Contribution of intertwined loop to membrane association revealed by Zika virus full‐length NS1 structure (original) (raw)
Abstract
The association of Zika virus (ZIKV) infections with microcephaly and neurological diseases has highlighted an emerging public health concern. Here, we report the crystal structure of the full‐length ZIKV nonstructural protein 1 (NS1), a major host‐interaction molecule that functions in flaviviral replication, pathogenesis, and immune evasion. Of note, a long intertwined loop is observed in the wing domain of ZIKV NS1, and forms a hydrophobic “spike”, which can contribute to cellular membrane association. For different flaviviruses, the amino acid sequences of the “spike” are variable but their common characteristic is either hydrophobic or positively charged, which is a beneficial feature for membrane binding. Comparative studies with West Nile and Dengue virus NS1 structures reveal conserved features, but diversified electrostatic characteristics on both inner and outer faces. Our results suggest different mechanisms of flavivirus pathogenesis and should be considered during the development of diagnostic tools.
Synopsis
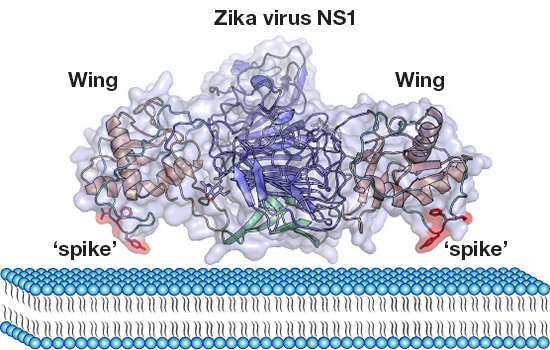
The Zika virus (ZIKV) nonstructural protein 1 (NS1) functions in viral replication, pathogenesis, and immune evasion. Here, we solved the crystal structure of full‐length NS1 protein, and found an extended membrane association interface contributed by the hydrophobic “spike” of a long intertwined loop, providing important information for ZIKV pathogenesis and development of diagnostic tools.
- Full‐length ZIKV NS1 structure has a similar overall structural fold to other flavivirus NS1 structures.
- A long intertwined loop forming a hydrophobic “spike” in the NS1 structure could contribute to cellular membrane association.
- The variable “spike” residues for different flaviviruses have common hydrophobic or positively charged characteristics.
- ZIKV NS1 has diversified electrostatic characteristics on both inner and outer faces.
- Conservation analysis of NS1 structure reveals vulnerable sites for the development of diagnostic tools.
Similar content being viewed by others
Introduction
Zika virus (ZIKV) has caused global concern, as growing clinical and experimental evidence showed that ZIKV infection is highly associated with microcephaly and serious neurological complications such as Guillain–Barré syndrome (GBS), even though ZIKV was previously considered a mild pathogen for humans (Cao‐Lormeau et al, 2016; Carteaux et al, 2016; Mlakar et al, 2016; Ventura et al, 2016; Wang et al, 2016; Zhang et al, 2016; Zhu et al, 2016). ZIKV belongs to Flaviviridae family and has a single‐strand positive‐sense RNA genome (Musso & Gubler, 2016). ZIKV can be transmitted by mosquitoes of the genus Aedes, and recently, several clinical investigations showed that it can also be transmitted by sexual contact (Ledermann et al, 2014; Musso et al, 2015; D'Ortenzio et al, 2016). To date, there are no clinically approved vaccines or antiviral drugs available to protect or treat ZIKV infections. To help accelerate the development of specific medical countermeasures, more studies are needed to understand the pathogenesis of ZIKV.
First described as a soluble complement fixing (SCF) antigen more than four decades ago (Brandt et al, 1970; Russell et al, 1970), the unusual properties of the flavivirus nonstructural protein 1 (NS1) have puzzled the flavivirus research field. NS1 protein is involved in many steps of the flavivirus life cycle, including viral replication, immune evasion, and pathogenesis (Amorim et al, 2014). It is thought that the glycosylated NS1 protein, which associates with lipids, forms a homodimer inside the cells, and separates into three distinct populations: a significant population participates in the viral replication complex, a second minor population is trafficked to the plasma membrane, and the third is secreted into the extracellular space as a hexameric lipoprotein particle (Flamand et al, 1999; Gutsche et al, 2011; Muller et al, 2012). Structural studies of dengue virus (DENV) and West Nile virus (WNV) NS1 have revealed the architecture of dimeric and hexameric NS1 proteins, providing molecular insight into the NS1 structure–function relationships (Akey et al, 2014). Recently, we have solved the C‐terminal region of ZIKV NS1 protein and elucidated a differential electrostatic potential in the host‐interaction interface, compared with other flavivirus proteins with known structures (Song et al, 2016a). However, molecular information about the full‐length ZIKV NS1 remains unknown.
Here, we have solved the crystal structure of full‐length ZIKV NS1, and found that the overall structural fold is similar to other flavivirus NS1 structures, including DENV and WNV. Importantly, a long intertwined loop was observed in the wing domain of ZIKV NS1, forming a hydrophobic “spike”. This hydrophobic “spike” can contribute to the membrane association of ZIKV NS1, together with the β‐roll and connector subdomain. Further sequence analysis revealed that the “spike” sequences of different flavivirus NS1 proteins are variable but have common features, consisting of hydrophobic or positively charged amino acids, which is beneficial for cellular membrane association. Moreover, ZIKV NS1 has unique electrostatic potential maps on both inner and outer faces, which might lead to interaction with host proteins different from those used by DENV and WNV NS1. Our results suggest that ZIKV pathogenesis may be distinct from other flaviviruses and should be considered in the development of specific diagnostic antigens.
Results
Overall structure of ZIKV full‐length NS1
We expressed the full‐length ZIKV NS1 (residues 2–352, BeH819015 strain isolated from Brazil in 2015) in insect cells using the baculovirus expression system, following the construct as previously reported (Brown et al, 2011). The recombinant NS1 protein was successfully purified and crystallized in 0.1 M Tris–HCl pH 8.5, 2.0 M ammonium sulfate, and then, we determined its X‐ray structure at 2.85 Å by molecular replacement using DENV NS1 structure (PDB ID: 4O6B) (Akey et al, 2014) as a search model (Table 1). The structure is complete for all amino acid residues, especially containing the internal loop (amino acids 108–128), which is not observed in either of the WNV or DENV2 structures. There are two potential N‐linked glycosylation sites (N130 and N207), and N207 is seen to be glycosylated with one sugar residue in clear electron density in our ZIKV NS1 structure. This observation of little carbohydrate does not reflect the real situation, and our insect cell‐produced NS1 protein would have simpler glycosylation than the mammalian cell‐produced NS1 protein. These two glycosylation sites are highly conserved for different flaviviruses.
Table 1 Data collection and refinement statistics
The overall structure of ZIKV NS1 protein is very similar to WNV and DENV2 NS1 structures (0.84 and 0.74 Å root mean square deviation (rmsd), respectively, of 539 Cα atoms). The ZIKV NS1 dimer is organized around an extended central β‐sheet domain, and each monomer has three domains, β‐hairpin domain, wing domain, and β‐ladder domain (Fig 1A). The first β‐hairpin domain (amino acids 1–30) forms a mini domain‐swap structure together with the β‐hairpin domain of the other NS1 monomer in the NS1 dimer structure, and these two β‐hairpin domains intertwine in a roll‐like structure, designated as “β‐roll” dimerization domain. The wing domain (amino acids 31–181) of each monomer protrudes from the central β‐ladder domain, containing one glycosylation site (N130) and two subdomains. An α/β subdomain (amino acids 38–151) consists of a four‐stranded β‐sheet, two α‐helices and a long intertwined loop between β5‐ and β6‐strands (amino acid residues 91–130) (Fig 1A and B). A discontinuous connector subdomain (amino acid residues 30–37 and 152–180) connects the wing domain to the central β‐roll and β‐ladder domains (Fig 1A and B). The predominant structural feature of NS1 is the third domain, a continuous β‐ladder domain (amino acid residues 182–352), extending along the length of the dimer with its 10 β‐strands (Fig 1A and B). The β‐ladder domains from two NS1 monomers from a long β‐sheet with 20 β‐strands arranged like the rungs of a ladder (Fig 1A). In the β‐ladder domain, most of the interstrand loops are short, except a long “spaghetti loop” between β13 and β14 that lacks repetitive secondary structures (such as α‐helix or β‐strand) but is ordered by extensive hydrogen bonds (Fig 1C, cyan). The β‐ladder domain of each monomer can be separated into two parts by the “spaghetti loop”, the central region containing the first four β‐strand rungs and the tip region containing the remaining six β‐strand rungs (Fig 1B).
Figure 1
Overall structure of ZIKV NS1 dimer and extended surface for membrane association
A. NS1 dimer with one subunit in gray and the other colored by domain (green, β‐roll; orange, wing with magenta connector subdomain; blue, central β‐ladder). N‐linked glycosylation sites are shown in sticks, and the glycans are shown in spheres.
B. Topology diagram for the NS1 monomer, colored as the pattern in (A). Glycosylation sites are indicated with red hexagons and disulfide bonds with yellow circles.
C. Side views of the NS1 dimer from the wing (left) and the end of the β‐ladder (right). The β‐roll (green), connector subdomain (magenta), and the intertwined loop (yellow) of the wing domain (orange) form a discontinuous protrusion on one face of the β‐ladder with the spaghetti loop (cyan) on the other face. The wing domain is omitted from the left image for clarity.
The overall dimensions of the ZIKV NS1 dimer are 85 Å along the β‐ladder, 95 Å in width from wingtip to wingtip, and 35 Å in the third dimension (Fig 1A). The β‐ladder defines a plane with the β‐roll domain and the second half part of intertwined loop of the wing domain on one side, and on the other side of the plane are the “spaghetti loop”, the N130‐glycosylation site, the first half part of intertwined loop of the wing domain and the C‐terminus, which is fused to the lumen‐side N terminus of viral protein NS2A before proteolytic cleavage (Fig 1C).
Intertwined loop may contribute to membrane association of ZIKV NS1
The β‐roll, the connector subdomain, and the second half part of the intertwined loop of the wing create a discontinuous linear hydrophobic protrusion with a markedly hydrophobic surface on the inner side of the dimer (Fig 1C). This hydrophobic surface is mainly constituted by aromatic residues such as F8, Y122, F123, F163, and H164 (Fig 1C). This “hydrophobic protrusion” provides a perfect interface to associate with the hydrophobic‐lipid membrane surface, and orients the outer hydrophilic face of the dimer toward the lumen, and subsequently the extracellular space. Previous work on WNV NS1 showed that the residues at the β‐roll (10–11 on a β‐turn) are implicated in a direct interaction with NS4B (Youn et al, 2012), implying the membrane surface interacting with other proteins of the replication complex. Residues in the connector subdomain (159–162 on a β‐turn) are also shown to be important for viral replication (Akey et al, 2014). In our ZIKV NS1 structure, we can clearly see the second half part of the intertwined loop of the wing, which exists between the β5 and β6 strands, clearly. This long loop (consists of Y122, F123, V124) forms a hydrophobic “spike” that may associate with the membrane. This exposed loop is considered to be critical in mediating NS1 interaction with the envelope protein, thus tethering the replication complex with virion formation (Scaturro et al, 2015).
In addition, we have modeled the ZIKV hexamer structure based on that previously reported for DENV2 NS1 (Fig 2). We found that the intertwined loop “spike” of one monomer has a very close distance to the β‐ladder domain of the neighboring NS1 monomer (Fig 2), indicating that the intertwined loop probably participates in the formation of the ZIKV NS1 hexamer. The modeled ZIKV NS1 hexamer reveals a hydrophobic hole surrounded by the “inner” faces of three NS1 dimers, containing the β‐roll domain and the intertwined loop “spike” (Fig 2). By contrast, the “outer” faces can be easily bound by host factors and reactive antibodies.
Figure 2
Model of the ZIKV NS1 hexamer
The ZIKV NS1 hexamer is modeled by using the full‐length DENV2 NS1 structure (PDB code: 4O6B) (Akey et al, 2014). Each monomer is colored by domains (green, β‐roll; orange, wing; magenta, connector subdomain; blue, central β‐ladder). The intertwined loop “spike” of ZIKV NS1 contributing to membrane association is colored in red and indicated by dotted lines.
Comparison with other flavivirus NS1 structures
We compared the ZIKV NS1 structure with the available NS1 structures of DENV2 and WNV (Akey et al, 2014). Previously, we found that the electrostatic surface potential maps of these NS1 proteins have group‐specific features in the loop surface of the β‐ladder domain (Song et al, 2016a). The DENV2 NS1 structure displays a positively charged surface in the central region of the loop surface, whereas the WNV NS1 structure has a negatively charged central region. For ZIKV, the loop surface exhibits a composite surface containing both a positively and negatively charged central region and displays a negatively charged region toward the two distal ends (Fig 3). Interestingly, in the “inner” face with the β‐roll domain, only ZIKV displays negatively charged surface, whereas DENV2 and WNV display surfaces of neutral charge. Moreover, we have observed different features in the “outer” face of the wing domain. ZIKV NS1 displays a positively charged surface in the tip region of the wing domain, whereas the WNV NS1 structure has a negatively charged region, and the DENV2 NS1 structure contains both positively and negatively charged regions (Fig 3).
Figure 3
Variation in electrostatic surface potential of flavivirus NS1 proteins
A–C Electrostatic surface views of NS1 from DENV2 (PDB: 4O6B, A) and WNV (PDB: 4O6C, B) (Akey et al, 2014), as well as ZIKV (C), showing diverse characteristics on certain regions. Surfaces are colored by electrostatic potential at neutral pH from −2 kT (red) to +2 kT (blue) using PyMOL (http://pymol.org/).
Conserved features of NS1 sequences relating to their structure–function relationships
In Fig 4A, the NS1 surface is colored according to sequence conservation from the most conserved (dark magenta) to the most divergent (dark cyan) based on an alignment of NS1 sequences from 73 flaviviruses (Fig 4A and Table EV1). The most conserved surfaces are on the β‐roll region and the C‐terminal tip of the central β‐ladder. The “outer” face of the wing domain is the most variable region. For the protruding tip (residues 122–124) of the intertwined loop, the amino acid sequence is variable (Fig 4B). Irrespective of amino acid variations, these residues are always hydrophobic or positively charged, which is beneficial for membrane association (Figs 5 and EV1).
Figure 4
NS1 sequence conservation mapped onto the protein surface and models of membrane association for ZIKV, WNV and DENV2
A. The NS1 surface is colored using the ConSurf server (Ashkenazy et al, 2016) according to sequence conservation from the most conserved (dark magenta) to the most divergent (dark cyan) based on an alignment of NS1 sequences from 73 flaviviruses.
B. Models of intertwined loop “spike” interacting with the cellular membrane. The disordered “spike” regions are indicated by dash lines in DENV2 and WNV NS1 structures, which consist of hydrophobic or positively charged amino acids.
Figure 5
Detailed analysis of the intertwined loop in the wing domain of ZIKV NS1 structure
A. B‐factor putty representation of the intertwined loop structure. Shades of blue and a narrower tube indicate regions with lower B‐factors, whereas orange to red and a wider tube indicate regions with higher B‐factors. The figure was prepared using PyMOL (http://pymol.org/).
B. The 2Fo‐Fc electron density map for the intertwined loop (yellow) “spike” region residues contoured at 1.0 sigma is represented in blue.
C. Detailed interactions of the highly conserved residues W115 and W118 with the neighboring residues to stabilize the conformation of the intertwined loop. W115 forms a hydrogen‐bonding interaction with E74 from α2‐helix. W118 inserts into the hydrophobic interspace between α1‐ and α2‐helix, formed by P36, L42, V71, L75, and I78, and W118 also interact with P36 and D37 by hydrogen‐bonding interactions.
D. Sequence alignment of the intertwined loop region (residues 91–130) of different flavivirus NS1 proteins. The intertwined loop “spike” residues of ZIKV NS1 contributing to membrane association are indicated by a dark blue arrowhead. Accession codes: ZIKV strain BeH819015, AMA12085; West Nile virus (WNV), Q9Q6P4; Dengue virus 1 Nauru/West Pac/1974 (DENV1), P17763; Dengue virus 2 Puerto Rico/PR159‐S1/1969 (DENV2), P12823; Dengue virus 3 Singapore/8120/1995 (DENV3), Q5UB51; Dengue virus 4 Thailand/0348/1991 (DENV4), Q2YHF0; Japanese encephalitis strain K94P05 (JEV), AAC02714; St. Louis encephalitis (SLEV), D8L537; Yellow fever 17D vaccine strain (YFV), CAA27332; Murray Valley encephalitis (MVEV), NP_051124; Tickborne encephalitis (TBEV), NP_043135.
The “outer” face has been suggested to play a crucial role in interactions of NS1 with host factors and antibodies (Chung et al, 2006a,b; Avirutnan et al, 2007; Edeling et al, 2014). A wide range of host proteins present on platelets and endothelial cells have been identified as cross‐reactive targets by NS1‐induced autoantibodies (Sun et al, 2007, 2015; Cheng et al, 2009; Chuang et al, 2011, 2014; Liu et al, 2011; Yin et al, 2013). ZIKV NS1 displays an interface with variable surface and divergent electrostatic potential that may result in altered binding properties to host factors and protective antibodies against flavivirus NS1.
Discussion
As the only viral protein which can be secreted into the extracellular space, the nonstructural protein NS1 plays diverse roles in immune evasion via binding to complement proteins and modifying or antagonizing their functions (Chung et al, 2006a; Avirutnan et al, 2010, 2011). NS1 was also shown to activate Toll‐like receptor 4 signaling in primary human myeloid cells, leading to secretion of pro‐inflammatory cytokines and vascular leakage (Beatty et al, 2015; Modhiran et al, 2015). Besides its immune evasive and endothelial permeability functions, NS1 modulates early events in viral RNA replication (Watterson et al, 2016). Indeed, deletion of NS1 completely abrogates viral RNA replication, but ectopic expression of NS1 in trans can efficiently rescue NS1‐deleted viruses (Lindenbach & Rice, 1997; Khromykh et al, 2000). It is thought that membrane‐associated NS1 dimer on the luminal face of the endoplasmic reticulum (ER) membrane plays an essential organizational role in the formation of the replication complex through interaction with NS4A and NS4B (Lindenbach & Rice, 1999; Youn et al, 2012). NS1 has also been proposed to interact with prM and the envelope protein, linking replicative complexes within the vesicle packets together with virion formation (Scaturro et al, 2015). In our ZIKV NS1 structure, we found that the intertwined loop “spike”, between the β5 and β6 strands of the wing domain, containing hydrophobic residues Y122, F123, and V124, forms an extended hydrophobic protrusion together with the β‐roll and the connector subdomain, and most likely involves in association with the membrane surface. It is intriguing that this intertwined loop is disordered in the previously reported DENV2 and WNV NS1 crystal structures. We speculate two possibilities for the discrepancy: First, we have further analyzed the crystal contacts of the intertwined loop in our ZIKV NS1 structure, and indeed found that the crystal packing can help stabilize the conformation of this intertwined loop; second, the different protein purification methods (the DENV2 and WNV NS1 proteins were purified with detergent) may also affect the conformation of the hydrophobic intertwined loop, as the detergent molecules could bind to the intertwined loop. Detailed analysis revealed that the intertwined loop does not have higher B‐factors (temperature factors) than the rest of the ZIKV NS1 structure, and is observed in the clear electron density map (Fig 5A and B), indicating that this loop is relatively stable in our structure. Moreover, highly conserved residues W115 and W118 could further stabilize the conformation of the intertwined loop by interaction with the neighboring residues (Fig 5C and D). Janet Smith and her colleagues recently reported the full‐length NS1 structure from the original 1947 Ugandan ZIKV strain (MR766). The structure is similar to that reported here, including an elongated hydrophobic region for membrane association in the wing domain, highlighting the importance of the intertwined loop (Brown et al, [2016](/article/10.15252/embj.201695290#ref-CR500 "Brown WC, Akey DL, Konwerski JR, Tarrasch JT, Skiniotis G, Kuhn RJ, Smith JL (2016) Extended surface for membrane association in Zika virus NS1 structure. Nat Struct Mol Biol 23: 865–867 ‡Correction added on 17 October 2016, after first online publication: the reference Brown et al (2016) has been added to the reference list.
")).Footnote 1
From the conservation analysis of all the flavivirus NS1 sequences mapped onto the protein surface, we found that the hydrophobic protrusion and the C‐terminal tip of the central β‐ladder regions are most conserved, while the “outer” face of the wing domain are most variable, indicating the latter has divergent characteristics among different flaviviruses. The electrostatic surface potential of that region in ZIKV NS1 also shows specific features compared with that of DENV2 and WNV NS1 structures. The unique surface characteristics of ZIKV NS1 may be related to ZIKV neurotropism and could be exploited in the development of novel therapeutic and diagnostic tools for ZIKV infection (Stettler et al, 2016).
The high level of secreted NS1 in the blood of flavivirus‐infected individuals during early infection has made NS1 a primary biomarker in disease diagnosis (Alcon et al, 2002). In addition, NS1 is a potential vaccination candidate against flavivirus infection (Amorim et al, 2014). Immunization of mice with DENV NS1 was protective against subsequent lethal DENV challenge (Schlesinger et al, 1987). However, autoantibodies against NS1 can elicit severe autoimmune diseases (Cheng et al, 2009; Chuang et al, 2011). More recently, a panel of monoclonal antibodies from ZIKV‐infected patients was isolated, and most of the antibodies were ZIKV‐specific, and memory T cells against NS1 proteins were poorly cross‐reactive, even in donors pre‐exposed to DENV, indicating that the ZIKV NS1 protein could induce a prominent ZIKV‐specific immune response (Stettler et al, 2016). Their results support our data that the ZIKV NS1 structure has unique surface characteristics, which has implications for developing a serological diagnostic tool against ZIKV NS1. Further research should focus on characterizing the diverse epitopes within NS1 for eliciting auto‐ or protective antibodies.
Materials and Methods
Gene cloning, expression, and protein purification
The ZIKV NS1 construct (amino acid residues 2–352, Accession Number: AMA12085) was cloned into the baculovirus transfer vector pFastBac Dual (Invitrogen) under control of the polyhedrin promoter, in‐frame with an N‐terminal gp67 signal peptide for secretion and a His6‐tag at the N‐terminus for purification. We placed green fluorescent protein (GFP) under control of the P10 promoter to visualize expression, which correlated well with expression of NS1. Recombinant pFastBac Dual plasmid was used to transform DH10Bac™Escherichia coli (Invitrogen). Transfection and virus amplification were performed according to the Bac‐to‐Bac baculovirus expression system manual (Invitrogen) (Song et al, 2016b). NS1 proteins were produced by infecting suspension cultures of Hi5 cells (Invitrogen) for 2 days. Soluble NS1 was recovered from cell supernatants by metal affinity chromatography using a HisTrap HP 5 ml column (GE Healthcare), and then purified by ion‐exchange chromatography using a RESOURCE™Q 6 ml column (GE Healthcare). For crystallization, the proteins were further purified by gel filtration chromatography using a Hiload 16/60 Superdex® 200 PG column (GE Healthcare) with a running buffer of 20 mM Tris–HCl and 50 mM NaCl (pH 8.5), and the collected protein fractions were concentrated to 10 mg/ml using a membrane concentrator with a molecular weight cutoff of 10 kDa (Millipore).
Crystallization, data collection, and structure determination
The initial screening trials were set up with commercial crystallization kits (Hampton Research and Molecular Dimensions) using the sitting‐drop vapor diffusion method with a Mosquito® LCP crystal screening robot (TTP Labtech). Normally, 0.5 μl protein was mixed with 0.5 μl reservoir solution. The resultant drop was then sealed, equilibrating against 70 μl reservoir solution at 4 or 18°C. Diffractable crystals were obtained in 0.1 M Tris–HCl pH 8.5, 2.0 M ammonium sulfate. Crystals were flash‐cooled in liquid nitrogen after a brief soak in reservoir solution with the addition of 17% (v/v) glycerol. X‐ray diffraction data were collected under cryogenic conditions (100 K) at the wavelength 0.97853 Å at Shanghai Synchrotron Radiation Facility (SSRF) beamline BL17U, and indexed, integrated, and scaled with HKL2000 (Otwinowski & Minor, 1997).
The ZIKV NS1 structure was solved by the molecular replacement method using Phaser (Read, 2001) from the CCP4 program suite (Winn et al, 2011), with the structure of DENV2 NS1 (PDB: 4O6B) as the search model (Akey et al, 2014). Initial restrained rigid‐body refinement and manual model building were performed using REFMAC5 (Murshudov et al, 1997) and COOT (Emsley & Cowtan, 2004), respectively. Further refinement was performed using Phenix (Adams et al, 2010). Final statistics for data collection and structure refinement are represented in Table 1.
Notes
- †Correction added on 17 October 2016, after first online publication: the last two sentences in this paragraph have been added.
- ‡Correction added on 17 October 2016, after first online publication: the reference Brown et al ([2016](/article/10.15252/embj.201695290#ref-CR500 "Brown WC, Akey DL, Konwerski JR, Tarrasch JT, Skiniotis G, Kuhn RJ, Smith JL (2016) Extended surface for membrane association in Zika virus NS1 structure. Nat Struct Mol Biol 23: 865–867
‡Correction added on 17 October 2016, after first online publication: the reference Brown et al (2016) has been added to the reference list.
")) has been added to the reference list.
References
- Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse‐Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) PHENIX: a comprehensive Python‐based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66: 213–221
Google Scholar - Akey DL, Brown WC, Dutta S, Konwerski J, Jose J, Jurkiw TJ, DelProposto J, Ogata CM, Skiniotis G, Kuhn RJ, Smith JL (2014) Flavivirus NS1 structures reveal surfaces for associations with membranes and the immune system. Science 343: 881–885
Google Scholar - Alcon S, Talarmin A, Debruyne M, Falconar A, Deubel V, Flamand M (2002) Enzyme‐linked immunosorbent assay specific to dengue virus type 1 nonstructural protein NS1 reveals circulation of the antigen in the blood during the acute phase of disease in patients experiencing primary or secondary infections. J Clin Microbiol 40: 376–381
Google Scholar - Amorim JH, Alves RP, Boscardin SB, Ferreira LC (2014) The dengue virus non‐structural 1 protein: risks and benefits. Virus Res 181: 53–60
Google Scholar - Ashkenazy H, Abadi S, Martz E, Chay O, Mayrose I, Pupko T, Ben‐Tal N (2016) ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res 44: W344–W350
Google Scholar - Avirutnan P, Zhang L, Punyadee N, Manuyakorn A, Puttikhunt C, Kasinrerk W, Malasit P, Atkinson JP, Diamond MS (2007) Secreted NS1 of dengue virus attaches to the surface of cells via interactions with heparan sulfate and chondroitin sulfate E. PLoS Pathog 3: e183
Google Scholar - Avirutnan P, Fuchs A, Hauhart RE, Somnuke P, Youn S, Diamond MS, Atkinson JP (2010) Antagonism of the complement component C4 by flavivirus nonstructural protein NS1. J Exp Med 207: 793–806
Google Scholar - Avirutnan P, Hauhart RE, Somnuke P, Blom AM, Diamond MS, Atkinson JP (2011) Binding of flavivirus nonstructural protein NS1 to C4b binding protein modulates complement activation. J Immunol 187: 424–433
Google Scholar - Beatty PR, Puerta‐Guardo H, Killingbeck SS, Glasner DR, Hopkins K, Harris E (2015) Dengue virus NS1 triggers endothelial permeability and vascular leak that is prevented by NS1 vaccination. Sci Transl Med 7: 304ra141
Google Scholar - Brandt WE, Chiewslip D, Harris DL, Russell PK (1970) Partial purification and characterization of a dengue virus soluble complement‐fixing antigen. J Immunol 105: 1565–1568
Google Scholar - Brown WC, DelProposto J, Rubin JR, Lamiman K, Carless J, Smith JL (2011) New ligation‐independent cloning vectors compatible with a high‐throughput platform for parallel construct expression evaluation using baculovirus‐infected insect cells. Protein Expr Purif 77: 34–45
Google Scholar - Brown WC, Akey DL, Konwerski JR, Tarrasch JT, Skiniotis G, Kuhn RJ, Smith JL (2016) Extended surface for membrane association in Zika virus NS1 structure. Nat Struct Mol Biol 23: 865–867Footnote
‡Correction added on 17 October 2016, after first online publication: the reference Brown et al ([2016](/article/10.15252/embj.201695290#ref-CR500 "Brown WC, Akey DL, Konwerski JR, Tarrasch JT, Skiniotis G, Kuhn RJ, Smith JL (2016) Extended surface for membrane association in Zika virus NS1 structure. Nat Struct Mol Biol 23: 865–867
‡Correction added on 17 October 2016, after first online publication: the reference Brown et al (2016) has been added to the reference list.
")) has been added to the reference list.
Google Scholar
- Cao‐Lormeau VM, Blake A, Mons S, Lastere S, Roche C, Vanhomwegen J, Dub T, Baudouin L, Teissier A, Larre P, Vial AL, Decam C, Choumet V, Halstead SK, Willison HJ, Musset L, Manuguerra JC, Despres P, Fournier E, Mallet HP et al (2016) Guillain‐Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: a case‐control study. Lancet 387: 1531–1539
Google Scholar - Carteaux G, Maquart M, Bedet A, Contou D, Brugières P, Fourati S, Cleret de Langavant L, de Broucker T, Brun‐Buisson C, Leparc‐Goffart I, Mekontso Dessap A (2016) Zika virus associated with meningoencephalitis. New Engl J Med 374: 1595–1596
Google Scholar - Cheng HJ, Lin CF, Lei HY, Liu HS, Yeh TM, Luo YH, Lin YS (2009) Proteomic analysis of endothelial cell autoantigens recognized by anti‐dengue virus nonstructural protein 1 antibodies. Exp Biol Med 234: 63–73
Google Scholar - Chung KM, Liszewski MK, Nybakken G, Davis AE, Townsend RR, Fremont DH, Atkinson JP, Diamond MS (2006a) West Nile virus nonstructural protein NS1 inhibits complement activation by binding the regulatory protein factor H. Proc Natl Acad Sci USA 103: 19111–19116
Google Scholar - Chung KM, Nybakken GE, Thompson BS, Engle MJ, Marri A, Fremont DH, Diamond MS (2006b) Antibodies against West Nile virus nonstructural protein NS1 prevent lethal infection through Fc gamma receptor‐dependent and ‐independent mechanisms. J Virol 80: 1340–1351
Google Scholar - Chuang YC, Lei HY, Lin YS, Liu HS, Wu HL, Yeh TM (2011) Dengue virus‐induced autoantibodies bind to plasminogen and enhance its activation. J Immunol 187: 6483–6490
Google Scholar - Chuang YC, Lin YS, Liu HS, Yeh TM (2014) Molecular mimicry between dengue virus and coagulation factors induces antibodies to inhibit thrombin activity and enhance fibrinolysis. J Virol 88: 13759–13768
Google Scholar - D'Ortenzio E, Matheron S, Yazdanpanah Y, de Lamballerie X, Hubert B, Piorkowski G, Maquart M, Descamps D, Damond F, Leparc‐Goffart I (2016) Evidence of sexual transmission of Zika virus. N Engl J Med 374: 2195–2198
Google Scholar - Edeling MA, Diamond MS, Fremont DH (2014) Structural basis of Flavivirus NS1 assembly and antibody recognition. Proc Natl Acad Sci USA 111: 4285–4290
Google Scholar - Emsley P, Cowtan K (2004) Coot: model‐building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60: 2126–2132
Google Scholar - Flamand M, Megret F, Mathieu M, Lepault J, Rey FA, Deubel V (1999) Dengue virus type 1 nonstructural glycoprotein NS1 is secreted from mammalian cells as a soluble hexamer in a glycosylation‐dependent fashion. J Virol 73: 6104–6110
Google Scholar - Gutsche I, Coulibaly F, Voss JE, Salmon J, d'Alayer J, Ermonval M, Larquet E, Charneau P, Krey T, Megret F, Guittet E, Rey FA, Flamand M (2011) Secreted dengue virus nonstructural protein NS1 is an atypical barrel‐shaped high‐density lipoprotein. Proc Natl Acad Sci USA 108: 8003–8008
Google Scholar - Khromykh AA, Sedlak PL, Westaway EG (2000) cis‐ and trans‐acting elements in flavivirus RNA replication. J Virol 74: 3253–3263
Google Scholar - Ledermann JP, Guillaumot L, Yug L, Saweyog SC, Tided M, Machieng P, Pretrick M, Marfel M, Griggs A, Bel M, Duffy MR, Hancock WT, Ho‐Chen T, Powers AM (2014) Aedes hensilli as a potential vector of Chikungunya and Zika viruses. PLoS Negl Trop Dis 8: e3188
Google Scholar - Lindenbach BD, Rice CM (1997) Trans‐complementation of yellow fever virus NS1 reveals a role in early RNA replication. J Virol 71: 9608–9617
Google Scholar - Lindenbach BD, Rice CM (1999) Genetic interaction of flavivirus nonstructural proteins NS1 and NS4A as a determinant of replicase function. J Virol 73: 4611–4621
Google Scholar - Liu IJ, Chiu CY, Chen YC, Wu HC (2011) Molecular mimicry of human endothelial cell antigen by autoantibodies to nonstructural protein 1 of dengue virus. J Biol Chem 286: 9726–9736
Google Scholar - Mlakar J, Korva M, Tul N, Popović M, Poljšak‐Prijatelj M, Mraz J, Kolenc M, Resman Rus K, Vesnaver Vipotnik T, Fabjan Vodušek V, Vizjak A, Pižem J, Petrovec M, Avšič Županc T (2016) Zika virus associated with microcephaly. New Engl J Med 374: 951–958
Google Scholar - Modhiran N, Watterson D, Muller DA, Panetta AK, Sester DP, Liu L, Hume DA, Stacey KJ, Young PR (2015) Dengue virus NS1 protein activates cells via Toll‐like receptor 4 and disrupts endothelial cell monolayer integrity. Sci Transl Med 7: 304ra142
Google Scholar - Muller DA, Landsberg MJ, Bletchly C, Rothnagel R, Waddington L, Hankamer B, Young PR (2012) Structure of the dengue virus glycoprotein non‐structural protein 1 by electron microscopy and single‐particle analysis. J Gen Virol 93: 771–779
Google Scholar - Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum‐likelihood method. Acta Crystallogr D Biol Crystallogr 53: 240–255
Google Scholar - Musso D, Roche C, Robin E, Nhan T, Teissier A, Cao‐Lormeau VM (2015) Potential sexual transmission of Zika virus. Emerg Infect Dis 21: 359–361
Google Scholar - Musso D, Gubler DJ (2016) Zika virus. Clin Microbiol Rev 29: 487–524
Google Scholar - Otwinowski Z, Minor W (1997) Processing of X‐ray diffraction data collected in oscillation mode. Methods Enzymol 276: 307–326
Google Scholar - Read RJ (2001) Pushing the boundaries of molecular replacement with maximum likelihood. Acta Crystallogr D 57: 1373–1382
Google Scholar - Russell PK, Chiewsilp D, Brandt WE (1970) Immunoprecipitation analysis of soluble complement‐fixing antigens of dengue viruses. J Immunol 105: 838–845
Google Scholar - Scaturro P, Cortese M, Chatel‐Chaix L, Fischl W, Bartenschlager R (2015) Dengue virus non‐structural protein 1 modulates infectious particle production via interaction with the structural proteins. PLoS Pathog 11: e1005277
Google Scholar - Schlesinger JJ, Brandriss MW, Walsh EE (1987) Protection of mice against dengue 2 virus encephalitis by immunization with the dengue 2 virus non‐structural glycoprotein NS1. J Gen Virol 68(Pt 3): 853–857
Google Scholar - Song H, Qi J, Haywood J, Shi Y, Gao GF (2016a) Zika virus NS1 structure reveals diversity of electrostatic surfaces among flaviviruses. Nat Struct Mol Biol 23: 456–458
Google Scholar - Song H, Qi J, Khedri Z, Diaz S, Yu H, Chen X, Varki A, Shi Y, Gao GF (2016b) An open receptor‐binding cavity of hemagglutinin‐esterase‐fusion glycoprotein from newly‐identified influenza D virus: basis for its broad cell tropism. PLoS Pathog 12: e1005411
Google Scholar - Stettler K, Beltramello M, Espinosa DA, Graham V, Cassotta A, Bianchi S, Vanzetta F, Minola A, Jaconi S, Mele F, Foglierini M, Pedotti M, Simonelli L, Dowall S, Atkinson B, Percivalle E, Simmons CP, Varani L, Blum J, Baldanti F et al (2016) Specificity, cross‐reactivity and function of antibodies elicited by Zika virus infection. Science 353: 823–826
Google Scholar - Sun DS, King CC, Huang HS, Shih YL, Lee CC, Tsai WJ, Yu CC, Chang HH (2007) Antiplatelet autoantibodies elicited by dengue virus non‐structural protein 1 cause thrombocytopenia and mortality in mice. J Thromb Haemost 5: 2291–2299
Google Scholar - Sun DS, Chang YC, Lien TS, King CC, Shih YL, Huang HS, Wang TY, Li CR, Lee CC, Hsu PN, Chang HH (2015) Endothelial cell sensitization by death receptor fractions of an anti‐dengue nonstructural protein 1 antibody induced plasma leakage, coagulopathy, and mortality in mice. J Immunol 195: 2743–2753
Google Scholar - Ventura CV, Maia M, Bravo‐Filho V, Gois AL, Belfort R Jr (2016) Zika virus in Brazil and macular atrophy in a child with microcephaly. Lancet 387: 228
Google Scholar - Wang Q, Yang Y, Zheng H, Bi Y, Song J, Li L, Gu D, Wang P, Li S, Liu S, Zhao Y, Liu L, Gao GF, Liu Y (2016) Genetic and biological characterization of Zika virus from human cases imported through Shenzhen Port (in Chinese). Chin Sci Bull 61: 2463–2474
Google Scholar - Watterson D, Modhiran N, Young PR (2016) The many faces of the flavivirus NS1 protein offer a multitude of options for inhibitor design. Antiviral Res 130: 7–18
Google Scholar - Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AGW, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Overview of the CCP4 suite and current developments. Acta Crystallogr D 67: 235–242
Google Scholar - Yin Y, Jiang L, Fang D, Jiang L, Zhou J (2013) Differentially expressed genes of human microvascular endothelial cells in response to anti‐dengue virus NS1 antibodies by suppression subtractive hybridization. Viral Immunol 26: 185–191
Google Scholar - Youn S, Li T, McCune BT, Edeling MA, Fremont DH, Cristea IM, Diamond MS (2012) Evidence for a genetic and physical interaction between nonstructural proteins NS1 and NS4B that modulates replication of West Nile virus. J Virol 86: 7360–7371
Google Scholar - Zhang Y, Chen W, Wong G, Bi Y, Yan J, Sun Y, Chen E, Yan H, Lou X, Mao H, Xia S, Gao GF, Shi W, Chen Z (2016) Highly diversified Zika viruses imported to China, 2016. Protein Cell 7: 461–464
Google Scholar - Zhu Z, Chan JF, Tee KM, Choi GK, Lau SK, Woo PC, Tse H, Yuen KY (2016) Comparative genomic analysis of pre‐epidemic and epidemic Zika virus strains for virological factors potentially associated with the rapidly expanding epidemic. Emerg Microbes Infect 5: e22
Google Scholar
Acknowledgements
This work was supported by the Emergency Task‐Force Project (Grant No. 81641001) of the National Natural Science Foundation of China (NSFC), the China Ministry of Science and Technology (MOST) National 973 Project (Grant No. 2014CB542503), the Task‐Force of Zika Virus Research from the Chinese Academy of Sciences (CAS), the National Key Plan for Scientific Research and Development of China (2016YFD0500300), Zika Special Project of the MOST Reform and Development Project, and Strategic Priority Research Program of the Chinese Academy of Sciences (XDB08020100). We thank the staff of beamline BL17U at Shanghai Synchrotron Radiation Facility, for assistance during data collection. Y.S. is supported by the Excellent Young Scientist Program from the NSFC (No. 81622031), the Excellent Young Scientist Program of the Chinese Academy of Sciences and the Youth Innovation Promotion Association CAS (2015078). G.F.G. is supported partly as a leading principal investigator of the NSFC Innovative Research Group (Grant No. 81321063).
Author information
Author notes
- These authors contributed equally to this work
Authors and Affiliations
- School of Life Sciences, University of Science and Technology of China, Hefei, China
Xiaoying Xu & George F Gao - CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing, China
Xiaoying Xu, Hao Song, Jianxun Qi, Yuqian Liu, Haiyuan Wang, Chao Su, Yi Shi & George F Gao - Research Network of Immunity and Health (RNIH), Beijing Institutes of Life Science, Chinese Academy of Sciences, Beijing, China
Hao Song & George F Gao - Savaid Medical School, University of Chinese Academy of Sciences, Beijing, China
Jianxun Qi, Yi Shi & George F Gao - Institute of Health Sciences, Anhui University, Hefei, China
Yuqian Liu - College of Animal Sciences and Technology, Guangxi University, Nanning, China
Haiyuan Wang - College of Veterinary Medicine, China Agricultural University, Beijing, China
Chao Su - Center for Influenza Research and Early‐warning (CASCIRE), Chinese Academy of Sciences, Beijing, China
Yi Shi & George F Gao - National Institute for Viral Disease Control and Prevention, Chinese Center for Disease Control and Prevention (China CDC), Beijing, China
George F Gao
Authors
- Xiaoying Xu
- Hao Song
- Jianxun Qi
- Yuqian Liu
- Haiyuan Wang
- Chao Su
- Yi Shi
- George F Gao
Contributions
GFG and YS designed and supervised the study. XX, HS, YL, HW, and CS conducted the experiments. JQ collected the data sets and solved the structures. YS, HS, and GFG analyzed the data and wrote the manuscript.
Corresponding authors
Correspondence toYi Shi or George F Gao.
Ethics declarations
The authors declare that they have no conflict of interest.
Additional information
See also: R Hilgenfeld (December 2016)
The EMBO Journal (2016) 35: 2170–2178
Supplementary Information
Rights and permissions
This is an open access article under the terms of the Creative Commons Attribution‐NonCommercial‐NoDerivs 4.0 License, which permits use and distribution in any medium, provided the original work is properly cited, the use is non‐commercial and no modifications or adaptations are made.
Copyright: The Author(s)
About this article
Cite this article
Xu, X., Song, H., Qi, J. et al. Contribution of intertwined loop to membrane association revealed by Zika virus full‐length NS1 structure.EMBO J 35, 2170–2178 (2016). https://doi.org/10.15252/embj.201695290
- Received: 19 July 2016
- Revised: 14 August 2016
- Accepted: 15 August 2016
- Published: 30 August 2016
- Issue date: 17 October 2016
- DOI: https://doi.org/10.15252/embj.201695290