Carrying out streamlined routine data analyses with reports for observational studies: introduction to a... (original) (raw)
Background
The traditional surgical approach for early-stage lung cancer is via a thoracotomy, and such open chest surgery is limited to certain eligible patients due to the increased risk of mortality, especially among elderly patients with multiple comorbidities. Over the past two decades, video-assisted thoracic surgery (VATS) has been increasingly used in clinics and provides excellent short-term advantages over thoracotomy, such as fewer complications, less pain, improved lung function, shorter recovery period and lower costs. On the other hand, incomplete lymph node evaluation with VATS or a high rate of the residual tumor may compromise the long-term efficacy of VATS vs. thoracotomy (open). The knowledge uncertainty still exists regarding the optimal surgical approach for early-stage lung cancer concerning long-term survival16. The goal of this study is to compare the overall survival (OS) between two surgical approaches among the target eligible lung cancer patients.
The National Cancer Database (NCDB)2, a joint project of the American Cancer Society and the Commission on Cancer of the American College of Surgeons, was queried for non-small cell lung cancer patients with clinical stage < T2N0M0 who underwent lobectomy in 2010-2013. After applying the exclusion/inclusion criteria, we identified 34,048 eligible patients. The study cohort is the surgical approach (open vs. minimally invasive). The clinical outcomes of interest were 30-day mortality after surgery (yes vs. no) and overall survival (OS): defined as months from the date of surgery to death or last follow up. Covariates considered included sex, Charlson-Deyo score (comorbidity), year of diagnosis, histology, AJCC clinic T stage, age at diagnosis and urban/rural 2003. For a detailed definition of these variables, users can refer to the online data dictionary (http://ncdbpuf.facs.org/node/259?q=print-pdf-all).
In this case study, we show sequential analytical steps along with interpretations to probe into the study question by using the proposed SAS® macros. To make it easy to follow, we only include a short but relevant list of variables and cover routine data analyses. A comprehensive and complete manuscript for the study question is under preparation for a peer-reviewed publication.
SAS® macros showcase – data analyses
Please note that these macros do not handle data cleaning or management directly. For the most interpretable results, it is highly recommended to properly format and label each variable before running these macros. All output tables are in Rich Text Format (RTF) and editable in Word.
Descriptive Statistics
The macro %DESCRIPTIVE was used to generate Table 1. The table provides an overview of the study population’s basic characteristics, in which the frequency and proportion are listed for categorical covariates and summary statistics (mean, median, min, max, standard deviation) for numerical covariates, along with the frequency of missing values. In the following SAS® code example, a few universal macro variables are implemented: DATASET for the data set name; CLIST for a list of categorical covariates separated by space; NLIST for a list of continuous covariates separated by spaces; OUTPATH for the file path to store the output RTF file; FNAME for the name of the RTF file; DEBUG to suppress deleting intermediate data sets for error checking by setting to T. Setting the DICTIONARY option to T creates two additional columns in the summary table, for the covariates’ SAS® names and unformatted values. These columns are useful for the programmer to connect the table with the dataset and code. In Table 1, we observe that 35.6% of the study population underwent a minimally invasive surgical approach, 45% were male, 80.9% resided in a metro area, and 85.4% had comorbidity score ≤1.
Table 1. Descriptive Statistics for All Variables.
| Variable | Level | N (%) = 34048 |
|---|---|---|
| Surgical Approach | Open | 21911 (64.4) |
| Minimally invasive | 12137 (35.6) | |
| Sex | Male | 15324 (45.0) |
| Female | 18724 (55.0) | |
| Urban/Rural 2003 | Metro | 26833 (80.9) |
| Urban | 5561 (16.8) | |
| Rural | 756 (2.3) | |
| Missing | 898 | |
| Charlson-Deyo Score | 0 | 16637 (48.9) |
| 1 | 12440 (36.5) | |
| 2+ | 4971 (14.6) | |
| Year of Diagnosis | 2010 | 8259 (24.3) |
| 2011 | 8175 (24.0) | |
| 2012 | 8591 (25.2) | |
| 2013 | 9023 (26.5) | |
| Histology | Adenocarcinomas | 21018 (61.7) |
| Other or Unknown | 4395 (12.9) | |
| Squamous cell carcinomas | 8635 (25.4) | |
| AJCC Clinical T | 1 | 22688 (66.6) |
| 2 | 11360 (33.4) | |
| Age at Diagnosis | Mean | 67.02 |
| Median | 68.00 | |
| Minimum | 40.00 | |
| Maximum | 90.00 | |
| Std Dev | 9.55 | |
| Missing | 0.00 |
************ SAS® code example for %DESCRIPTIVE **************************;
/* TIP 1: Create the macro variable &DIR to specify the location used to store all results. If you are conducting an updated analysis, you need to change this path, and all updated results will be saved to a new location without modifying it in every macro. */
%let dir = C:\Desired Location to Save All Results\;
/* TIP 2: Create the macro variables &cat_var and &num_var to store categorical and numerical variable lists outside macros, and reference them in all related macros. All related tables can be updated by changing those two macro variables.
/* TIP 3: Also note that the order of variables in CLIST is the same as the order in which they will appear in the final output table. Listing similar types of variables together, e.g., demographic variables, tumor characteristic variables, lab values, etc., will improve the readability of the results. */
/* Variable Label:
SEX - Sex
UR_CD_03 -- Urban/Rural 2003
CDCC_TOTAL -- Charlson Deyo score (comorbidity)
YEAR_OF_DIAGNOSIS -- Year of Diagnosis
HIS_CAT -- Histology
TNM_CLIN_T -- AJCC Clinical T stage
SURG_APP -- Surgical Approach
*/
%let cat_var = sex UR_CD_03 CDCC_TOTAL YEAR_OF_DIAGNOSIS HIS_CAT TNM_CLIN_T;
%let num_var = age ;
TITLE 'Table 1 Descriptive Statistics for All Variables';
%DESCRIPTIVE(DATASET=anal,
CLIST = surg_app &cat_var,
NLIST = &num_var,
OUTPATH= &dir,
FNAME=Table 1 Descriptive Statistics for All Variables,
DEBUG=F);
TITLE;
**************************************************************************;
Univariate Association with a categorical outcome/exposure variable.
We examined the univariate association between each covariate and surgical approach (Table 2) using %UNI_CAT and 30-day mortality after surgery (Table 3) using %UNI_LOGREG. Each covariate was processed separately, but summarized all together in one table. %UNI_CAT is suitable to compare multiple covariates specified in CLIST and NLIST between two or more cohorts defined by a categorical variable (OUTCOME) (see following SAS® code example). For each categorical covariate, frequencies from a contingency table are reported along with row percentages (Row%) or column percentages (Col%), which can be controlled by the ROWPERCENT option based on the desired interpretation. For each numeric covariate, summary statistics will be generated for each level of OUTCOME. The univariate associations can be tested by either parametric or non-parametric tests using the NONPAR option. The name of the tests will appear in the footnote of the table. Analyses can be performed using logistic regression models if the outcome/exposure variable is binary or ordinal using %UNI_LOGREG. As shown in Table 3, the probability of the EVENT = ‘Yes’ was modeled and the odds ratio (OR) was reported with a 95% confidence interval (CI). The reference level of a categorical covariate in CLIST can be chosen to aid interpretation, which can be done by separating the CLIST variables by an asterisk (*), and then adding “**(DESC)”** or “**(ref = “Reference level in formatted value”)”** after each desired variable name (see following code). If the outcome/explanatory variable is numeric, users can refer to SAS® macro %UNI_NUM. In all report tables, a p-value < 0.05 will be shown in bold for easy visualization.
Table 2. Univariate association with study cohort.
| | Surgical Approach | | | | | | | ------------------- | ------------------------ | --------------- | ------------- | ------------------------- | --------------------------- | | Covariate | Statistics | Level | OpenN=21911 | Minimallyinvasive N=12137 | ParametricP-value* | | Sex | N (Col %) | Male | 10128 (46.22) | 5196 (42.81) | <.001 | | N (Col %) | Female | 11783 (53.78) | 6941 (57.19) | | | | Urban/Rural 2003 | N (Col %) | Metro | 16863 (79.14) | 9970 (84.18) | <.001 | | N (Col %) | Urban | 3894 (18.28) | 1667 (14.08) | | | | N (Col %) | Rural | 550 (2.58) | 206 (1.74) | | | | Charlson-DeyoScore | N (Col %) | 0 | 10630 (48.51) | 6007 (49.49) | 0.014 | | N (Col %) | 1 | 7993 (36.48) | 4447 (36.64) | | | | N (Col %) | 2+ | 3288 (15.01) | 1683 (13.87) | | | | Year of Diagnosis | N (Col %) | 2010 | 6107 (27.87) | 2152 (17.73) | <.001 | | N (Col %) | 2011 | 5436 (24.81) | 2739 (22.57) | | | | N (Col %) | 2012 | 5233 (23.88) | 3358 (27.67) | | | | N (Col %) | 2013 | 5135 (23.44) | 3888 (32.03) | | | | Histology | N (Col %) | Adenocarcinomas | 13186 (60.18) | 7832 (64.53) | <.001 | | N (Col %) | Other or Unknown | 2878 (13.13) | 1517 (12.5) | | | | N (Col %) | Squamous cell carcinomas | 5847 (26.69) | 2788 (22.97) | | | | AJCC Clinical T | N (Col %) | 1 | 14132 (64.5) | 8556 (70.5) | <.001 | | N (Col %) | 2 | 7779 (35.5) | 3581 (29.5) | | | | Age at Diagnosis | N | | 21911 | 12137 | 0.520 | | Mean | | 66.99 | 67.06 | | | | Median | | 68 | 68 | | | | Min | | 40 | 40 | | | | Max | | 90 | 90 | | | | Std Dev | | 9.54 | 9.57 | | |
In Table 2, 42.8% of patients that had minimally invasive surgery were male, and 46.2% among patients that had open surgery. Compared to open surgical patients, minimally invasive surgical patients were more likely to reside in metro areas (84.2% vs. 79.1%), to be diagnosed in recent years, to have a histology of adenocarcinomas (64.5% vs. 60.2%), and to be at clinical T stage 1 (70.5% vs. 64.5%) (all p < 0.001). In Table 3, the factors associated with a higher risk of 30-day mortality after surgery were having open surgery (OR = 1.48), male gender (OR = 2.16), older age (OR = 1.06 for every 1-year increase), a comorbidity score larger than one (OR = 1.8 compared to score = 0), squamous cell carcinomas (OR = 2.49 compared to adenocarcinomas), and clinical T stage 2 (OR = 1.34) (all p < 0.001). By interpreting the information in Table 2 and Table 3 together, the univariate association between surgical type and 30-day mortality could be confounded by the effect from histology and clinical T stage. Open surgery was more likely to be conducted among patients who had squamous cell carcinomas and clinical T2, and these two factors were linked to a higher rate of 30-day mortality. The observed higher probability of 30-day mortality in open surgery patients might be partially due to the imbalanced distribution of histology and clinical T stage in the surgical cohorts. The common statistical approaches to control for confounding effects include multivariable analysis, subgroup analysis, and propensity score methods.
Table 3. Univariate association with 30-day mortality.
| | 30 Day Mortality=Yes | | | | | | | ------------------------ | ---------------- | ---------------- | ------------------ | --------- | ------------ | | Covariate | Level | N | Odds ratio(95% CI) | ORP-value | Type3P-value | | Surgical approach | Open | 21911 | 1.48 (1.23-1.77) | <.001 | <.001 | | Minimally invasive | 12137 | - | - | | | | Sex | Male | 15324 | 2.16 (1.83-2.55) | <.001 | <.001 | | Female | 18724 | - | - | | | | Urban/rural 2003 | Rural | 756 | 1.28 (0.77-2.12) | 0.334 | 0.013 | | Urban | 5561 | 1.34 (1.10-1.64) | 0.004 | | | | Metro | 26833 | - | - | | | | Charlson-Deyo score | 2+ | 4971 | 1.80 (1.46-2.22) | <.001 | <.001 | | 1 | 12440 | 1.09 (0.90-1.31) | 0.373 | | | | 0 | 16637 | - | - | | | | Year of diagnosis | 2010 | 8259 | 1.30 (1.05-1.62) | 0.018 | 0.031 | | 2011 | 8175 | 1.00 (0.79-1.26) | 0.990 | | | | 2012 | 8591 | 0.99 (0.78-1.24) | 0.915 | | | | 2013 | 9023 | - | - | | | | Histology | Other or Unknown | 4395 | 1.13 (0.86-1.48) | 0.393 | <.001 | | Squamous cell carcinomas | 8635 | 2.49 (2.10-2.95) | <.001 | | | | Adenocarcinomas | 21018 | - | - | | | | AJCC clinical T | 2 | 11360 | 1.34 (1.14-1.58) | <.001 | <.001 | | 1 | 22688 | - | - | | | | Age at diagnosis | | 34048 | 1.06 (1.05-1.07) | <.001 | <.001 |
********** SAS® code example for %UNI_CAT and %UNI_LOGREG *************;
/* TIP 4: In the following example, the NONPAR option is turned off because it will take more time to compute Fisher’s exact test on a large sample. */
TITLE 'Table 2 Univariate Association with Study Cohort';
%UNI_CAT (
- DATASET = anal,
- OUTCOME = surg_app,
- CLIST = &cat_var,
- NLIST = &num_var,
- NONPAR = F,
- ROWPERCENT = F,
- SPREAD = T,
- OUTPATH = &dir,
- FNAME =Table 2 Univariate Association with Study Cohort);
/* TIP 5: If you need to specify the reference level of a categorical variable, separate CLIST by “*” and add (DESC) or (ref = “Reference level in formatted value”) after each desired variable name. Otherwise, CLIST can be separated by spaces alone. */
%let cat_var_ref = SEX * UR_CD_03(DESC) * CDCC_TOTAL(DESC) * YEAR_OF_DIAGNOSIS * HIS_CAT(ref="Adenocarcinomas") * TNM_CLIN_T(DESC);
TITLE 'Table 3 Univariate Association with 30-day Mortality';
%UNI_LOGREG(
- DATASET = anal,
- OUTCOME = PUF_30_DAY_MORT_CD,
- EVENT = 'Yes',
- CLIST = surg_app * &cat_var_ref,
- NLIST = &num_var,
- OUTPATH = &dir,
- FNAME = Table 3 Univariate Association with 30-day Mortality);
TITLE;
****************************************************************************;
Univariate Association with a time-to-event outcome
We examined the association of the study cohorts and each covariate with OS using %UNI_PHREG, as shown in Table 4. Each covariate was individually fit in a proportional hazards model (PROC PHREG in SAS®), and the hazard ratios (95% CI) and p-values were summarized in one table. In the following SAS® code example, the survival time variable is specified in EVENT, and the censoring indicator variable in CENSOR. Note that it requires that ‘1’ is used for the event and ‘0’ for the censored cases in the data (DATASET). Other options include LOGRANK to output log-rank p-value, TYPE3 to output the Type3 p-values for categorical covariates, and PHA for proportional hazard assumption checks. Similar to %UNI_LOGREG, one can set the reference level of a categorical covariate in CLIST. %UNI_PHREG can also handle time to event data in the counting process data form by using the START and STOP options, especially when modeling data with time-varying covariates or recurrent event data. It can also handle competing risk data by using the EVENTCODE option to activate the Fine and Gray’s model17.
Table 4. Univariate association with overall survival (OS).
| | Months (OS) | | | | | | | ------------------------ | ---------------- | ---------------- | --------------------- | --------- | ------------ | | Covariate | Level | N | Hazard ratio (95% CI) | HRP-value | Type3P-value | | Surgical approach | Open | 21911 | 1.18 (1.13-1.24) | <.001 | <.001 | | Minimally invasive | 12137 | - | - | | | | Sex | Male | 15324 | 1.68 (1.61-1.76) | <.001 | <.001 | | Female | 18724 | - | - | | | | Urban/rural 2003 | Rural | 756 | 1.23 (1.06-1.42) | 0.006 | <.001 | | Urban | 5561 | 1.19 (1.13-1.27) | <.001 | | | | Metro | 26833 | - | - | | | | Charlson-Deyo score | 2+ | 4971 | 1.77 (1.67-1.89) | <.001 | <.001 | | 1 | 12440 | 1.32 (1.26-1.39) | <.001 | | | | 0 | 16637 | - | - | | | | Year of diagnosis | 2010 | 8259 | 1.00 (0.93-1.08) | 0.900 | 0.922 | | 2011 | 8175 | 1.00 (0.93-1.08) | 0.982 | | | | 2012 | 8591 | 0.98 (0.91-1.06) | 0.648 | | | | 2013 | 9023 | - | - | | | | Histology | Other or unknown | 4395 | 1.05 (0.97-1.13) | 0.211 | <.001 | | Squamous cell carcinomas | 8635 | 1.64 (1.56-1.73) | <.001 | | | | Adenocarcinomas | 21018 | - | - | | | | AJCC clinical T | 2 | 11360 | 1.63 (1.56-1.71) | <.001 | <.001 | | 1 | 22688 | - | - | | | | Age at diagnosis | | 34048 | 1.04 (1.03-1.04) | <.001 | <.001 |
In the univariate analysis shown in Table 4, we see that patients who underwent open surgery had 18% more chance to die before those who underwent minimally invasive surgery (HR = 1.18; 95%CI = (1.13-1.24); p < 0.001). Also, residing in rural or urban areas, higher comorbidity scores, squamous cell carcinomas, clinical stage T2, and older age were all risk factors linked to worse overall survival in this study population. In combination with the results in Table 2, those prognostic factors except for age were also more likely to present in patients who underwent open surgery and should be controlled for as confounders in multivariable analyses.
************ SAS® code example for %UNI_PHREG and its options *****************;
TITLE 'Table 4 Univariate association with Overall Survival';
%UNI_PHREG (
- DATASET = anal,
- EVENT = os_surg,
- CENSOR = os_censor,
- CLIST = surg_app * &cat_var_ref,
- NLIST = &num_var,
- LOGRANK = F,
- TYPE3 = T,
- PHA = F,
- OUTPATH = &dir,
- FNAME = Table 4 Univariate association with Overall Survival);
TITLE;
*****************************************************************************;
Multivariable model for the binary or time-to-event outcome
We performed multivariable analysis with a logistic regression model for the 30-day mortality outcome using %LOGREG_SEL (Table 5) and a Cox proportional hazards model for overall survival using %PHREG_SEL (Table 6). The adjusted association of the surgical approaches with the two clinical outcomes was estimated after controlling for observed confounding variables. The odds ratio and hazard ratio were reported along with 95% CI and p-values. In both macros, a manual backward elimination procedure was implemented by dropping one variable at a time until all variables left satisfied a pre-specified alpha level (e.g., SLSTAY = 0.1). The selection process in the macros allows the sample size to adjust and always uses the maximum available sample as the number of variables in the model drops, which differs from the automatic selection procedure built into PROC LOGISTIC or PROC PHREG, where a complete and fixed data set is used throughout the selection procedure. In the following example code, DSN is for specifying the data set name, and EVENT in %LOGREG_SEL is to specify the event of interest, for which the probability will be modeled. VAR is for the list of all variables or terms in the initial model, separated by a space. CVAR is for the list of the categorical variables in VAR with the option to specify reference levels as shown in Table 4. INC = k is set to protect the first k variables in VAR from being eliminated, such as a primary exploratory variable and important confounding variables. In this case study, we want to keep surgical approach in the model as it is the study cohort, which was done by putting surg_app on the first position in VAR and setting INC = 1. Setting CLNUM = T outputs the frequency of categorical variables in the final model. The summary information of the selection procedure was presented in the footnote of the final report table. A separate macro for multivariable analysis of competing risk data is %FINEGRAY_SEL. In reality, there are many approaches to build a multivariable model. If a user wants to customize the model building process, they can use these two macros for reporting purposes only by setting VAR = final model variables selected by other approaches and INC = total number of items in the final model.
Table 5. Multivariable logistic regression model for 30-day mortality.
| | 30 day mortality=Yes | | | | | | ------------------------ | ---------------- | ------------------ | --------- | ------------ | | Covariate | Level | Odds ratio(95% CI) | ORP-value | Type3P-value | | Surgical approach | Open | 1.41 (1.18-1.69) | <.001 | <.001 | | Minimally invasive | - | - | | | | Sex | Male | 1.84 (1.55-2.18) | <.001 | <.001 | | Female | - | - | | | | Charlson-Deyo score | 2+ | 1.44 (1.17-1.79) | <.001 | 0.001 | | 1 | 1.00 (0.83-1.20) | 0.996 | | | | 0 | - | - | | | | Histology | Other or unknown | 1.23 (0.93-1.61) | 0.149 | <.001 | | Squamous cell carcinomas | 1.94 (1.63-2.31) | <.001 | | | | Adenocarcinomas | - | - | | | | Age at diagnosis | | 1.06 (1.05-1.07) | <.001 | <.001 |
Table 6. Multivariable cox proportional hazard model for overall survival.
| | Months (OS) | | | | | | | ------------------------ | ---------------- | ---------------- | -------------------- | --------- | ------------ | | Covariate | Level | N | Hazard ratio(95% CI) | HRP-value | Type3P-value | | Surgical approach | Open | 21307 | 1.12 (1.06-1.18) | <.001 | <.001 | | Minimally invasive | 11843 | - | - | | | | Sex | Male | 14960 | 1.50 (1.43-1.58) | <.001 | <.001 | | Female | 18190 | - | - | | | | Urban/rural 2003 | Rural | 756 | 1.16 (1.00-1.34) | 0.049 | <.001 | | Urban | 5561 | 1.12 (1.06-1.19) | <.001 | | | | Metro | 26833 | - | - | | | | Charlson-Deyo score | 2+ | 4866 | 1.59 (1.49-1.70) | <.001 | <.001 | | 1 | 12153 | 1.26 (1.20-1.33) | <.001 | | | | 0 | 16131 | - | - | | | | Histology | Other or unknown | 4287 | 1.11 (1.03-1.20) | 0.004 | <.001 | | Squamous cell carcinomas | 8424 | 1.28 (1.21-1.35) | <.001 | | | | Adenocarcinomas | 20439 | - | - | | | | AJCC clinical T | 2 | 11077 | 1.49 (1.42-1.56) | <.001 | <.001 | | 1 | 22073 | - | - | | | | Age at diagnosis | | 33150 | 1.03 (1.03-1.04) | <.001 | <.001 |
In Table 5, the backward elimination drops clinical T stage, urban/rural and year of diagnosis from the 30-day mortality model at an alpha level of removal of 0.1, which means all variables in the final model have a p-value < 0.1. After controlling for the selected variables in the model, open surgery patients had a 41% higher odds to die within 30-days after surgery comparing to minimally invasive surgery patients (OR = 1.41; 95%CI = (1.18 - 1.69); p < 0.001). In Table 6, year of diagnosis was dropped from the final model to predict overall survival. Patients that underwent open surgery had 12% more chance to die before those that underwent minimally invasive surgery (HR = 1.12; 95% CI = (1.06-1.18); p < 0.001).
Stratified Multivariable model
When fitting the data into a main-effect model, as in Table 5 or Table 6, an imposed assumption is that the effect of treatment on outcomes is the same across all subgroups defined by the controlled variables. This assumption may or may not hold, and exploring and identifying subgroups that may benefit more from treatment can lead us to a deeper insight. Instead of splitting the dataset into smaller, separated data sets, fitting an interaction model on the entire data set is more appropriate18. As shown in Table 7, we fit a multivariable model including an interaction term between surgical approaches and histology, still implementing the backward elimination procedure. In the related %PHREG_SEL code, the first three terms in VAR were protected from elimination by setting INC = 3, to keep the interaction of interest (surg_app*his_cat) in the model. The macro parameters EFFECT and SLICEBY allow users to specify the variables for the treatment effect and subgroups (the two variables have to be categorical variables for the macro to run correctly). Setting SHORTREPORT = T, only reports the hazard ratio (HR) and the p-value of surgical approach by histology subgroup, and if set to F, the HR for all other control variables in the model will be reported. This macro can only handle one interaction at a time but is useful as an initial exploration for the treatment effect in subgroups.
Table 7. Multivariable Cox model for overall survival (OS), stratified by histology.
| | Months (OS) | | | | | | ---------------------------------- | -------------------------- | -------------------- | --------- | ------------ | | Covariate | Level | Hazard ratio(95% CI) | HRP-value | Type3P-value | | Comparisons stratifiedby histology | Surgical approach | - | - | 0.069 | | Adenocarcinomas | Minimally invasivevs. Open | 0.85 (0.79-0.91) | <.001 | - | | Other or unknown | Minimally invasivevs. open | 0.95 (0.82-1.10) | 0.511 | - | | Squamous cellcarcinomas | Minimally invasivevs. open | 0.96 (0.88-1.04) | 0.321 | - |
In Table 7, detailed information about variable selection in the model building process is shown in the footnote. We see that, overall, minimally invasive surgery shows a protective effect on survival compared to open surgery, but such protection is more pronounced among adenocarcinoma patients (HR = 0.85; p < 0.001) than squamous cell carcinomas (HR = 0.96; p = 0.321). The p-value for interaction term is 0.069, and it tests the difference among the HR of 0.85, 0.95, and 0.96 for the three histology groups. If using the significance level of 0.05, the interaction p-value may confirm that the hazard ratio for the surgical approaches is the same across the histology groups. In the case of a significant interaction effect, researchers should report the interaction model and discuss the treatment effect in each subgroup.
******** SAS® code example for %PHREG_SEL and %LOGREG_SEL **************;
TITLE 'Table 5 Multivariable Logistic Regression Model for 30-day Mortality';
%LOGREG_SEL(
- DSN = anal,
- OUTCOME = PUF_30_DAY_MORT_CD,
- EVENT = 'Yes',
- VAR = surg_app &cat_var &num_var,
- CVAR = surg_app*&cat_var_ref,
- INC = 1,
- SLSTAY = 0.1,
- CLNUM = F,
- OUTPATH = &dir,
- FILENAME = Table 5 Multivariable Logistic Regression Model for 30-day Mortality);
TITLE 'Table 6 Multivariable Cox Proportional Hazard Model for Overall Survival';
%PHREG_SEL (
- DSN=anal,
- EVENT = os_surg,
- CENSOR = os_censor,
- VAR = surg_app &cat_var &num_var,
- CVAR = surg_app * &cat_var_ref ,
- INC = 1,
- SLSTAY = .10,
- CLUNM=T,
- OUTPATH = &dir,
- FILENAME = Table 6 Multivariable Cox Proportional Hazard Model for Overall Survival);
TITLE 'Table 7 Multivariable Cox Model for OS Stratify by Histology';
%PHREG_SEL(
- DSN=anal,
- EVENT = os_surg,
- CENSOR = os_censor,
- VAR = surg_app his_cat surg_app*his_cat &cat_var &num_var,
- CVAR = surg_app &cat_var ,
- INC = 3,
- SLSTAY = .10,
- EFFECT = surg_app,
- SLICEBY = his_cat,
- SHORTREPORT = T,
- OUTPATH = &dir,
- FILENAME = Table 7 Multivariable Cox Model for OS Stratify by Histology);
TITLE;
*****************************************************************************;
Kaplan–Meier analysis
It is standard practice in many time-to-event studies to report Kaplan–Meier (KM) plots, median survival times, and accrual survival rates for time-to-event outcomes stratified by treatments, as a straightforward and intuitive way to assess the survival profile. %KM_PLOT was used to generate Figure 2. The KM plot for overall survival (specified in EVENTS, CENSORS) stratified by surgical approaches (specified in GRPLIST) was carried out with the key information of interest reported in a summary table. This macro has many options that allow the user to control the appearance of the plot (TITLE, XTICK, XMAX, NONCENSORED, and ATRISK) and output the estimated accrual survival rate at pre-specified time points (TIMELIST). This macro becomes handier when there are multiple datasets, several outcomes, or multiple variables of interest. Users can produce multiple KM plots in one macro call for all combinations of the parameter values by listing multiple values separated by spaces in DSN, EVENTS/CENSORS, and GRPLIST.
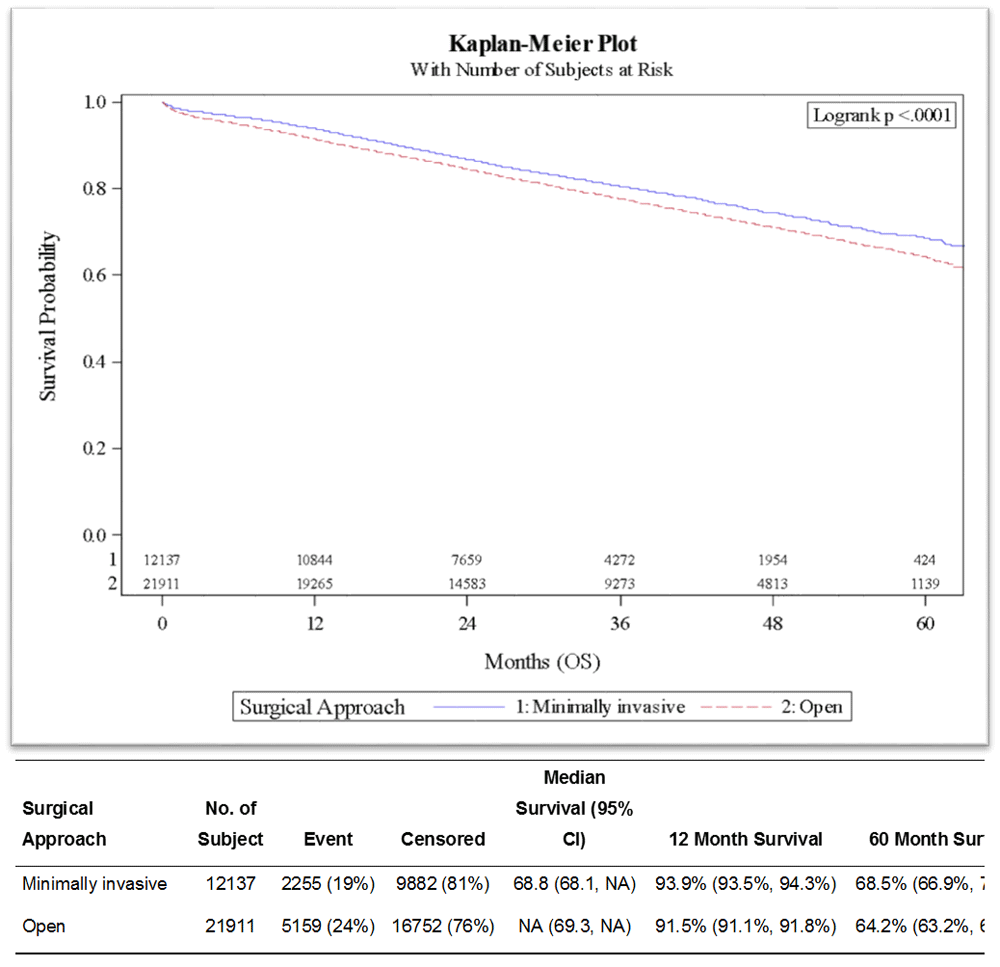
Figure 2. Kaplan–Meier plot of overall survival by surgical approach.
In Figure 2, the two survival curves by surgical approaches do not reach the survival probability of 0.5, and hence the median survival time is not estimable (will show as NAs). The 60-month (5-year) survival rate is 68.5% in the minimally invasive surgery group, while 64.2% in the open surgery group. Overall, the log-rank p < 0.001 indicates a significantly improved overall survival in the minimally invasive surgery group over the open surgery group (note that the KM method is only univariate analysis). However, a gain of 4.3% improvement in 5-year survival rate may or may not be clinically significant, and a large sample size may easily lead to small p-values. In such cases, researchers may shed light on both the clinical relevance and statistical significance to interpret results.
************ SAS® code example for %KM_PLOT *********** *******************;
%KM_PLOT(DSN = anal,
- EVENTS = os_surg,
- CENSORS = os_censor,
- GRPLIST = surg_app,
- TITLE = "Figure 2 KM plot of Overall Survival by Surgical Approach",
- UNIT = Month, JOIN = T, PLOT = T, TABLE = T,
- TIMELIST = 12 60, NONCENSORED = T,
- XTICK = (0 12 24 36 48 60), XMAX = 60, ATRISK = T,
- OUTPATH = &dir,
- FNAME = Figure 2 KM plot of Overall Survival by Surgical Approaches);
*****************************************************************************;