostic and biologic challenges. Am. J. Respir. Crit. Care Med. 194, 1448–1451.
https://doi.org/10.1164/rccm.201607-1431ED
(2016)."),[5](#ref-CR5 "Garton, N. J. et al. Cytological and transcript analyses reveal fat and lazy persister-like bacilli in tuberculous sputum. PLoS Med. 5, e75.
https://doi.org/10.1371/journal.pmed.0050075
(2008)."),[6](/articles/s41598-021-86054-z#ref-CR6 "Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G. & Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle Bacilli in Sputum. Am. J. Respir. Crit. Care Med. 181, 174–180.
https://doi.org/10.1164/rccm.200905-0661OC
(2009)."). A commonly used diagnostic method for detection of _Mtb_ in patient sputum is enumerating mycobacterial colony forming units (CFU) of culture dilution suspensions on solid agar media. However, this methodology is limited to detecting only replicating mycobacterial populations and is unable to account for the non-replicating population, which has important implications for TB diagnostics. This is illustrated by prior work that demonstrated the presence of differentially culturable tubercle bacteria (DCTB) in sputum that do not grow on solid media and are detected only in liquid media supplemented with culture filtrate (CF)[3](/articles/s41598-021-86054-z#ref-CR3 "Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540.
https://doi.org/10.1164/rccm.201604-0769OC
(2016)."),[6](/articles/s41598-021-86054-z#ref-CR6 "Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G. & Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle Bacilli in Sputum. Am. J. Respir. Crit. Care Med. 181, 174–180.
https://doi.org/10.1164/rccm.200905-0661OC
(2009)."). This demonstrates that TB infection presents with a much higher level of bacterial phenotypic complexity than previously thought and a deeper understanding of the clinical relevance of DCTB populations is vital for the advancement of improved TB diagnosis and development of novel drug regimens[4](/articles/s41598-021-86054-z#ref-CR4 "Dartois, V., Saito, K., Warrier, T. & Nathan, C. New evidence for the complexity of the population structure of Mycobacterium tuberculosis increases the diagnostic and biologic challenges. Am. J. Respir. Crit. Care Med. 194, 1448–1451.
https://doi.org/10.1164/rccm.201607-1431ED
(2016).").The growth stimulatory activity of CF has been ascribed to the presence of resuscitation promoting factors (Rpfs), a group of secreted enzymes in Mtb[6](/articles/s41598-021-86054-z#ref-CR6 "Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G. & Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle Bacilli in Sputum. Am. J. Respir. Crit. Care Med. 181, 174–180. https://doi.org/10.1164/rccm.200905-0661OC
(2009)."),[7](/articles/s41598-021-86054-z#ref-CR7 "Kana, B. D. & Mizrahi, V. Resuscitation-promoting factors as lytic enzymes for bacterial growth and signaling. FEMS Immunol. Med. Microbiol. 58, 39–50.
https://doi.org/10.1111/j.1574-695X.2009.00606.x
(2010).") however, use of this approach is limited by the availability of freshly prepared _Mtb_ CF in diagnostic laboratories. To investigate alternative approaches for detection of DCTB, we explored the use of CF from _Mycobacterium smegmatis_ (_Msm_), a non-pathogenic close relative of _Mtb,_ with the hypothesis that _Msm_ CF would provide a more safer and practical approach in routine laboratories. Bioinformatic analysis identified three _rpf_ genes with a duplicate _rpfE_ homolog (_rpfA_, _rpfB_, _rpfE1_ and _rpfE2_) in _Msm_ compared to the five homologous (_rpfA_\-_rpfE_) in _Mtb_[8](/articles/s41598-021-86054-z#ref-CR8 "Machowski, E. E., Senzani, S., Ealand, C. & Kana, B. D. Comparative genomics for mycobacterial peptidoglycan remodelling enzymes reveals extensive genetic multiplicity. BMC Microbiol. 14, 75.
https://doi.org/10.1186/1471-2180-14-75
(2014).") (Figure [S1](/articles/s41598-021-86054-z#MOESM1)). In prior work, we generated a mutant of _Msm_ devoid of all Rpfs[9](/articles/s41598-021-86054-z#ref-CR9 "Ealand, C. et al. Resuscitation-promoting factors are required for Mycobacterium smegmatis biofilm formation. Appl. Environ. Microbiol. 8, 4.
https://doi.org/10.1128/AEM.00687-18
(2018).") and herein, we exploited this strain to generate complemented mutant strains that carry _Mtb_ Rpfs in various combinations. These recombinant strains were used to generate CF for detection of DCTB in sputum. We found that _Mtb_ CF was superior at detecting DCTB when compared to CF from _Msm_ and whilst the addition of _Mtb_ Rpfs to _Msm_ did improve recovery of DCTB, the bacterial yield was not comparable to that observed with _Mtb_ CF. It has also been demonstrated that fatty acids and cyclic-AMP (cAMP) are able to stimulate the growth of DCTB in an in vitro model of mycobacterial dormancy[10](/articles/s41598-021-86054-z#ref-CR10 "Shleeva, M. et al. Cyclic AMP-dependent resuscitation of dormant Mycobacteria by exogenous free fatty acids. PLoS ONE 8, e82914.
https://doi.org/10.1371/journal.pone.0082914
(2013)."). Hence, we also tested various fatty acids and cAMP for the ability to resuscitate DCTB and found that no single agent was able to stimulate bacterial growth in a manner comparable to _Mtb_ CF.Results
Construction of Msm mutant strains expressing Mtb rpf genes
To assess the role of Mtb Rpfs in stimulation of DCTB, we generated strains of Msm that expressed Mtb rpf genes. The inclusion of antibiotics during the culture of these strains to ensure that _rpf_-expressing plasmids are retained was problematic as the antibiotic would be carried into the resulting CF, leading to inhibition of bacterial growth in sputum cultures. To address this, we used integrating plasmids as these carry an integrase to facilitate plasmid integration into the bacterial chromosome at phage attachment sites11,[12](/articles/s41598-021-86054-z#ref-CR12 "Pham, T. T., Jacobs-Sera, D., Pedulla, M. L., Hendrix, R. W. & Hatfull, G. F. Comparative genomic analysis of mycobacteriophage Tweety: evolutionary insights and construction of compatible site-specific integration vectors for mycobacteria. Microbiology (Reading) 153, 2711–2723. https://doi.org/10.1099/mic.0.2007/008904-0
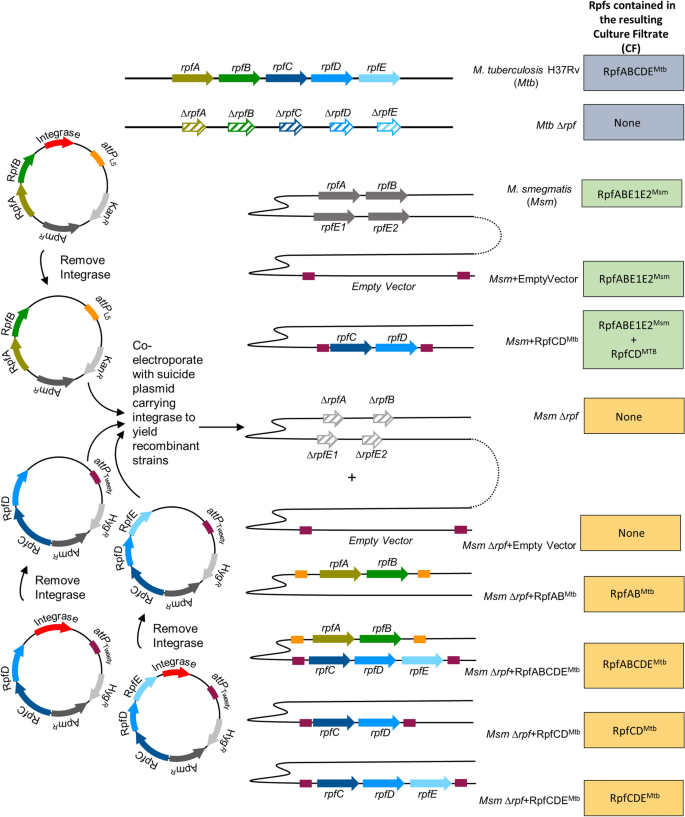
(2007)."). However, these integrases can also spontaneously excise vectors in the absence of antibiotic selection. To overcome this, we generated versions of conditionally integrative plasmids that lacked the integrase gene and then co-electroporated these with the suicide plasmid that provided the integrase _in trans_ (Fig. [1](/articles/s41598-021-86054-z#Fig1)). With this approach, the integrase is present during the initial integration event and is then lost during subsequent cell division cycles. We used a mutant of _Msm_, deleted for all four of the _Msm rpf_ genes to build recombinant strains expressing the different _Mtb rpf_ genes. The resulting strains were then used to generate CF from strains expressing _rpf_ genes from _Mtb_ for use in the MPN assay for the detection of DCTB. Wild type _Msm_, carrying the _Msm_ Rpfs was also used as a control. The strains generated and predicted complement of Rpf proteins from the resulting CF are shown in Fig. [1](/articles/s41598-021-86054-z#Fig1).Figure 1
The alternative text for this image may have been generated using AI.
Strategy for construction of recombinant Msm strains expressing Mtb Rpfs in various combinations. The Mtb H37Rv wild type and Rpf defective mutant (Mtb ∆rpf) were used as a source of Mtb culture filtrates (CFs). Wild type Msm was used as a source of Msm Rpfs and this strain was transformed with a vector carrying RpfCD_Mtb_ to mimic the Rpf complement from Mtb. The Msm mutant defective for Rpfs (Msm ∆rpf) was used as a control and within this mutant background, vectors carrying various combinations of Mtb Rpfs were inserted to create strains that express Mtb Rpfs in Msm CF as a source of growth stimulatory molecules in MPN DCTB assays. The panel on the right shows the rpf gene complement present in the various Mtb and Msm strains. The vector diagram on the left depicts the strategy to construct vectors with different combinations of Mtb Rpfs that are stably carried in the resulting recombinant strains. The presence of the integrase on mycobacterial vectors that integrate phages at attachments sites, facilitates the first integration event but then can also lead to excision of the vector in the absence of antibiotic selection. For DCTB assays, CF that is free of antibiotic was required as antibiotics will inhibit the growth of sputum-derived bacteria. To facilitate this, the integrase was first removed from vectors expressing Mtb Rpfs and was provided in trans on another suicide vector during transformation. With this approach, the integrase is present for the first integration event and is then lost as it does not carry a mycobacterial origin of replication.
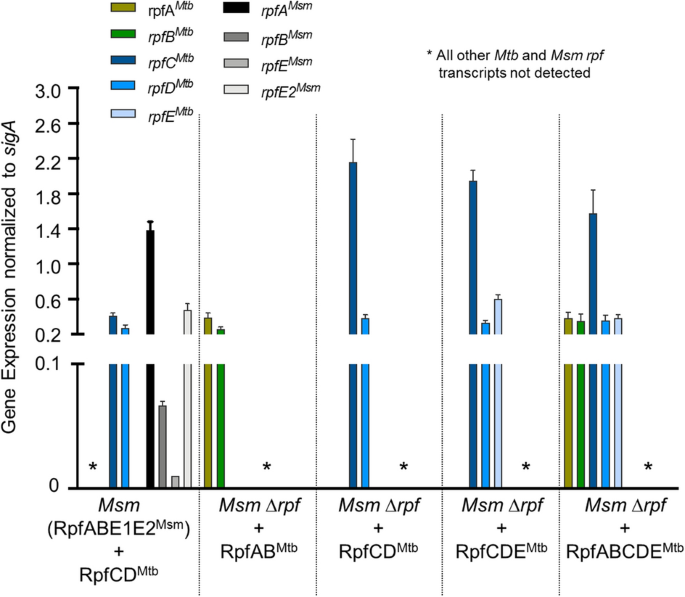
To confirm that the _rpf_-like genes from Mtb are expressed in Msm from the modified integrative plasmids, we conducted transcriptional analysis using quantitative real-time PCR. Recombinant Msm strains expressed all Mtb rpf genes provided on integrative plasmids. With the exception of rpfC Mtb, all Mtb rpf genes were expressed at equivalent levels, Fig. 2. As we had used native Mtb promoters, we expected some difference in rpf gene expression in recombinant Msm strains. As the Mtb genes were expressed in Msm, we also assessed expression of _Msm rpf_-like genes for comparative purposes. The rpfA Msm gene was expressed at the highest level relative to the other _Msm rpf_-like genes and the rpfE2 Msm gene was expressed at levels similar to the rpfABDE Mtb genes.
Figure 2
The alternative text for this image may have been generated using AI.
Assessment of Mtb and Msm rpf gene expression in recombinant strains. Transcriptional analysis was carried out on recombinant strains of Msm and the cognate parent strains. In all strains shown, transcriptional analysis was carried out for all five Mtb and four Msm rpf genes. Where these genes were deleted, no transcripts were detected. Transcript levels were assessed in each strain when axenic cultures reached mid-exponential growth (OD600nm ~ 0.6). Gene expression was normalized against Msm sigA transcript levels. Data are representative of three independent biological repeats. Graph was generated using Graphpad Prism software.
We anticipated that both recombinant Mtb and Msm Rpfs would be secreted as analysis of the domain structure of these proteins confirmed the presence of a SecA-dependent signal sequences in all of them, except Mtb RpfC (Figure S1). We further confirmed the presence of these signal peptides using Signal P analysis, which identified clear Sec-dependent signal peptides in all proteins, except RpfC (Figure S2). In addition, both SecA1 and SecA2 are conserved in Msm and Mtb (Figure S3), which when combined with the demonstrated secretion of Rpfs in prior work[13](/articles/s41598-021-86054-z#ref-CR13 "Mukamolova, G. V. et al. A family of autocrine growth factors in Mycobacterium tuberculosis. Mol. Microbiol. 46, 623–635. https://doi.org/10.1046/j.1365-2958.2002.03184.x
(2002)."), suggests that recombinant Rpfs are most likely secreted from the cell and these proteins, or the products of their catalytic activity, are present in the CF used in this study.Quantification of DCTB in sputum specimens using different mycobacterial CF preparations
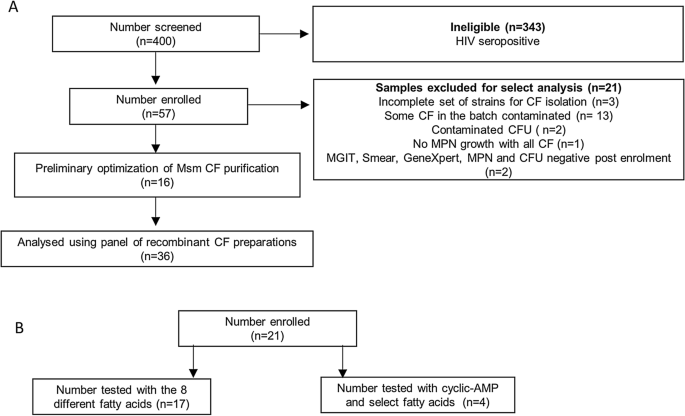
To assess the capacity of CF from the various mycobacterial strains generated to detect DCTB, we established a cross-sectional observation cohort of HIV-negative individuals with drug susceptible TB. The participant disposition flow chart for our cohort is given in Fig. 3A. We screened 400 individuals and enrolled 57 individuals. A large number of screen failures were due to co-incident HIV infection and these were excluded from our study due to the low bacterial burden that is expected to prevail in sputum. A summary of participant demographics and laboratory data is included in Table 1. All included participants were sputum smear positive and the majority had medium or high bacterial loads by GeneXpert, with a median time to culture positivity in the MGIT system of 5.5 days.
Figure 3
The alternative text for this image may have been generated using AI.
Participant disposition flow chart for individuals recruited to this study. (A) Sputum specimens obtained from these individuals were decontaminated and the resulting bacteria grown in limiting dilution assays with various CF combinations. (B) Sputum specimens obtained from these individuals were decontaminated and the ability of the various fatty acids and cyclic-AMP to resuscitate the DCTB was assessed by MPN and CFU assays.
Table 1 Demographics and laboratory diagnostic data for participant specimens tested with various CFs and FAs.
Sixteen specimens were used for methods optimization as we had contamination of the CF with bacteria, requiring iterative revision of the filtering protocol. Two specimens yielded contamination on CFU plates, 1 specimen gave no Mtb on MPN assays and 2 specimens were negative for MGIT, Smear, GeneXpert, MPN and CFU after enrolment (Fig. 3A). After these exclusions, 36 specimens were tested and analysed for DCTB using the approach outlined in Fig. 4A. As reported previously[3](/articles/s41598-021-86054-z#ref-CR3 "Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540. https://doi.org/10.1164/rccm.201604-0769OC
(2016)."), we were able to demonstrate that inclusion of _Mtb_ CF resulted in detection of higher number of specimens with DCTB (24/36), together with a higher DCTB quantum when compared to using media with no CF (13/36), Fig. [4](/articles/s41598-021-86054-z#Fig4)B. We next sought to determine if addition of Rpfs from _Mtb_ to _Msm_ would yield a CF with comparable growth stimulatory properties as that obtained from _Mtb_. The genome of _Msm_ encodes four _rpf_\-like genes, _rpfA_, _rpfB_, _rpfE1_ and _rpfE2_[8](/articles/s41598-021-86054-z#ref-CR8 "Machowski, E. E., Senzani, S., Ealand, C. & Kana, B. D. Comparative genomics for mycobacterial peptidoglycan remodelling enzymes reveals extensive genetic multiplicity. BMC Microbiol. 14, 75.
https://doi.org/10.1186/1471-2180-14-75
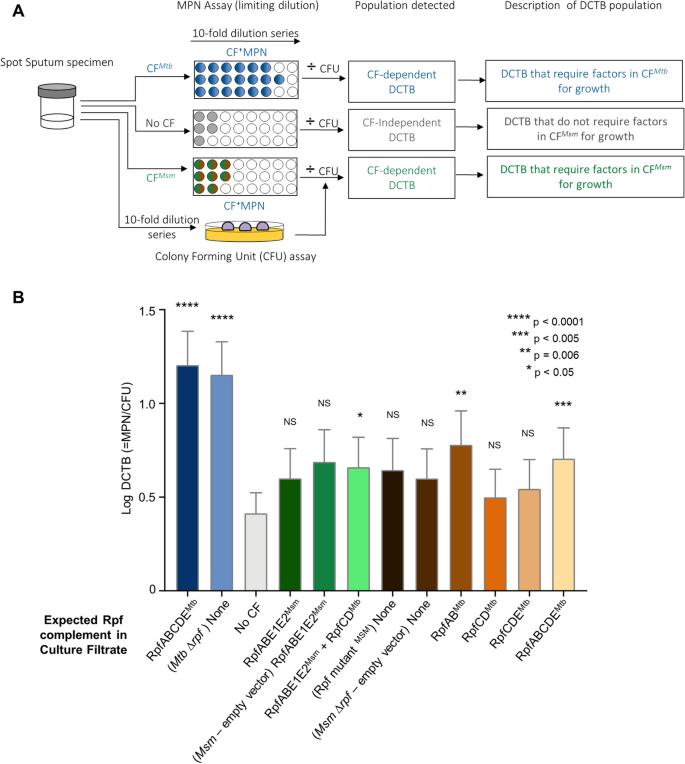
(2014)."). As there were no counterparts for _rpfC_ _Mtb_ and _rpfD_ _Mtb_ in _Msm_, we first inserted these two genes into wild type _Msm_ to create a CF that would contain all five mycobacterial Rpfs, RpfABCDE, by combining _Msm_ and _Mtb_ homologues (albeit with a duplicate copy of _rpfE1_ \[_rpfE2_\]). When compared to the no CF control, inclusion of CF from wild type _Msm_ in MPN assays did not yield a comparable increase in DCTB quantum and appeared to be indistinguishable from assays with no CF. The addition of RpfCD_Mtb_ to _Msm_ yielded a significant increase in the mean quantum of DCTB, Fig. [4](/articles/s41598-021-86054-z#Fig4)B, but this was not associated with detection of DCTB in more specimens when compared to the no CF control (both yielding 13/36 DCTB positive specimens).Figure 4
The alternative text for this image may have been generated using AI.
Bacterial yield in DCTB assays using CF from Msm strains expressing Mtb Rpfs in various combinations. (A) Assessment of DCTB recovery from sputum cultures using CF supplementation. DCTB were detected using the MPN limiting dilution assay. For comparison, CF containing and deficient in Mtb and Msm Rpf was used to assess the role of Rpfs in growth stimulation. To control for the effect of CF in growth stimulation, fresh Middlebrook media was used. CF from various recombinant Msm strains was used to assess if expression of Mtb Rpfs in Msm yields CF that can be used to stimulate DCTB in sputum specimens in a manner that is comparable to Mtb CF. To obtain the DCTB count, MPN values (with or without CF) were divided by CFU counts, which depict conventionally culturable bacteria. (B) Histogram depicting mean DCTB counts from assays using CF from Mtb and Msm. The mean DCTB counts were compared to that obtained from no CF assays using a Wilcoxon test to determine the growth stimulatory effect of the various CFs. Error bars depict the standard error of the mean. P-values depict the comparison of any given CF to the no CF or empty vector control. Graph was generated using Graphpad Prism software, Version 6.
Following this, we next asked if CF derived from Msm containing only Mtb Rpfs was able to stimulate growth of DCTB, as Msm would be a more tractable source of Rpfs in the diagnostic setting. For this, we used our previously reported mutant of Msm deleted for all four _Msm rpf_-like genes[9](/articles/s41598-021-86054-z#ref-CR9 "Ealand, C. et al. Resuscitation-promoting factors are required for Mycobacterium smegmatis biofilm formation. Appl. Environ. Microbiol. 8, 4. https://doi.org/10.1128/AEM.00687-18
(2018)."), _Msm_ Δ_rpf_, and inserted _Mtb rpf_ genes into this genetic background in different combinations. We found that only the addition of RpfABMtb and RpfABCDEMtb to _Msm_ Δ_rpf_ yielded a marginal but statistically significant increase in the quantum of DCTB recovered, together with a statistically non-significant increase in the number of positive specimens detected (15/36 for both compared to 13/36 for the no CF control). However, whilst the mean DCTB values for CF containing RpfABMtb and RpfABCDEMtb were higher than the empty vector control (mean DCTB yield of 0.8 log and 0.7 log for RpfABMtb and RpfABCDEMtb compared to 0.6 log for the empty vector control), the difference was not statistically significant. Addition of RpfCDMtb and RpfDCEMtb to the _Msm_ Δ_rpf_ mutant did not yield CF preparations that were able to stimulate more DCTB when compared to the no CF control (Fig. [4](/articles/s41598-021-86054-z#Fig4)B).Data from our heterologous complementation experiments suggested that addition of Mtb Rpfs to Msm does not yield growth stimulatory effects comparable to that seen with the Mtb CF, suggesting that other factors could also contribute to the growth stimulatory effect. To further investigate this, we compared DCTB yields using Mtb CF derived from wild type Mtb or a mutant defective for the five _rp_f genes. There was no statistical difference in the quantum of DCTB detected with CF derived from these two strains.
Assessment of the ability of fatty acids and cyclic-AMP (cAMP) to resuscitate DCTB
We next set out to evaluate whether other molecules present in the CF are able to contribute to the growth stimulatory effect seen with sputum specimens and tested the ability of free fatty acids and cAMP to stimulate DCTB. For this component of the study, we obtained sputum (n = 21) from a second collection of patients with drug susceptible TB of whom, 62% were HIV-coinfected (Table 1). The participant disposition flow chart and demographics, together with laboratory data, are shown in Fig. 3B and Table 1 respectively. Twenty specimens were smear positive, the majority had high or medium bacterial load as assessed by GeneXpert, with a median time to positivity on MGIT culture of 6 days. Of these, 4 specimens were used to interrogate the effect of cAMP and select fatty acids, whilst the remainder were used for further analysis of a broader range of fatty acids.
In prior work, cAMP appeared to facilitate recovery of DCTB from an in vitro model of differential culturabilty, together with fatty acids such as arachidonic acid, whereas addition of stearic acid did not yield the same effect[10](/articles/s41598-021-86054-z#ref-CR10 "Shleeva, M. et al. Cyclic AMP-dependent resuscitation of dormant Mycobacteria by exogenous free fatty acids. PLoS ONE 8, e82914. https://doi.org/10.1371/journal.pone.0082914
(2013)."). Hence, we first tested these agents (as outlined in Fig. [5](/articles/s41598-021-86054-z#Fig5)A) in a small selection of sputum specimens and found that neither cAMP nor arachidonic acid facilitated recovery of DCTB in a manner comparable to _Mtb_ CF (Fig. [5](/articles/s41598-021-86054-z#Fig5)B). We also noted that stearic acid did not enhance recovery of DCTB (Fig. [5](/articles/s41598-021-86054-z#Fig5)B). Following this, we opted to test a broader range of fatty acids, with the hypothesis that sputum resident organisms may respond to different stimuli when compared to differentially culturable bacteria generated in vitro. We chose a variety of chemically distinct molecules and first optimised the concentrations using the previously reported in vitro model of _M. smegmatis_ differential culturability[14](/articles/s41598-021-86054-z#ref-CR14 "Shleeva, M., Mukamolova, G. V., Young, M., Williams, H. D. & Kaprelyants, A. S. Formation of “non-culturable” cells of Mycobacterium smegmatis in stationary phase in response to growth under suboptimal conditions and their Rpf-mediated resuscitation. Microbiology (Reading) 150, 1687–1697.
https://doi.org/10.1099/mic.0.26893-0
(2004)."). The concentration that yielded the best bacterial recovery (detailed in Table [S1](/articles/s41598-021-86054-z#MOESM1)) were then applied to sputum DCTB assays (Fig. [5](/articles/s41598-021-86054-z#Fig5)A). Certain fatty acids such as palmitoleic, petroselenic and linolenic acid yielded a marginal, statistically non-significant increase in bacterial recovery when compared to media alone (Fig. [5](/articles/s41598-021-86054-z#Fig5)C). However, no fatty acid yielded growth stimulation in a manner comparable to CF from _Mtb_. We noted that in this collection of sputum specimens, the yield of DCTB with CF from _Mtb_ was higher when compared to yields shown in Figure 4B. As these specimens were collected from two distinct cohorts of individuals, at different times, these differences may relate to sputum collection procedures or individual characteristics of the participants. Figure 5
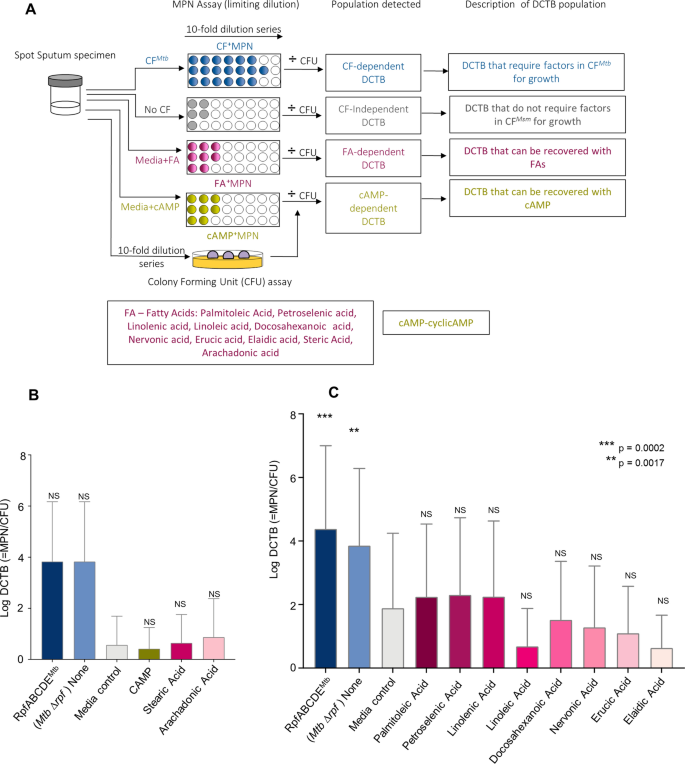
The alternative text for this image may have been generated using AI.
Bacterial yield in DCTB assays using CF containing Fatty acids and cyclic-AMP. (A) Assessment of DCTB recovery from sputum cultures using CF, fatty acid and cyclic-AMP (CAMP) supplementation. DCTB were detected using the MPN limiting dilution assay. For comparison, CF containing and deficient in Mtb Rpfs was used to compare the growth stimulatory effects of fatty acids and cAMP. To control for the effect of CF in growth stimulation, fresh Middlebrook media was used, which standardly contains oleic acid. To obtain the DCTB count, MPN values (with or without CF) were divided by CFU counts, which depict conventionally culturable bacteria. (B) Histogram depicting mean DCTB counts from MPN assays supplemented with standard media containing cAMP or select fatty acids. Statistical comparisons were conducted between CFs and the no CF or cAMP/fatty acid assays. There was an increase in bacterial recovery from MPN assays containing CF, which was not significant, most likely due to the small samples size (n = 4), no significant increase in DCTB recovery was noted in the media control and cAMP or fatty acid assays. Error bars depict the standard error of the mean. (C) Histogram depicting mean DCTB counts from assays using CF from Mtb and standard media supplemented with fatty acids_._ The mean DCTB counts were compared to that obtained from the media controls assay using a Wilcoxon test to determine the growth stimulatory affect of fatty acids. P-values depict the comparison of the CFs to the no CF or fatty acid assays. Graphs were generated using Graphpad Prism software, Version 6.
Discussion
A growing number of studies point to the presence of a differentially culturable population of tubercle bacteria in sputum which require CF to grow in liquid or on solid media[3](/articles/s41598-021-86054-z#ref-CR3 "Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540. https://doi.org/10.1164/rccm.201604-0769OC
(2016)."),[5](/articles/s41598-021-86054-z#ref-CR5 "Garton, N. J. et al. Cytological and transcript analyses reveal fat and lazy persister-like bacilli in tuberculous sputum. PLoS Med. 5, e75.
https://doi.org/10.1371/journal.pmed.0050075
(2008)."),[15](/articles/s41598-021-86054-z#ref-CR15 "Dusthackeer, A. et al. Differential culturability of Mycobacterium tuberculosis in culture-negative sputum of patients with pulmonary tuberculosis and in a simulated model of dormancy. Front. Microbiol. 10, 2381.
https://doi.org/10.3389/fmicb.2019.02381
(2019)."),[16](/articles/s41598-021-86054-z#ref-CR16 "Saito, K. et al. Rifamycin action on RNA polymerase in antibiotic-tolerant Mycobacterium tuberculosis results in differentially detectable populations. Proc. Natl. Acad. Sci. U. S. A. 114, E4832–E4840.
https://doi.org/10.1073/pnas.1705385114
(2017)."), this growth stimulatory effect of CF has been ascribed to the presence of Rpfs[6](/articles/s41598-021-86054-z#ref-CR6 "Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G. & Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle Bacilli in Sputum. Am. J. Respir. Crit. Care Med. 181, 174–180.
https://doi.org/10.1164/rccm.200905-0661OC
(2009)."),[15](/articles/s41598-021-86054-z#ref-CR15 "Dusthackeer, A. et al. Differential culturability of Mycobacterium tuberculosis in culture-negative sputum of patients with pulmonary tuberculosis and in a simulated model of dormancy. Front. Microbiol. 10, 2381.
https://doi.org/10.3389/fmicb.2019.02381
(2019)."). A single Rpf was identified in _Micrococcus luteus_ as an essential secreted protein able to stimulate the growth of dormant bacterial cultures and represented a group of bacterial growth modulators[17](/articles/s41598-021-86054-z#ref-CR17 "Mukamolova, G. V. et al. The rpf gene of Micrococcus luteus encodes an essential secreted growth factor. Mol. Microbiol. 46, 611–621.
https://doi.org/10.1046/j.1365-2958.2002.03183.x
(2002)."). Subsequent studies have highlighted a role for these proteins in exit from dormancy in vitro and during virulence and reactivation in the murine model of TB infection[18](#ref-CR18 "Russell-Goldman, E., Xu, J., Wang, X., Chan, J. & Tufariello, J. M. A Mycobacterium tuberculosis Rpf double-knockout strain exhibits profound defects in reactivation from chronic tuberculosis and innate immunity phenotypes. Infect. Immun. 76, 4269–4281.
https://doi.org/10.1128/IAI.01735-07
(2008)."),[19](#ref-CR19 "Tufariello, J. M. et al. Deletion of the Mycobacterium tuberculosis resuscitation-promoting factor Rv1009 gene results in delayed reactivation from chronic tuberculosis. Infect. Immun. 74, 2985–2995.
https://doi.org/10.1128/IAI.74.5.2985-2995.2006
(2006)."),[20](#ref-CR20 "Downing, K. J. et al. Mutants of Mycobacterium tuberculosis lacking three of the five rpf-like genes are defective for growth in vivo and for resuscitation in vitro. Infect. Immun. 73, 3038–3043.
https://doi.org/10.1128/IAI.73.5.3038-3043.2005
(2005)."),[21](/articles/s41598-021-86054-z#ref-CR21 "Kana, B. D. et al. The resuscitation-promoting factors of Mycobacterium tuberculosis are required for virulence and resuscitation from dormancy but are collectively dispensable for growth in vitro. Mol. Microbiol. 67, 672–684.
https://doi.org/10.1111/j.1365-2958.2007.06078.x
(2008)."). It has been shown that these proteins exert their effects through breakdown of the peptidoglycan component of the cell wall, leading to the release of growth stimulating muropeptides suggesting that the growth stimulatory effect can be applied to bacterial cultures in a paracrine manner, through supplementation of growth media with the Rpf proteins themselves[22](/articles/s41598-021-86054-z#ref-CR22 "Nikitushkin, V. D. et al. A product of RpfB and RipA joint enzymatic action promotes the resuscitation of dormant mycobacteria. FEBS J. 282, 2500–2511.
https://doi.org/10.1111/febs.13292
(2015)."),[23](/articles/s41598-021-86054-z#ref-CR23 "Nikitushkin, V. D., Demina, G. R., Shleeva, M. O. & Kaprelyants, A. S. Peptidoglycan fragments stimulate resuscitation of “non-culturable” mycobacteria. Antonie Van Leeuwenhoek 103, 37–46.
https://doi.org/10.1007/s10482-012-9784-1
(2013)."). Consistent with this, it has been demonstrated that mycobacterial Rpfs are secreted into the culture media hence, making the CF amenable for the identification of DCTB[13](/articles/s41598-021-86054-z#ref-CR13 "Mukamolova, G. V. et al. A family of autocrine growth factors in Mycobacterium tuberculosis. Mol. Microbiol. 46, 623–635.
https://doi.org/10.1046/j.1365-2958.2002.03184.x
(2002).").In a previous study we demonstrated the utility of CF derived from Mtb to detect DCTB in individuals from whom the sputum specimen yielded negative cultures using standard techniques[3](/articles/s41598-021-86054-z#ref-CR3 "Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540. https://doi.org/10.1164/rccm.201604-0769OC
(2016)."). This suggested that CF-supplementation in the diagnostic setting could improve TB detection in difficult to diagnose cases with paucibacillary TB however, use of _Mtb_ CF is limited by the requirement to perform these processes in containment laboratories and need for expert skills to culture the pathogen with subsequent separation of the CF from the bacteria. In this context, _Msm_ provide a tractable avenue to generate CF but it remained unclear if this would yield a similar growth stimulatory effect. As the genome of _Msm_ lacks homologues for _rpfC_ and _rpfD_, we first added the _rpfC_ _Mtb_ and _rpfD_ _Mtb_ genes into the genome of wild type _Msm_, to create a CF that was comparable to that derived from _Mtb_. Thereafter, we tested various combinations of _Mtb rpf_\-genes in the _Msm_ Δ_rpf_ mutant for growth stimulatory capabilities. None of the CF preparations from _Msm_ yielded growth stimulatory effects that were comparable to CF from _Mtb_.The addition of CF containing RpfAB_Mtb_ and RpfABCDE_Mtb_ yielded a small but significant increase in recovery of DCTB and given that addition of CF with RpfCDE_Mtb_ did not yield comparable effects, the increased growth stimulatory effect by the former CF may be due to RpfAB_Mtb_. These results should be interpreted with caution as in cases where addition of select Mtb Rpfs appeared to significantly increase the yield of DCTB compared to the no CF control, these differences were not significantly different when compared to CF from the no vector control strain. In our prior work, we demonstrated that the combinatorial effect of RpfA and RpfB from Msm played an important role in biofilm formation[9](/articles/s41598-021-86054-z#ref-CR9 "Ealand, C. et al. Resuscitation-promoting factors are required for Mycobacterium smegmatis biofilm formation. Appl. Environ. Microbiol. 8, 4. https://doi.org/10.1128/AEM.00687-18
(2018)."), suggesting that these two proteins may have an essential function in bacterial communication.In addition to the Rpf proteins, cAMP and fatty acids have also been reported to stimulate the growth of dormant Msm[10](/articles/s41598-021-86054-z#ref-CR10 "Shleeva, M. et al. Cyclic AMP-dependent resuscitation of dormant Mycobacteria by exogenous free fatty acids. PLoS ONE 8, e82914. https://doi.org/10.1371/journal.pone.0082914
(2013)."). These factors in CF, together with other as yet unidentified molecules may work in concert to allow for paracrine stimulation of DCTB in sputum specimens. We tested the effect of cAMP and a variety of fatty acids and whilst these appear to differentially facilitate recovery of differentially culturable _Msm_ in vitro, they had no benefit in recovery of DCTB in sputum. When combined with the observations on recombinant CFs (with and without _Mtb_ Rpfs), our data suggests that the growth stimulatory effect observed in sputum specimens cultured with _Mtb_ CF is most likely the result of a combination of factors. Further work to identify the interplay between these factors and how they enhance bacterial growth may yield useful new supplements for diagnostic media and would facilitate the development of faster culture-based diagnostics for TB.Materials and methods
All methods were performed in accordance with the relevant guidelines and regulations for growth of Mtb and handling of human specimens. All procedures were conducted in a BioSafety Level III laboratory, registered with the South African Department of Agriculture Forestry and Fisheries (Registration Number: 39.2/NHLS-20/010). All procedures were approved by the Institutional BioSafety Committee of the University of the Witwatersrand (approval number: 20200502Lab).
Bacterial strains and culture conditions
Bacterial strains and plasmids used in this study are listed in Table 2. E. coli and M. smegmatis strains were grown as previously described[9](/articles/s41598-021-86054-z#ref-CR9 "Ealand, C. et al. Resuscitation-promoting factors are required for Mycobacterium smegmatis biofilm formation. Appl. Environ. Microbiol. 8, 4. https://doi.org/10.1128/AEM.00687-18
(2018).").Table 2 Plasmids and strains used in this study.
Cloning of shuttle plasmids for integration into M. smegmatis
To prevent spontaneous excision of the integrated vector in the absence of antibiotic selection, previously generated integrating vectors carrying the rpf genes[21](/articles/s41598-021-86054-z#ref-CR21 "Kana, B. D. et al. The resuscitation-promoting factors of Mycobacterium tuberculosis are required for virulence and resuscitation from dormancy but are collectively dispensable for growth in vitro. Mol. Microbiol. 67, 672–684. https://doi.org/10.1111/j.1365-2958.2007.06078.x
(2008).") were manipulated to remove the integrase gene. Plasmid pMV-RPFAB was digested with _Mlu_I and _Bst_BI and the 6313 bp blunt ended fragment was cloned into pTTP1B at the _Acc_65I site to generate pTT-RPFAB. The resulting plasmid was subsequently partially digested with _Nco_I, and the 10,937 bp fragment was circularised to yield plasmid pTT-RPFABΔi (Table [2](/articles/s41598-021-86054-z#Tab2)). Plasmids pH-RPFCD and pH-RPFCDE[21](/articles/s41598-021-86054-z#ref-CR21 "Kana, B. D. et al. The resuscitation-promoting factors of Mycobacterium tuberculosis are required for virulence and resuscitation from dormancy but are collectively dispensable for growth in vitro. Mol. Microbiol. 67, 672–684.
https://doi.org/10.1111/j.1365-2958.2007.06078.x
(2008).") were digested with _Hin_dIII/_Bam_HI and _Hin_dIII/_Bsr_GI respectively and the resulting 9496 bp and 10,907 bp fragments were blunt ended and circularised to yield plasmids pH-RPFCDΔi and pH-RPFCDEΔi (Table [2](/articles/s41598-021-86054-z#Tab2)). The empty vector control, pHINT-attP-hyg, containing no _rpf_ genes was generated by digesting pHINT with _Hin_dIII/_Nde_I and the resulting 4781 bp fragment, was blunted and circularised (Table [2](/articles/s41598-021-86054-z#Tab2)). The integrase protein was provided in _trans_, by generating suicide plasmids from pHINT and pTTP1B which lack the phage attachment sites _attP_L5 and _attP_Tweety, respectively (Table [2](/articles/s41598-021-86054-z#Tab2)). The pHINT was digested with _Eco_RI/ _Nde_I, and pTTP1B was digested with _Hin_dIII and partially with _Acc_I to generate the 4204 bp fragment and the 4533 bp fragment respectively. These fragments were blunt ended and circularised to yield plasmid pHINT-int-bla and pTTP1B-int-bla (Table [2](/articles/s41598-021-86054-z#Tab2)). These suicide vectors bearing the integrase gene of either pHINT or pTTP1B were co-electroporated with the conditionally integrating plasmids carrying the _rpf_ genes in various combinations into _Msm_ and the _Msm_ Δ_rpf_ mutant to generate recombinant strains of _Msm_ carrying various combinations of _Mtb rpf_ genes, detailed in Table [2](/articles/s41598-021-86054-z#Tab2). Hygromycin or kanamycin resistant transformants were picked and screened for expression of the corresponding _Msm_ and _Mtb rpf_ genes by RT-PCR using the primers listed in Table [S1](/articles/s41598-021-86054-z#MOESM1).Gene expression analysis by real time, qRT-PCR
To monitor the expression of the rpf genes, all recombinant strains were grown to early exponential phase (OD600nm 0.6). Gene expression analysis was conducted as previously described[9](/articles/s41598-021-86054-z#ref-CR9 "Ealand, C. et al. Resuscitation-promoting factors are required for Mycobacterium smegmatis biofilm formation. Appl. Environ. Microbiol. 8, 4. https://doi.org/10.1128/AEM.00687-18
(2018).") using primers indicated in Table [S2](/articles/s41598-021-86054-z#MOESM1).Recruitment of participants to obtain sputum specimens
Ethics approval for participant recruitment was obtained from the Human Research Ethics Committee of the University of the Witwatersrand, South Africa, with clearance number M110833, subsequently revised to M200164 (SNT study). Individuals 18 years and older attending the Perinatal HIV Research Unit (PHRU), Soweto, South Africa were approached for participation into the study. Fifty seven HIV sero-negative patients with drug sensitive TB, as determined from either an auramine stained smear or by GeneXpert obtained from the public sector (National Health Laboratory Service, Johannesburg, South Africa) were eligible for enrolment in the study. Patients willing to participate were approached and written informed consent was obtained using Informed Consent Forms reviewed and approved by the Human Research Ethics Committee of the University of the Witwatersrand. After written informed consent was granted, a spot sputum was collected for analysis by the most probable number (MPN) assay (otherwise referred to as limiting dilution assays) as described previously[3](/articles/s41598-021-86054-z#ref-CR3 "Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540. https://doi.org/10.1164/rccm.201604-0769OC
(2016)."). The MPN assay is a limiting dilution series based on a Poisson distribution for the quantification of bacterial growth in liquid media, described in further detail in the Supplementary Information. Sputum samples were serially diluted in a 48 well microtitre plate in media with culture filtrate (1:1 ratio) purified from wild type _Msm, Mtb_ or a variety of mutant/recombinant strains. Where relevant, fatty acids or CAMP were added to media in the MPN assay.Optimization of fatty acid concentrations to use in sputum DCTB assays
The concentrations of fatty acids for use in sputum were optimised using an in vitro model of dormancy that generates differentially culturable Msm as previously described[14](/articles/s41598-021-86054-z#ref-CR14 "Shleeva, M., Mukamolova, G. V., Young, M., Williams, H. D. & Kaprelyants, A. S. Formation of “non-culturable” cells of Mycobacterium smegmatis in stationary phase in response to growth under suboptimal conditions and their Rpf-mediated resuscitation. Microbiology (Reading) 150, 1687–1697. https://doi.org/10.1099/mic.0.26893-0
(2004)."). _Msm_ was pre-cultured in 20 ml of nutrient rich broth for 16 h at 37 °C in an orbital shaker (250 rpm). Pre-cultures were sub-cultured into modified Hartman’s–de Bont medium. Trace elements solution was prepared as previously described[24](/articles/s41598-021-86054-z#ref-CR24 "Smeulders, M. J., Keer, J., Speight, R. A. & Williams, H. D. Adaptation of Mycobacterium smegmatis to stationary phase. J. Bacteriol. 181, 270–283.
https://doi.org/10.1128/JB.181.1.270-283.1999
(1999)."). The OD600nm of the pre-culture was adjusted to 0.9 and 20 ml was added to 130 ml of the Hartmans de Bont media and incubated at 37 °C for 14 days without shaking. The unsaturated fatty acids kit (Sigma Catalogue number UN10-1KT) was used for the resuscitation of differentially culturable _Msm_. Each fatty acid was initially resuspended in 1 ml of dimethyl sulfoxide to create a working stock. The starved _Msm_ was added to Middlebrook media (supplemented with ADC—no oleic acid was used) and the OD600nm adjusted to 0.1\. Thereafter, 9 ml aliquots of these cells was used to test the different concentrations of fatty acids. Fatty acids were tested at the following concentrations: 0.125 µM, 0.25 µM, 0.5 µM, 1 µM, 2.5 µM, 5 µM, 7.5 µM, 10 µM, 12.5 µM, 15 µM, 17.5 µM, 20 µM, 22.5 µM, 25 µM, 27.5 µM, 30 µM and 32.5 µM. The OD600nm of each culture was assessed after 72 h and the bacterial recovery was measure as the fold increased in OD600nm compared to the inoculum. For cAMP (3 mM), steric acid (4 µM), arachinodic acid (1.6 µM) and Linoleic acid (1.7 µM) the concentrations used were derived from previous similar work[10](/articles/s41598-021-86054-z#ref-CR10 "Shleeva, M. et al. Cyclic AMP-dependent resuscitation of dormant Mycobacteria by exogenous free fatty acids. PLoS ONE 8, e82914.
https://doi.org/10.1371/journal.pone.0082914
(2013).").Data analysis
MPN values from assays containing distinct CF preparations were compared in a pairwise fashion using a student’s t test. All statistical analysis was done using GraphPad Prism software, version 6.
References
- WHO. Global Tuberculosis Report 2019 (World Health Organization, Geneva, 2019).
Google Scholar - Dheda, K., Barry, C. E. 3rd. & Maartens, G. Tuberculosis. Lancet 387, 1211–1226. https://doi.org/10.1016/S0140-6736(15)00151-8 (2016).
Article PubMed Google Scholar - Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540. https://doi.org/10.1164/rccm.201604-0769OC (2016).
Article CAS PubMed PubMed Central Google Scholar - Dartois, V., Saito, K., Warrier, T. & Nathan, C. New evidence for the complexity of the population structure of Mycobacterium tuberculosis increases the diagnostic and biologic challenges. Am. J. Respir. Crit. Care Med. 194, 1448–1451. https://doi.org/10.1164/rccm.201607-1431ED (2016).
Article PubMed PubMed Central Google Scholar - Garton, N. J. et al. Cytological and transcript analyses reveal fat and lazy persister-like bacilli in tuberculous sputum. PLoS Med. 5, e75. https://doi.org/10.1371/journal.pmed.0050075 (2008).
Article CAS PubMed PubMed Central Google Scholar - Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G. & Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle Bacilli in Sputum. Am. J. Respir. Crit. Care Med. 181, 174–180. https://doi.org/10.1164/rccm.200905-0661OC (2009).
Article CAS PubMed PubMed Central Google Scholar - Kana, B. D. & Mizrahi, V. Resuscitation-promoting factors as lytic enzymes for bacterial growth and signaling. FEMS Immunol. Med. Microbiol. 58, 39–50. https://doi.org/10.1111/j.1574-695X.2009.00606.x (2010).
Article CAS PubMed Google Scholar - Machowski, E. E., Senzani, S., Ealand, C. & Kana, B. D. Comparative genomics for mycobacterial peptidoglycan remodelling enzymes reveals extensive genetic multiplicity. BMC Microbiol. 14, 75. https://doi.org/10.1186/1471-2180-14-75 (2014).
Article PubMed PubMed Central Google Scholar - Ealand, C. et al. Resuscitation-promoting factors are required for Mycobacterium smegmatis biofilm formation. Appl. Environ. Microbiol. 8, 4. https://doi.org/10.1128/AEM.00687-18 (2018).
Article Google Scholar - Shleeva, M. et al. Cyclic AMP-dependent resuscitation of dormant Mycobacteria by exogenous free fatty acids. PLoS ONE 8, e82914. https://doi.org/10.1371/journal.pone.0082914 (2013).
Article ADS CAS PubMed PubMed Central Google Scholar - O’Gaora, P. et al. Mycobacteria as immunogens: development of expression vectors for use in multiple mycobacterial species. Med. Princ. Pract. 6, 91–96 (1997).
Article Google Scholar - Pham, T. T., Jacobs-Sera, D., Pedulla, M. L., Hendrix, R. W. & Hatfull, G. F. Comparative genomic analysis of mycobacteriophage Tweety: evolutionary insights and construction of compatible site-specific integration vectors for mycobacteria. Microbiology (Reading) 153, 2711–2723. https://doi.org/10.1099/mic.0.2007/008904-0 (2007).
Article CAS Google Scholar - Mukamolova, G. V. et al. A family of autocrine growth factors in Mycobacterium tuberculosis. Mol. Microbiol. 46, 623–635. https://doi.org/10.1046/j.1365-2958.2002.03184.x (2002).
Article CAS PubMed Google Scholar - Shleeva, M., Mukamolova, G. V., Young, M., Williams, H. D. & Kaprelyants, A. S. Formation of “non-culturable” cells of Mycobacterium smegmatis in stationary phase in response to growth under suboptimal conditions and their Rpf-mediated resuscitation. Microbiology (Reading) 150, 1687–1697. https://doi.org/10.1099/mic.0.26893-0 (2004).
Article CAS Google Scholar - Dusthackeer, A. et al. Differential culturability of Mycobacterium tuberculosis in culture-negative sputum of patients with pulmonary tuberculosis and in a simulated model of dormancy. Front. Microbiol. 10, 2381. https://doi.org/10.3389/fmicb.2019.02381 (2019).
Article PubMed PubMed Central Google Scholar - Saito, K. et al. Rifamycin action on RNA polymerase in antibiotic-tolerant Mycobacterium tuberculosis results in differentially detectable populations. Proc. Natl. Acad. Sci. U. S. A. 114, E4832–E4840. https://doi.org/10.1073/pnas.1705385114 (2017).
Article CAS PubMed PubMed Central Google Scholar - Mukamolova, G. V. et al. The rpf gene of Micrococcus luteus encodes an essential secreted growth factor. Mol. Microbiol. 46, 611–621. https://doi.org/10.1046/j.1365-2958.2002.03183.x (2002).
Article CAS PubMed Google Scholar - Russell-Goldman, E., Xu, J., Wang, X., Chan, J. & Tufariello, J. M. A Mycobacterium tuberculosis Rpf double-knockout strain exhibits profound defects in reactivation from chronic tuberculosis and innate immunity phenotypes. Infect. Immun. 76, 4269–4281. https://doi.org/10.1128/IAI.01735-07 (2008).
Article CAS PubMed PubMed Central Google Scholar - Tufariello, J. M. et al. Deletion of the Mycobacterium tuberculosis resuscitation-promoting factor Rv1009 gene results in delayed reactivation from chronic tuberculosis. Infect. Immun. 74, 2985–2995. https://doi.org/10.1128/IAI.74.5.2985-2995.2006 (2006).
Article CAS PubMed PubMed Central Google Scholar - Downing, K. J. et al. Mutants of Mycobacterium tuberculosis lacking three of the five _rpf_-like genes are defective for growth in vivo and for resuscitation in vitro. Infect. Immun. 73, 3038–3043. https://doi.org/10.1128/IAI.73.5.3038-3043.2005 (2005).
Article CAS PubMed PubMed Central Google Scholar - Kana, B. D. et al. The resuscitation-promoting factors of Mycobacterium tuberculosis are required for virulence and resuscitation from dormancy but are collectively dispensable for growth in vitro. Mol. Microbiol. 67, 672–684. https://doi.org/10.1111/j.1365-2958.2007.06078.x (2008).
Article ADS CAS PubMed Google Scholar - Nikitushkin, V. D. et al. A product of RpfB and RipA joint enzymatic action promotes the resuscitation of dormant mycobacteria. FEBS J. 282, 2500–2511. https://doi.org/10.1111/febs.13292 (2015).
Article CAS PubMed Google Scholar - Nikitushkin, V. D., Demina, G. R., Shleeva, M. O. & Kaprelyants, A. S. Peptidoglycan fragments stimulate resuscitation of “non-culturable” mycobacteria. Antonie Van Leeuwenhoek 103, 37–46. https://doi.org/10.1007/s10482-012-9784-1 (2013).
Article CAS PubMed Google Scholar - Smeulders, M. J., Keer, J., Speight, R. A. & Williams, H. D. Adaptation of Mycobacterium smegmatis to stationary phase. J. Bacteriol. 181, 270–283. https://doi.org/10.1128/JB.181.1.270-283.1999 (1999).
Article CAS PubMed PubMed Central Google Scholar - Snapper, S. B., Melton, R. E., Mustafa, S., Kieser, T. & Jacobs, W. R. Jr. Isolation and characterization of efficient plasmid transformation mutants of Mycobacterium smegmatis. Mol. Microbiol. 4, 1911–1919 (1990).
Article CAS Google Scholar
Acknowledgements
This work was supported by funding from the South African National Research Foundation (to BDK and CE), the South African Medical Research Council (to BDK and CE), the National Health Laboratory Service Research Trust (to BGG) and the Centre for Aids Prevention Research in South Africa (CAPRISA, to CE). We thank the participants of this study.
Author information
Author notes
- These authors contributed equally: Bhavna G. Gordhan, Julian S. Peters, Amanda McIvor and Edith E. Machowski
Authors and Affiliations
- Department of Science and Technology/National Research Foundation Centre of Excellence for Biomedical TB Research, School of Pathology, Faculty of Health Sciences, University of the Witwatersrand and the National Health Laboratory Service, P. O. Box 1038, Johannesburg, 2000, South Africa
Bhavna G. Gordhan, Julian S. Peters, Amanda McIvor, Edith E. Machowski, Christopher Ealand, Neil Martinson & Bavesh D. Kana - Perinatal HIV Research Unit, Faculty of Health Sciences, University of the Witwatersrand, Johannesburg, South Africa
Ziyaad Waja & Neil Martinson - Center for Tuberculosis Research, Johns Hopkins University, Baltimore, MD, USA
Neil Martinson
Authors
- Bhavna G. Gordhan
- Julian S. Peters
- Amanda McIvor
- Edith E. Machowski
- Christopher Ealand
- Ziyaad Waja
- Neil Martinson
- Bavesh D. Kana
Contributions
B.K. conceived the overall concept of the study. B.G.G., J.S.P., A.M., E.E.M. and C.E. executed the laboratory aspects of the study. Z.W. and N.M. recruited participants. B.K. and B.G. wrote the first draft of the manuscript. B.K. provided further input on the concept, critical review and edited various drafts.
Corresponding author
Correspondence toBavesh D. Kana.
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gordhan, B.G., Peters, J.S., McIvor, A. et al. Detection of differentially culturable tubercle bacteria in sputum using mycobacterial culture filtrates.Sci Rep 11, 6493 (2021). https://doi.org/10.1038/s41598-021-86054-z
- Received: 05 November 2020
- Accepted: 01 March 2021
- Published: 22 March 2021
- Version of record: 22 March 2021
- DOI: https://doi.org/10.1038/s41598-021-86054-z