Protein-Protein Interactions in the Membrane: Sequence, Structural, and Biological Motifs (original) (raw)
. Author manuscript; available in PMC: 2013 Sep 12.
Published in final edited form as: Structure. 2008 Jul;16(7):991–1001. doi: 10.1016/j.str.2008.05.007
Abstract
Single-span transmembrane (TM) helices have structural and functional roles well beyond serving as mere anchors to tether water-soluble domains in the vicinity of the membrane. They frequently direct the assembly of protein complexes and mediate signal transduction in ways analogous to small modular domains in water-soluble proteins. This review highlights different sequence and structural motifs that direct TM assembly and discusses their roles in diverse biological processes. We believe that TM interactions are potential therapeutic targets, as evidenced by natural proteins that modulate other TM interactions and recent developments in the design of TM-targeting peptides.
Introduction
Helical integral membrane proteins are involved in diverse biological processes, from the fusion of sperm and egg at conception to apoptotic events during cell death. The rapidly expanding list of polytopic membrane protein structures has provided insight into how folding and packing of their transmembrane (TM) segments contribute to their diverse functions. More recently, we have begun to understand the importance of single-span TM helices beyond being anchors for more “interesting” water-soluble domains or mere models for multipass protein folding. Like small modular domains, such as leucine zippers that mediate protein complex formation in water-soluble proteins, TMs can direct protein-protein interactions within the membrane (Bechinger, 2000; Call and Wucherpfennig, 2005; Curran and Engelman, 2003; MacKenzie, 2006; Senes et al., 2004) and participate in signal transduction across lipid bilayers (Klein-Seetharaman, 2005).
Here, we will discuss sequence and geometric motifs that are important for interactions between single-pass TM helices, with a special focus on biological function. In addition to organizing these interactions by sequence and structure, they can be broken down into two broad functional categories (Figure 1): (1) relatively static, kinetically stable contacts between TM helices that are important for the assembly of multiprotein complexes; and (2) regulated-switchable interactions involved in signal transduction across bilayers via changes in oligomeric state and/or conformation. Thermodynamically, these groupings are likely to exist in a continuum and be affected by interactions within the water-soluble compartment. However, biologically (and medically) these differences can be significant, because a number of diseases result from changes in these equilibria.
Figure 1. Two General Categories of TM Interactions.
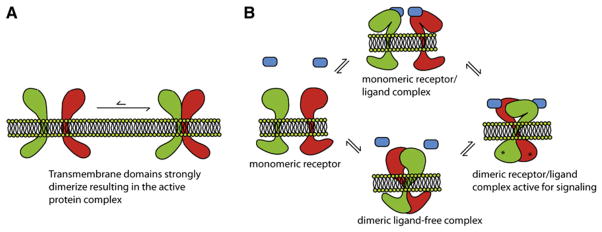
(A) Interacting TM helices can form static associations in the membrane or (B) participate in regulated switchable interactions in a dynamic equilibrium that regulates signaling. A simplified schematic of the coupled equilibria involving a dimeric receptor-ligand complex is depicted; however, signal transduction can potentially travel in both directions across the membrane. The receptor is in equilibrium between the monomeric and dimeric states, and ligand can bind to either. Along the bottom path, ligand binding to a dimeric receptor leads to a conformational change and signal transduction across the membrane. On the top path, the binding of ligand to a monomeric receptor can induce dimerization of the receptor and the subsequent conformational changes. Depending on the local ligand concentration and the expression level of the receptor in the membrane, different cell types and tissues can potentially fine-tune their responses as needed.
We will also discuss a special subclass of TM helices that function as modulators of other TM proteins. Examples of this class of proteins include channel-regulating peptides, such as phos-pholamban, as well as viral proteins that activate growth factor receptors. The surprising sequence specificity of these natural peptides and recent computational methods for the design of novel TM regulating agents point toward potential therapeutic avenues for the targeting of TM interactions.
Geometric Motifs in Multipass Membrane Proteins
What are the geometric characteristics of helix-helix interactions in membrane proteins (Chamberlain et al., 2003) and how does an amino acid sequence predispose the helices to adopt these geometries? Early work focused on distributions of the interhelical angles and distances and on the composition of the side chains packed at the interface. These studies have shown that helices tend to approach somewhat more closely in membrane-soluble proteins than in water-soluble proteins, although the distributions tend to be broad and overlapping (Bowie, 1997; Gimpelev et al., 2004). Distributions of interhelical angles also overlap, but the extent to which this reflects geometric preferences, versus biases associated with random distributions of two vectors, is not clear (Bowie, 1997). Another difficulty with early statistical surveys is that they considered only two parameters (interhelical distance and crossing angle). Six parameters (three Eulerian angles and three distances) are required to define the mutual orientation of two helices (Engel and DeGrado, 2005); more would be required for nonideal helices.
An alternate approach (Figure 2) involves clustering interacting helical pairs according to their three-dimensional similarity (Gimpelev et al., 2004; Walters and DeGrado, 2006). Surprisingly, two-thirds of the helical pairs selected from high-resolution crystal structures of membrane proteins fell within just four clusters, in which Cα atoms from each helix in the pair were within 1.5 Å rmsd of the centroid structure (Walters and DeGrado, 2006). The largest cluster, which comprises 29% of the library, consists of antiparallel helices with left-handed crossing angles of approximately 20° and packing of amino acids with small side-chains that are spaced every seven residues in the sequence. Because Gly, Ala, and Ser (G, A, S) are frequently found at this helix-helix interface, the cluster was designated the antiparallel-GASLeft motif. Right-handed motifs with approximately 40° crossing angles show a similar tendency to segregate small residues to the helix-helix interface, but spaced at four-residue intervals leading to the designation of parallel and antiparallel GASRight motifs.
Figure 2. The Majority of Interacting TM Pairs Use a Limited Number of Structural Motifs.
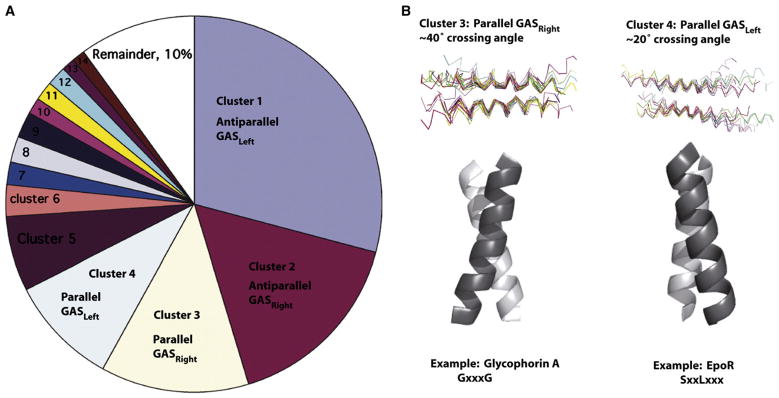
(A) Geometric clustering of interacting TM pairs in the crystallographic database by rmsd places two-thirds of TM helices into four categories: parallel or antiparallel GASLeft, and parallel or antiparallel GASRight (Walters and DeGrado, 2006).
(B) The majority of known interacting monotypic TMs fall into parallel clusters. Parallel GASRight pairs have a ~40° crossing angle and a GX3G-like motif. Parallel GASLeft helices have a ~20° crossing angle and often a heptad repeat of small residues (G/A/S).
It is important to note that kinked structures can be accommodated adjacent to or within many of these motifs, and they might also establish unique packing motifs and hydrogen-bonding networks because of unpaired backbone carbonyls (Senes et al., 2004). Proline residues, which could introduce a bend in an alpha helix are common in membrane proteins (Senes et al., 2000), are overrepresented in interacting TM libraries (Dawson et al., 2002) and have been demonstrated to be critical for membrane protein function (Yohannan et al., 2004).
Sequence Motifs in Single-Pass Membrane Proteins
To date, most interactions between single-span TM helices involve parallel helix packing. For homomeric bundles, this is a consequence of all known single-span proteins being unidirectionally inserted into membranes. In the case of heteromeric interactions, the individual proteins are often evolutionarily related and share a common orientation leading to parallel bundles when their TM helices interact.
Right-Hand-Parallel Helix Packing Motifs
The core geometric and sequence elements of the GASRight motif were first recognized by Engelman and coworkers, who demonstrated that a GX3G sequence in the glycophorin A (GpA) TM domain was necessary for homodimerization and adopted an approximately 40° right-handed crossing angle (Lemmon et al., 1992; MacKenzie et al., 1997; Senes et al., 2000). This motif has since been observed in water-soluble proteins (Kleiger et al., 2002). In the classic GpA GX3G motif, the small Gly residues create a shallow groove that complements the surface of a second helix. The association is stabilized by van der Waals interactions resulting from the excellent geometric fit and also by weak hydrogen bonding between CαH groups on one helix and the carbonyls of a neighboring helix (Senes et al., 2001). Besides the original GpA NMR structure (MacKenzie et al., 1997), there have been recent studies of GX3G containing TM homodimers (Bocharov et al., 2007, 2008), including the structure of the Ff bacteriophage major coat protein, which adopts a noncanonical 27° crossing angle (Wu et al., 2007).
Extensive mutagenesis of the GpA TM has shown that the sequence L75IxxGVxxGVxxT87 is most important for dimerization in SDS-micelles (Lemmon et al., 1992). Subsequent studies by analytical ultracetrifugation and in bacterial reporter systems have largely supported this motif, with some differences between micellar and membrane systems, depending on the hydrophobicity of the mutations (Duong et al., 2007). It is interesting to note the presence of beta-branched amino acids—Ile, Val, and Thr—which have fewer low-energy rotameric conformations and allow helix dimerization to occur at minimal entropic cost (MacKenzie et al., 1997). Recently, it also has been shown that phenylalanine can reinforce GX3G dimerization (Unterreitmeier et al., 2007). Consistent with the GASRight motif found in multi-pass proteins, the small residues Ala and Ser can replace one or both of the Gly residues (Senes et al., 2000, 2004; Walters and DeGrado, 2006), or small amino acids can occur repetitively every four residues to create extended “glycine zippers” (Kim et al., 2005).
Left-Hand-Parallel Helix Packing Motifs
TM interactions consistent with GASLeft packing are also very common in single-pass proteins. Like water-soluble coiled coils (O’Shea et al., 1991), they often have 20° crossing angles and heptad sequence repeats (Langosch and Heringa, 1998; Walters and DeGrado, 2006). For example, the class II MHC α– and β-chains contain heptad repeats of Gly residues at three consecutive “a” positions, with the remaining “a” position being filled by a small residue (Cys or Ser) (Cosson and Bonifacino, 1992). Mutation to many of the Gly residues disrupts the formation of heterodimers between the two chains (Cosson and Bonifacino, 1992). Similarly, Gly residues fill several of the “d” positions. Synthetic peptides containing a similar heptad repeat associate to form dimers in micelles (Lear et al., 2004).

Heptad repeats with Ser at the “a” position have also been found in a number of other structures. Such motifs have been used in the design of ion channel peptides (Lear et al., 1988) and drive dimerization of the erythropoietin receptor TM (Ebie and Fleming, 2007; Kubatzky et al., 2001; Ruan et al., 2004a). Adamian and Liang (2002) have also described an antiparallel “serine zipper” motif in which interacting Ser sidechains are spaced at seven-residue increments in the crystal structures of many transmembrane proteins. Model peptides with a SaxxLdxxx repeat were found to associate to form both parallel and antiparallel dimers by equilibrium thiol-disulfide exchange (North et al., 2006). A variant containing an AaxxLdxxx repeat dimerized with similar affinity, indicating that the Ser might form intrachain H-bonds with the carbonyl at position i-4 and that its small size was the dominant factor for oligomerization (North et al., 2006). By use of a genetic selection from a randomized library of TM sequences, Engelman and coworkers also demonstrated that SxxSSxxT and SxxxSSxxT motifs could strongly drive dimerization in bacterial membranes. Such motifs are potentially consistent with either GASRight or GASLeft (Dawson et al., 2002).
Compared with optimized GX3G motifs in the GASRight family, the stabilities of the left-handed dimers discussed in this section are frequently rather weak. However, as we will see in the next section, hydrogen bonds between more polar groups, such as Asp, Asn, Gln, or Glu, can work in concert with these packing interactions to strongly stabilize left-handed motifs.
Polar Motifs
Strongly polar side chains can play an important role in the association of TM helices in a variety of packing motifs. In some cases, they help to augment or provide geometric specificity to an energetically dominant folding motif. For example, a Thr residue at the helix-helix interface of GpA appears to cooperate with the GX3G motif to increase the affinity and specificity of the inter-action (Duong et al., 2007; Lemmon et al., 1992). A recent NMR structure of the BNIP3 TM homodimer found glycophorin-like packing between small amino acids, as well as extensive hydrogen bonding between Ser and His residues (Bocharov et al., 2007). This structure was consistent with previous mutagenesis and computational studies (Metcalf et al., 2007; Sulistijo and MacKenzie, 2006).
In other cases, a single polar residue can play a dominant role in stabilizing helix-helix associations. Model peptides containing a single Asn, Asp, Gln, or Glu dimerize and/or trimerize in detergents, liposomes (Choma et al., 2000; Gratkowski et al., 2001; Zhou et al., 2000, 2001), and bacterial membranes (Dawson et al., 2003; Ruan et al., 2004b). Oligomerization of an Asn-containing peptide was increased by adding a second Asn at position i+7 (Lear et al., 2003) or an adjacent Ser or Thr residue to form a hydrogen-bonded network (Tatko et al., 2006).
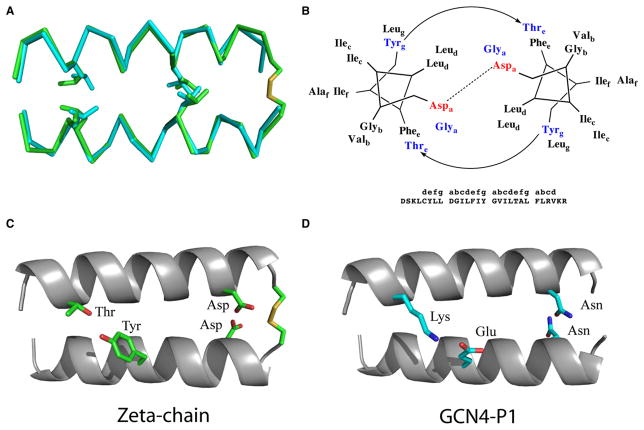
The solution structure of the TM helix of the T cell receptor (TCR) ζζ-chain dimer (Figure 3) nicely demonstrates how H-bonding can establish a left-handed homodimer (Call et al., 2006). Previous studies showed that homodimerization was dependent on an N-terminal interchain disulfide bond and a conserved Asp residue at position i+4 relative to the Cys (Call et al., 2002). The nature of this residue was found to be extremely important, because a Glu mutation maintained the homodimer, whereas Asn, Ser, and Ala mutations reduced dimerization by 60%, 60%, and >80%, respectively. Mixed heterodimers Asp/ Glu, Asp/Asn, and Asp/Ser all formed equally well, whereas Asp/Ala dimerization was reduced by nearly 40%. On the basis of NOEs with water and the mutagenesis data, it was proposed that the two aspartic acids form an extensive hydrogen-bonding network with at least one water molecule, the cysteine carbonyls, and aspartic acid backbone amides. Although the protonation state of the Asp is not yet known, the carboxylates on neighboring chains are in very close proximity (~2.5 Å oxygen-to-oxygen distance) in the computed structure, suggesting that they might form a carboxylate-carboxyl pair with a single proton shared between the two carboxylates (Jasti et al., 2007; Wohlfahrt, 2005).
Figure 3. The TCR-ζ Chain Has a Geometric Motif Similar to the Leucine Zipper GCN4-P1.
(A) Superposition of the TCR-ζ chain TM dimer (Call et al., 2006) and the GCN4 backbones (O’Shea et al., 1991).
(B) Schematic of the crucial interactions within the TCR-ζ helical dimer.
(C) TCR-ζ chain interactions including a Cys-disulfide, Asp-Asp hydrogen bonding, and Thr-Tyr hydrogen bonding.
(D) The analogous GCN4-P1 interactions, including the Asn-Asn and Lys-Glu pairs.
The NMR structure of the ζζ-dimer has a left-handed crossing angle of 20°, which is quite similar to the value seen in left-handed coiled coil peptides. Indeed, the structure superimposes quite well onto the structure of GCN4-P1 (Figure 3A), a two-stranded coiled coil from the yeast transcription factor GCN4. GCN4-P1 and the ζ-chain contain analogous heptads—NxxLxxx and DxxLxxx (Figure 3B), respectively. Although water-soluble coiled-coils often have salt bridges between positions “g” and “e” of successive heptads, a hydrogen bond is formed between Tyr and Thr residues in the ζ-chain, as can be seen by comparing Figure 3C with Figure 3D. Thus, the same basic framework is retained in both cases, but different interactions are used to stabilize the membrane-soluble versus the water-soluble structures.
Biological Significance
The geometric and sequence motifs described above can participate in two broad categories of interactions. In the first, the TMs form relatively static contacts that are necessary for the assembly of a functional protein complex and for proper folding and export from endoplasmic reticulum. In the second, TMs undergo dynamic conformational changes important for signaling; this process can involve a change in association state and/or lateral, vertical, and rotational motions in the membrane (Qiu et al., 2006). We refer to this class of TMs as “switchable.” Switchable TM interactions may not be the dominant force regulating protein-protein interactions, but rather fine-tune the system’s energetics. The biological importance of these interactions is highlighted by the number of disease mutations within single-pass TMs (Table 1). Although many of these mutations are uncharacterized, the involvement of polar or small residues points toward changes in protein-protein interactions within the membrane.
Table 1.
Disease-Associated Single-Pass TM Mutations
| Ectodysplasin | H54T | LLFLGFFGLSLALHLLTLCCYL | X-linked hypohidrotic ectodermal dysplasia (Vincent et al., 2001) |
|---|---|---|---|
| Y | |||
| FXYD2 | G41A | LIFAGLAFIVGLLILLSRRF | Primary hypomagnasemia |
| R | (Meij et al., 2000) | ||
| TACI | A181E | VALVYSTLGLCLCAVLCCFLVAVA | Common variable immunodeficiency |
| E | (Castigli et al., 2005) (Salzer et al., 2005) | ||
| GPIX | A146T | GVLWDVALVAVAALGLALLAGLL | Bernard-Soulier syndrome |
| T | (Wang et al., 2004) | ||
| MPL | W515L | ISLVTALHLVLGLSAVLGLLLLRW | Myelofibrosis with myeloid metaplasia |
| L | (Pikman et al., 2006) | ||
| MPZ | G134R | YGVVLGAVIGGVLGVVLLLLLLF | Charcot-Marie-Tooth disease |
| G138R | R | (Shy et al., 2004) | |
| G138A | R | ||
| A | |||
| Caveolin-3 | P104L | LLSTLLGVPLALLWGFLFACISF | Hypertrophic cardiomyopathy |
| (mouse) | L | (Ohsawa et al., 2004) | |
| Neu/ErbB-2 | V664E | VTFIIATVVGVLLFILVVVVGILI | Oncogenic |
| (rat) | E | (Bargmann et al., 1986) | |
| ErbB-2 | I654V | IISAVIGILLVVVLGVVFGILI | Increased risk of breast cancer |
| (human) | V | (Frank et al., 2005) | |
| FGFR1 | Y372C | YLEIIIYCTGAFLISCMVGSVIVY | Osteoglophonic dysplasia |
| C379R | C | ||
| R | |||
| FGFR2 | S372C | SPDYLEIAIYCIGVFLIACMVVTV | Beare-Stevenson syndrome |
| Y375C | C | ||
| C | |||
| FGFR3 | G370C | GSVYAGILSYGVGFFLFILVVAAV | Thanatophoric dysplasia I, bladder cancer |
| S371C | C | Achondroplasia | |
| Y373C | C | Achondroplasia | |
| G375C | C | Hypochondroplasia | |
| G380R | C | Multiple myeloma | |
| V381E | R | Crouzon syndrome | |
| G382D | E | Bladder cancer | |
| A391E | D | ||
| E | |||
| FGFR4 | G388R | IILYASGSLALAVLLLLAGLY | Tumor progression |
| R | (Li and Hristova, 2006) | ||
| TREM-2 | K186N | SILLLLACIFLIKILAASALWA | Nasu-Hakola disease |
| N | (Paloneva et al., 2002) | ||
| FcγRIIb | I232T | IIVAVVTGIAVAAIVAAVVALIY | Lupus |
| T | (Kono et al., 2005) | ||
| KIT | F522C | LFTPLLIGFVIVAGMMCIIVMILT | Mastocytosis |
| C | (Akin et al., 2004) | ||
| G-CSFR | T617N | IILGLFGLLLLLTCLCGTAWLCC | Acute myeloid leukemia |
| N | (Forbes et al., 2002) |
Static Interactions
Activating Immune Receptors
The biological importance of static TM interactions is clearly illustrated by a system of modular receptors expressed on the surface of hematopoeitic cells. These complexes use a network of charged residues to connect at least one ligand-binding protein that generally lacks intracellular signaling domains with a second group of signaling adaptor proteins (Call et al., 2002; Feng et al., 2005, 2006). The best-characterized member of this family is the TCR. The ζζ-homodimer structure (Call et al., 2006) described above is just one component of the larger TCR complex. Positively charged Arg and Lys residues on the TCRαβ TMs interact with negatively charged Asp and Glu residues on the CD3δε and CD3γε heterodimers and the ζζ-homodimer (Call et al., 2002). These interactions are necessary for proper assembly of the complex in the ER and subsequent transport to the cell surface (Call et al., 2002).
This high degree of specificity is also present in the other activating immune receptors that interact with the signaling modules Fcγ, DAP10, and DAP12 (Table 2; Feng et al., 2005, 2006). The importance of these interactions is illustrated by a Lys to Asn mutation in the TREM-2 receptor that blocks assembly with the DAP12 adaptor protein and results in Nasu-Hakola disease (Paloneva et al., 2002). Like the TCR, the B cell receptor (BCR) also pairs a signaling dimer (Ig-αβ) with a ligand binding receptor (mIg). Although the Ig-αβ chains contain Glu and Gln residues that may be analogous to the Asp and Glu residues in the CD3 complexes, there are no corresponding positively charged residues in the mIg TM (Table 2). Recent studies (Dylke et al., 2007) point toward specific interactions between the mIgM and Ig-αβ TM domains and early mutagenesis to polar Ser, Thr, and Tyr residues support a role for polar interactions within the membrane (Grupp et al., 1993; Pleiman et al., 1994; Stevens et al., 1994).
Table 2.
TM Sequences from Ligand-Binding Immune Receptors and Their Interacting Signaling Modules
| Ligand-binding module | Signaling module | ||
|---|---|---|---|
| TCRα | VIGFRILLLKVAGFNLLMTL | ζ | LCYLLDGILFIYGVILTALFL |
| VIGFRILLLKVAGFNLLMTL | CD3ε | VMSVATIVIVDICITGGLLLLVYYW | |
| CD3δ | PATVAGIIVTDVIATLLLALGVFCF | ||
| TCRβ | TILYEILLGKATLYAVLVSALVL | CD3ε | VMSVATIVIVDICITGGLLLLVYYW |
| CD3γ | ISGFLFAEIVSIFVLAVGVYFI | ||
| NKG2C | VLGIICIVLMATVLKTIVLIPFL | ||
| KIR2DS2 | (Type II TM, C-type lectin) | ||
| VLIGTSVVKIPFTILLFFLL | |||
| KIR3DS1 | ILIGTSVVKIPFTILLFFLL | ||
| NKp44 | LVPVFCGLLVAKSLVLSALLVWW | ||
| TREM-1 | VILLAGGFLSKSLVFSVLFAVTL | ||
| TREM-2 | LLLLACIFLIKILAASALWAAAW | DAP12 | VLAGIVMGDLVLTVLIALAVYFL |
| IREM-2 | FLLVVLLKLPLLLSMLGAVFWV | ||
| SIRPβ1 | LVALLLGPKLLLVVGVSAIYICW | ||
| SILRβ | VALAVALKTVILGLLCLLLLWW | ||
| CD200RLa | LIILYVKLSLFVVILVTTGFVFF | ||
| NKG2D | PFFFCCFIAVAMGIRFIIMVAIW (Type II TM, C-type lectin) | DAP10 | LLAGLVAADAVASLLIVGAVFL |
| FcαRI | LIRMAVAGLVLVALLAILV | ||
| NKp46 | LLRMGLAFLVLVALVWFLV | ||
| ILT1 | LIRMGVAGLVLVVLGILLF | Fcγ | LCYILDAILFLYGIVLTLLYC |
| GPVI | LVRICLGAVILIILAGFLA | ||
| OSCAR | LVRLGLAGLVLISLGALVTF | ||
| mIgM | LWATASTFIVLFLLSLFYSTTVT | ||
| mIgG | LWTTLSTFVALFILTLLYSGIVT | Igα | IITAEGIILLFCAVVPGTLLL |
| mIgA | LWTTITIFITLFLLSVCYSATVT | Igβ | IILIQTLLIILFIIVPIFLLL |
| mIgD | LWTGLCIFAALFLLSVSYSAALT |
The mechanism of signal transduction by activating immune receptors may differ between family members and is beyond the scope of this review. Receptor clustering and conformational changes are likely important, but other modes have also been proposed and reviewed (Choudhuri and van der Merwe, 2007). Therefore, it is possible that these seemingly static associations between charged residues in the membrane, which are necessary for assembly, are flexible enough to propagate information across the bilayer.
Switchable Interactions
The TM helices described in this section participate with soluble domains in finely tuned equilibria that govern signal transduction across biological membranes. In each case, conformational changes are propagated through the membrane via changes in the oligomerization and/or orientation of the TM helices. Because they must be able to switch between states without large energetic penalties, it is not surprising that these proteins contain GASRight or GASLeft motifs that rely on van der Waals interactions, as opposed to strongly polar or charged residues. In fact, there are a number of activating disease mutations in Table 1 that place Cys, Asp, or Glu residues in the TM, which could lock the proteins in active conformations.
Ion channels represent one class of proteins in which conformational changes give rise to gating events that open and close the pore. Although work in this field is beyond the scope of this review, the M2 proton channel from influenza virus should be mentioned, because the transmembrane domain of this protein is a homotetramer formed by association of four copies of a single TM helix. Recent NMR and crystal structures have provided possible mechanisms for the pH-dependent conduction of the channel (Schnell and Chou, 2008; Stouffer et al., 2008).
Cytokine Receptors and Receptor Tyrosine Kinases
Cytokine receptors and receptor tyrosine kinases (RTKs) are unrelated families of proteins that regulate cell proliferation through evolutionarily convergent mechanisms involving ligand-dependent activation of intracellular kinases. The simplest model for their signaling is an equilibrium between inactive monomers and ligand-bound dimers. However, several bodies of evidence discussed below suggest that TM interactions may play important roles in more complicated equilibria (Figure 1).
In the case of the erythropoietin receptor (EpoR), which is a cytokine receptor that directly regulates the production of red blood cells, it has been hypothesized that TM interactions may be responsible for the apparent increased activity of the mouse receptor relative to the human receptor (Ebie and Fleming, 2007). The human receptor lacks an SaxxdLxxx heptad repeat that is found in the murine protein and is necessary for increased TM dimerization in vitro (Ebie and Fleming, 2007; Kubatzky et al., 2001; Ruan et al., 2004a) and for the ligand-independent oligomerization of inactive receptors and increased signal transduction at low ligand concentrations (Constantinescu et al., 2001b). This interaction is quite specific, and activation is believed to be coupled to rotation of the TM in the membrane (Constantinescu et al., 2001a; Seubert et al., 2003). In contrast to EpoR, the biological significance of TM interactions in normal RTK physiology remains unclear. However the prevalence of TM sequences consistent with GASRight packing and the number of activating disease mutations in growth factor receptor TMs (Table 1) are suggestive. These topics are discussed extensively in a recent review (Li and Hristova, 2006).
Integrins
Like activating immune receptors, such as the TCR, integrins are modular signaling complexes that recognize diverse ligands. There are 24 known integrin α/β heterodimers composed of combinations of 18 α subunits and eight β subunits (Bennett, 2005; Luo and Springer, 2006). Most integrins exist in equilibrium between an inactive resting state and an activated state capable of binding its extracellular ligands (Figure 4). Integrin TMs and their short cytosolic domains are in close contact in the inactive receptor, and activation is accompanied by their separation (Kim et al., 2003; Litvinov et al., 2004). This process is reversible and is only part of a larger network of linked equilibria involving integrin extracellular, transmembrane, and cytoplasmic domains and a host of interacting proteins. A push/pull mechanism (Li et al., 2005b) emphasizes these linked equilibria and postulates that any interaction or mutation that physically disrupts and αβ-chain TM helix-helix interactions will push the integrin toward the activated state, whereas cytoplasmic or TM proteins that preferentially interact with the cytoplasmic domains in either the inactive or active state of the integrin will pull the equilibrium toward one of these states. Most integrin TM domains contain GASRight-like sequences consistent with right-handed helical packing. In the case of the platelet integrin αIIbβ3, there are important TM interactions between a GX3G sequence in the α chain and bulky hydrophobic residues in the β3-chain (Li et al., 2005b; Luo et al., 2004, 2005; Partridge et al., 2004). Interestingly, the β3-chain contains an SX3A on the opposite face of the helix that may participate in as of yet unknown interactions.
Figure 4. Integrin TMs Make Switchable Heterodimeric Interactions.
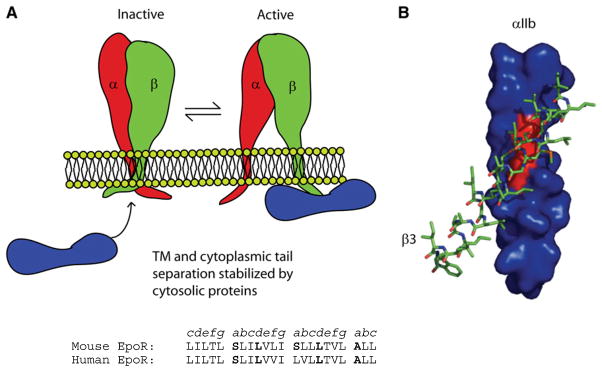
(A) Integrins are held inactive by interactions between α- and β-chain TM and soluble domains. Interactions between cytosolic proteins and integrin cytosolic tails stabilize TM separation and activation of the extracellular integrin ligand-binding site (Bennett, 2005).
(B) A computational model of the β3-integrin TM domain packing against the GX3G motif of the αIIb-TM domain. Adapted from Li et al. (2005b).
TM-Regulating Peptides
Switchable interactions between TM domains can be regulated by other TM proteins that either inhibit or stabilize TM oligomerization. Viruses have evolved peptides that interact with the TM domains of growth factor and cytokine receptors to cause cellular transformation. The spleen focal forming virus (SFFV) protein gp55-P transforms cells through ligand-independent TM interactions with mouse EpoR (Constantinescu et al., 1999). The type 1 bovine papillomavirus (BPV) protein E5 also causes cellular transformation by activating the RTK platelet-derived growth factor β receptor (PDGFRβ) (DiMaio and Mattoon, 2001; Mattoon et al., 2001). E5 is a disulfide-linked type II TM protein that is predicted to make a homodimeric scaffold for the assembly of the PDGFβ receptor TMs into an antiparallel tetrameric coiled coil, bringing the receptor kinase domains in close proximity (Freeman-Cook et al., 2004, 2005).
Single-span TM proteins have also evolved to regulate ion channels through specific TM-TM interactions. Phospholamban (PLB) and sarcolipin (SLN) are homologous 52 aa and 31 aa TM proteins that regulate the sarcoendoplasmic reticulum calcium transport ATPase (SERCA), which pumps calcium from the cytosol into the sarcoendoplasmic reticulum in cardiac muscle cells. Key residues within the TM domain of PLB drive pentamerization in detergents (Arkin et al., 1994). The solution NMR structure (Figure 5) of Oxenoid and Chou (2005) demonstrated a pentameric left handed-coiled coil with a heptad Leu/Ile zipper motif in which Ile residues occupy the “a” positions (Ile33, Ile40, and Ile47) and Leu residues occupy “d” positions (Leu37, Leu44, and Leu51). This motif had previously been shown to mediate tetramer formation in water-soluble coiled coils (Harbury et al., 1993) and had been proposed to stabilize the pentameric form of phospholamban B (Karim et al., 1998; Simmerman et al., 1996). Mutations to the TM that blocked oligomerization result in increased inhibition of SERCA by PLB (Fujii et al., 1989; Kimura et al., 1997), whereas mutations to the opposite face of the TM helix prevented inhibition of SERCA by PLB (Kimura et al., 1997), suggesting that SERCA inhibition may depend on the monomer form of PLB. Recent studies, however, are consistent with the PLB pentamer interacting with SERCA, suggesting that the mechanisms of transporter regulation may be more complicated than previously thought (Stokes et al., 2006). SLN also appears to make similar direct interactions with SERCA, though it is not known whether it too has a homo-oligomeric form, as does PLB (Hughes et al., 2007).
Figure 5. Phospholamban, Sarcolipin, and the FXYD Proteins Regulate Channel Proteins through TM Interactions.
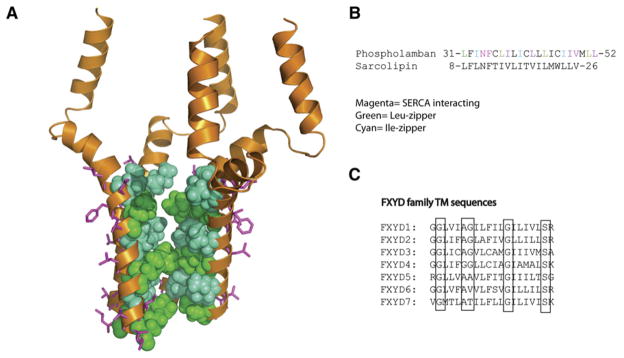
(A) An NMR structure of the phospholamban homopentamer (Oxenoid and Chou, 2005). In magenta are residues that mediate interactions with the SERCA channel. They are on the opposite face from the Leu- and Ile-zippers that mediate pentamerization (red and orange, respectively).
(B) TM sequence of phospholamban showing important residues and the related TM sequence from sarcolipin.
(C) Alignment of the FXYD protein family TM sequences, which regulate the Na/K-ATPase. Highly conserved and functionally important residues are highlighted. A mutation in the second conserved glycine, Gly41Arg, in the FXYD2 TM results in dominant negative primary hypomagnesemia due to aggregation of FXYD2 in the cytoplasm and misrouting of the Na,K-ATPase (Meij et al., 2000).
The seven FXYD proteins (FXYD1–FXYD7) are evolutionarily unrelated to PLB and SLN but also function to regulate ion channels through TM interactions. They are tissue-specific modulators of the Na,K-ATPase, which governs the basic membrane potential that powers membrane transporters and maintains cell volume (Geering, 2006). With the exception of FXYD3, which has an uncleaved signal sequence, all FXYD proteins are single-pass type I TM proteins that contain an extracellular FXYD domain and conserved TM domains that are necessary for their functions (Figure 5C; Geering, 2006). Mutagenesis suggests that small residues at positions i, i+4, and i+11 along one face of the TM helix are important for interactions with the ATPase (Crambert et al., 2004; Li et al., 2005a) and that flanking residues specifically and differentially regulate Na+ and K+ affinity (Li et al., 2005a; Lindzen et al., 2003).
Exogenous Regulatory Peptides
A number of laboratories have used exogenous TM peptides to perturb signaling pathways believed to be dependent on TM interactions. Peptides containing the ErbB2 TM were shown to partially inhibit signaling at low concentrations of ligand in a GX3G-dependent fashion (Bennasroune et al., 2004). TM peptides that mimic the TCR TM have also been used to inhibit T cell function (Manolios et al., 1997; Quintana et al., 2007). Exogenous peptides based on the phospholamban (Afara et al., 2006) and FXYD2 (Zouzoulas et al., 2003) sequences also perturb SERCA and Na/K-ATPase function, respectively.
We have targeted the heterodimeric interface of the integrin αIIbβ3 with TM peptides from the wild-type integrin (Yin et al., 2006). Peptides that block the heteromeric association between αIIb and β3 should activate the integrin. Indeed, a peptide spanning the sequence of the αIIb TM specifically activates αIIbβ3, presumably by competing for this interface through homodimeric interactions. To design peptides with greater specificity and affinity, we developed a computational method for designing TM peptides to specifically target protein TM domains: Computed Helical Anti-Membrane Protein (CHAMP) (Yin et al., 2007). The GX3G-like motifs within the αIIb- and α_v_-integrin TM domains were targeted to demonstrate the specificity of the technique. Each of these integrin subunits is known to pair with a common β3 to create either αIIbβ3 or α_v_β3, which recognize different extracellular ligands. Sequence analysis showed that αIIb- and α_v_-integrin TM domains each have a high propensity to associate with a second helix in a parallel GASRight motif, indicating that a peptide that also had a strong propensity to adopt this motif would be likely to have a high affinity for the integrin TM sequence (Figure 2). Thus, backbones of helical pairs taken from the crystallographic database interacting with this geometry were chosen as potential templates for the design of a CHAMP peptide. The sequence of the integrin TM helical target was threaded onto one of the two helices, and the amino acid residues on the opposite helix was selected using a side chain repacking algorithm. The anti-αIIb and anti-α_v_ CHAMP peptides bound specifically to their intended integrin TMs and specifically activated receptor adhesion to their respective ligands.
Conclusions
As discussed above, there are a host of pathologic mutations in single-span transmembrane proteins that likely affect TM interactions. As the structural and sequence motifs that govern TM interactions become increasingly apparent, we will better understand the pathophysiology of these mutations. Similarly, we are beginning to better understand the roles of wild-type TM sequences in such medically important processes as hemostasis and cell proliferation. These structural and mechanistic insights are just now beginning to allow us to develop technologies such as CHAMPs and potentially even small molecules that can interact with transmembrane domains as molecular probes of biological function or new classes of targeted therapeutics.
Acknowledgments
We sincerely thank Joel S. Bennett, Jason Donald, and Alessandro Senes for their thoughtful discussions and helpful input during the preparation of this manuscript.
References
- Adamian L, Liang J. Interhelical hydrogen bonds and spatial motifs in membrane proteins: polar clamps and serine zippers. Proteins. 2002;47:209–218. doi: 10.1002/prot.10071. [DOI] [PubMed] [Google Scholar]
- Afara MR, Trieber CA, Glaves JP, Young HS. Rational design of peptide inhibitors of the sarcoplasmic reticulum calcium pump. Biochemistry. 2006;45:8617–8627. doi: 10.1021/bi0523761. [DOI] [PubMed] [Google Scholar]
- Akin C, Fumo G, Yavuz AS, Lipsky PE, Neckers L, Metcalfe DD. A novel form of mastocytosis associated with a transmembrane c-kit mutation and response to imatinib. Blood. 2004;103:3222–3225. doi: 10.1182/blood-2003-11-3816. [DOI] [PubMed] [Google Scholar]
- Arkin IT, Adams PD, MacKenzie KR, Lemmon MA, Brünger AT, Engelman DM. Structural organization of the pentameric transmembrane α-helices of phospholamban, a cardiac ion channel. EMBO J. 1994;13:4757–4764. doi: 10.1002/j.1460-2075.1994.tb06801.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bargmann CI, Hung MC, Weinberg RA. Multiple independent activations of the neu oncogene by a point mutation altering the transmembrane domain of p185. Cell. 1986;45:649–657. doi: 10.1016/0092-8674(86)90779-8. [DOI] [PubMed] [Google Scholar]
- Bechinger B. Biophysical investigations of membrane perturbations by polypeptides using solid-state NMR spectroscopy. Mol Membr Biol. 2000;17:135–142. doi: 10.1080/09687680050197365. [DOI] [PubMed] [Google Scholar]
- Bennasroune A, Fickova M, Gardin A, Dirrig-Grosch S, Aunis D, Cremel G, Hubert P. Transmembrane peptides as inhibitors of ErbB receptor signaling. Mol Biol Cell. 2004;15:3464–3474. doi: 10.1091/mbc.E03-10-0753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennett JS. Structure and function of the platelet integrin α(IIb)β(3) J Clin Invest. 2005;115:3363–3369. doi: 10.1172/JCI26989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bocharov EV, Pustovalova YE, Pavlov KV, Volynsky PE, Goncharuk MV, Ermolyuk YS, Karpunin DV, Schulga AA, Kirpichnikov MP, Efremov RG, et al. Unique dimeric structure of BNip3 transmembrane domain suggests membrane permeabilization as a cell death trigger. J Biol Chem. 2007;282:16256–16266. doi: 10.1074/jbc.M701745200. [DOI] [PubMed] [Google Scholar]
- Bocharov EV, Mineev KS, Volynsky PE, Ermolyuk YS, Tkach EN, Sobol AG, Chupin VV, Kirpichnikov MP, Efremov RG, Arseniev AS. Spatial structure of the dimeric transmembrane domain of the growth factor receptor ErbB2 presumably corresponding to the receptor active state. J Biol Chem. 2008;283:6950–6956. doi: 10.1074/jbc.M709202200. [DOI] [PubMed] [Google Scholar]
- Bowie JU. Helix packing angle preferences. Nat Struct Biol. 1997;4:915–917. doi: 10.1038/nsb1197-915. [DOI] [PubMed] [Google Scholar]
- Call ME, Wucherpfennig KW. The T cell receptor: critical role of the membrane environment in receptor assembly and function. Annu Rev Immunol. 2005;23:101–125. doi: 10.1146/annurev.immunol.23.021704.115625. [DOI] [PubMed] [Google Scholar]
- Call ME, Pyrdol J, Wiedmann M, Wucherpfennig KW. The organizing principle in the formation of the T cell receptor-CD3 complex. Cell. 2002;111:967–979. doi: 10.1016/s0092-8674(02)01194-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Call ME, Schnell JR, Xu CQ, Lutz RA, Chou JJ, Wucherpfennig KW. The structure of the ζζ transmembrane dimer reveals features essential for its assembly with the T cell receptor. Cell. 2006;127:355–368. doi: 10.1016/j.cell.2006.08.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castigli E, Wilson SA, Garibyan L, Rachid R, Bonilla F, Schneider L, Geha RS. TACI is mutant in common variable immunodeficiency and IgA deficiency. Nat Genet. 2005;37:829–834. doi: 10.1038/ng1601. [DOI] [PubMed] [Google Scholar]
- Chamberlain AK, Faham S, Yohannan S, Bowie JU. Construction of helix-bundle membrane proteins. Adv Protein Chem. 2003;63:19–46. doi: 10.1016/s0065-3233(03)63002-0. [DOI] [PubMed] [Google Scholar]
- Choma C, Gratkowski H, Lear JD, DeGrado WF. Asparagine-mediated self-association of a model transmembrane helix. Nat Struct Biol. 2000;7:161–166. doi: 10.1038/72440. [DOI] [PubMed] [Google Scholar]
- Choudhuri K, van der Merwe PA. Molecular mechanisms involved in T cell receptor triggering. Semin Immunol. 2007;19:255–261. doi: 10.1016/j.smim.2007.04.005. [DOI] [PubMed] [Google Scholar]
- Constantinescu SN, Liu XD, Beyer W, Fallon A, Shekar S, Henis YI, Smith SO, Lodish HF. Activation of the erythropoietin receptor by the gp55-P viral envelope protein is determined by a single amino acid in its transmembrane domain. EMBO J. 1999;18:3334–3347. doi: 10.1093/emboj/18.12.3334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Constantinescu SN, Huang LJS, Nam HS, Lodish HF. The erythropoietin receptor cytosolic juxtamembrane domain contains an essential, precisely oriented, hydrophobic motif. Mol Cell. 2001a;7:377–385. doi: 10.1016/s1097-2765(01)00185-x. [DOI] [PubMed] [Google Scholar]
- Constantinescu SN, Keren T, Socolovsky M, Nam H, Henis YI, Lodish HF. Ligand-independent oligomerization of cell-surface erythropoietin receptor is mediated by the transmembrane domain. Proc Natl Acad Sci USA. 2001b;98:4379–4384. doi: 10.1073/pnas.081069198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cosson P, Bonifacino JS. Role of transmembrane domain interactions in the assembly of class II MHC molecules. Science. 1992;258:659–662. doi: 10.1126/science.1329208. [DOI] [PubMed] [Google Scholar]
- Crambert G, Li CM, Swee LK, Geering K. FXYD7, mapping of functional sites involved in endoplasmic reticulum export, association with and regulation of Na,K-ATPase. J Biol Chem. 2004;279:30888–30895. doi: 10.1074/jbc.M313494200. [DOI] [PubMed] [Google Scholar]
- Curran AR, Engelman DM. Sequence motifs, polar interactions and conformational changes in helical membrane proteins. Curr Opin Struct Biol. 2003;13:412–417. doi: 10.1016/s0959-440x(03)00102-7. [DOI] [PubMed] [Google Scholar]
- Dawson JP, Weinger JS, Engelman DM, Sturgis JN. Motifs of serine and threonine can drive association of transmembrane helices. J Mol Biol. 2002;316:799–805. doi: 10.1006/jmbi.2001.5353. [DOI] [PubMed] [Google Scholar]
- Dawson JP, Melnyk RA, Deber CM, Engelman DM. Sequence context strongly modulates association of polar residues in transmembrane helices. J Mol Biol. 2003;331:255–262. doi: 10.1016/s0022-2836(03)00714-9. [DOI] [PubMed] [Google Scholar]
- DiMaio D, Mattoon D. Mechanisms of cell transformation by papillomavirus E5 proteins. Oncogene. 2001;20:7866–7873. doi: 10.1038/sj.onc.1204915. [DOI] [PubMed] [Google Scholar]
- Duong MT, Jaszewski TM, Fleming KG, MacKenzie KR. Changes in apparent free energy of helix-helix dimerization in a biological membrane due to point mutations. J Mol Biol. 2007;371:422–434. doi: 10.1016/j.jmb.2007.05.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dylke J, Lopes J, Dang-Lawson M, Machtaler S, Matsuuchi L. Role of the extracellular and transmembrane domain of Ig-α/β in assembly of the B cell antigen receptor (BCR) Immunol Lett. 2007;112:47–57. doi: 10.1016/j.imlet.2007.06.005. [DOI] [PubMed] [Google Scholar]
- Ebie AZ, Fleming KG. Dimerization of the erythropoietin receptor transmembrane domain in micelles. J Mol Biol. 2007;366:517–524. doi: 10.1016/j.jmb.2006.11.035. [DOI] [PubMed] [Google Scholar]
- Engel DE, DeGrado WF. A-α linking motifs and interhelical orientations. Proteins. 2005;61:325–337. doi: 10.1002/prot.20522. [DOI] [PubMed] [Google Scholar]
- Feng J, Call ME, Wucherpfennig KW. The assembly of diverse immune receptors is focused on a polar membrane-embedded interaction site. PLoS Biol. 2006;4:768–779. doi: 10.1371/journal.pbio.0040142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng J, Garrity D, Call ME, Moffett H, Wucherpfennig KW. Convergence on a distinctive assembly mechanism by unrelated families of activating immune receptors. Immunity. 2005;22:427–438. doi: 10.1016/j.immuni.2005.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forbes LV, Gale RE, Pizzey A, Pouwels K, Nathwani A, Linch DC. An activating mutation in the transmembrane domain of the granulocyte colony-stimulating factor receptor in patients with acute myeloid leukemia. Oncogene. 2002;21:5981–5989. doi: 10.1038/sj.onc.1205767. [DOI] [PubMed] [Google Scholar]
- Frank B, Hemminki K, Wirtenberger M, Bermejo JL, Bugert P, Klaes R, Schmutzler RK, Wappenschmidt B, Bartram CR, Burwinkel B. The rare ERBB2 variant Ile654Val is associated with an increased familial breast cancer risk. Carcinogenesis. 2005;26:643–647. doi: 10.1093/carcin/bgh342. [DOI] [PubMed] [Google Scholar]
- Freeman-Cook LL, Dixon AM, Frank JB, Xia Y, Ely L, Gerstein M, Engelman DM, DiMaio D. Selection and characterization of small random transmembrane proteins that bind and activate the platelet-derived growth factor β receptor. J Mol Biol. 2004;338:907–920. doi: 10.1016/j.jmb.2004.03.044. [DOI] [PubMed] [Google Scholar]
- Freeman-Cook LL, Edwards APB, Dixon AM, Yates KE, Ely L, Engelman DM, DiMaio D. Specific locations of hydrophilic amino acids in constructed transmembrane ligands of the platelet-derived growth factor β receptor. J Mol Biol. 2005;345:907–921. doi: 10.1016/j.jmb.2004.10.072. [DOI] [PubMed] [Google Scholar]
- Fujii J, Maruyama K, Tada M, MacLennan DH. Expression and site-specific mutagenesis of phospholamban: studies of residues involved in phosphorylation and pentamer formation. J Biol Chem. 1989;264:12950–12955. [PubMed] [Google Scholar]
- Geering K. FXYD proteins: new regulators of Na-k-ATPase. Am J Physiol Renal Physiol. 2006;290:F241–F250. doi: 10.1152/ajprenal.00126.2005. [DOI] [PubMed] [Google Scholar]
- Gimpelev M, Forrest LR, Murray D, Honig B. Helical packing patterns in membrane and soluble proteins. Biophys J. 2004;87:4075–4086. doi: 10.1529/biophysj.104.049288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gratkowski H, Lear JD, DeGrado WF. Polar sidechains drive the association of model, transmembrane peptides. Proc Natl Acad Sci USA. 2001;98:880–885. doi: 10.1073/pnas.98.3.880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grupp SA, Campbell K, Mitchell RN, Cambier JC, Abbas AK. Signaling-defective mutants of the B lymphocyte antigen receptor fail to associate with Ig-α and Ig-β/γ. J Biol Chem. 1993;268:25776–25779. [PubMed] [Google Scholar]
- Harbury PB, Zhang T, Kim PS, Alber T. A switch between two-, three-, and four-stranded coiled coils. Science. 1993;262:1401–1407. doi: 10.1126/science.8248779. [DOI] [PubMed] [Google Scholar]
- Hughes E, Clayton JC, Kitmitto A, Esmann M, Middleton DA. Solid-state NMR and functional measurements indicate that the conserved tyrosine residues of sarcolipin are involved directly in the inhibition of SERCA1. J Biol Chem. 2007;282:26603–26613. doi: 10.1074/jbc.M611668200. [DOI] [PubMed] [Google Scholar]
- Jasti J, Furukawa H, Gonzales EB, Gouaux E. Structure of acid-sensing ion channel 1 at 1.9A resolution and low pH. Nature. 2007;449:293–294. doi: 10.1038/nature06163. [DOI] [PubMed] [Google Scholar]
- Karim CB, Stamm JD, Karim J, Jones LR, Thomas DD. Cysteine reactivity and oligomeric structures of phospholamban and its mutants. Biochemistry. 1998;37:12074–12081. doi: 10.1021/bi980642n. [DOI] [PubMed] [Google Scholar]
- Kim M, Carman CV, Springer TA. Bidirectional transmembrane signaling by cytoplasmic domain separation in integrins. Science. 2003;301:1720–1725. doi: 10.1126/science.1084174. [DOI] [PubMed] [Google Scholar]
- Kim S, Jeon TJ, Oberai A, Yang D, Schmidt JJ, Bowie JU. Transmembrane glycine zippers: physiological and pathological roles in membrane proteins. Proc Natl Acad Sci USA. 2005;102:14278–14283. doi: 10.1073/pnas.0501234102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimura Y, Kurzydlowski K, Tada M, MacLennan DH. Phospholamban inhibitory function is activated by depolymerization. J Biol Chem. 1997;272:15061–15064. doi: 10.1074/jbc.272.24.15061. [DOI] [PubMed] [Google Scholar]
- Kleiger G, Grothe R, Mallick P, Eisenberg D. GXXXG and AXXXA: common α-helical interaction motifs in proteins, particularly in extremophiles. Biochemistry. 2002;41:5990–5997. doi: 10.1021/bi0200763. [DOI] [PubMed] [Google Scholar]
- Klein-Seetharaman J. Dual role of interactions between membranous and soluble portions of helical membrane receptors for folding and signaling. Trends Pharmacol Sci. 2005;26:183–189. doi: 10.1016/j.tips.2005.02.009. [DOI] [PubMed] [Google Scholar]
- Kono H, Kyogoku C, Suzuki T, Tsuchiya N, Honda H, Yamamoto K, Tokunaga K, Honda ZI. Fc γ RIIB Ile232Thr transmembrane polymorphism associated with human systemic lupus erythematosus decreases affinity to lipid rafts and attenuates inhibitory effects on B cell receptor signaling. Hum Mol Genet. 2005;14:2881–2892. doi: 10.1093/hmg/ddi320. [DOI] [PubMed] [Google Scholar]
- Kubatzky KF, Ruan W, Gurezka R, Cohen J, Ketteler R, Watowich SS, Neumann D, Langosch D, Klingmuller U. Self assembly of the transmembrane domain promotes signal transduction through the erythropoietin receptor. Curr Biol. 2001;11:110–115. doi: 10.1016/s0960-9822(01)00018-5. [DOI] [PubMed] [Google Scholar]
- Langosch D, Heringa J. Interaction of transmembrane helices by a knobs-into-holes packing characteristic of soluble coiled coils. Proteins. 1998;31:150–159. doi: 10.1002/(sici)1097-0134(19980501)31:2<150::aid-prot5>3.0.co;2-q. [DOI] [PubMed] [Google Scholar]
- Lear JD, Wasserman ZR, DeGrado WF. Synthetic amphiphilic peptide modes for protein ion channels. Science. 1988;240:1177–1181. doi: 10.1126/science.2453923. [DOI] [PubMed] [Google Scholar]
- Lear JD, Gratkowski H, Adamian L, Liang J, DeGrado WF. Position-dependence of stabilizing polar interactions of asparagine in transmembrane helical bundles. Biochemistry. 2003;42:6400–6407. doi: 10.1021/bi020573j. [DOI] [PubMed] [Google Scholar]
- Lear JD, Stouffer AL, Gratkowski H, Nanda V, Degrado WF. Association of a model transmembrane peptide containing gly in a heptad sequence motif. Biophys J. 2004;87:3421–3429. doi: 10.1529/biophysj.103.032839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemmon MA, Flanagan JM, Treutlein HR, Zhang J, Engelman DM. Sequence specificity in the dimerization of transmembrane α helices. Biochemistry. 1992;31:12719–12725. doi: 10.1021/bi00166a002. [DOI] [PubMed] [Google Scholar]
- Li E, Hristova K. Role of receptor tyrosine kinase transmembrane domains in cell signaling and human pathologies. Biochemistry. 2006;45:6241–6251. doi: 10.1021/bi060609y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li C, Crambert G, Thuillard D, Roy S, Schaer D, Geering K. Role of the transmembrane domain of FXYD7 in structural and functional interactions with Na,K-ATPase. J Biol Chem. 2005a;280:42738–42743. doi: 10.1074/jbc.M508451200. [DOI] [PubMed] [Google Scholar]
- Li W, Metcalf DG, Gorelik R, Li R, Mitra N, Nanda V, Law PB, Lear JD, Degrado WF, Bennett JS. A push-pull mechanism for regulating integrin function. Proc Natl Acad Sci USA. 2005b;102:1424–1429. doi: 10.1073/pnas.0409334102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindzen M, Aizman R, Lifshitz Y, Lubarski I, Karlish SJD, Garty H. Structure-function relations of interactions between Na,K-ATPase, the γ subunit, and corticosteroid hormone-induced factor. J Biol Chem. 2003;278:18738–18743. doi: 10.1074/jbc.M213253200. [DOI] [PubMed] [Google Scholar]
- Litvinov RI, Nagaswami C, Vilaire G, Shuman H, Bennett JS, Weisel JW. Functional and structural correlations of individual α IIb β 3 molecules. Blood. 2004;104:3979–3985. doi: 10.1182/blood-2004-04-1411. [DOI] [PubMed] [Google Scholar]
- Luo BH, Springer TA. Integrin structures and conformational signaling. Curr Opin Cell Biol. 2006;18:579–586. doi: 10.1016/j.ceb.2006.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo BH, Springer TA, Takagi J. A specific interface between integrin transmembrane helices and affinity for ligand. PLoS Biol. 2004;2:776–786. doi: 10.1371/journal.pbio.0020153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo BH, Carman CV, Takagi J, Springer TA. Disrupting integrin transmembrane domain heterodimerization increases ligand binding affinity, not valency or clustering. Proc Natl Acad Sci USA. 2005;102:3679–3684. doi: 10.1073/pnas.0409440102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacKenzie KR. Folding and stability of α-helical integral membrane proteins. Chem Rev. 2006;106:1931–1977. doi: 10.1021/cr0404388. [DOI] [PubMed] [Google Scholar]
- MacKenzie KR, Prestegard JH, Engelman DM. A transmembrane helix dimer: structure and implications. Science. 1997;276:131–133. doi: 10.1126/science.276.5309.131. [DOI] [PubMed] [Google Scholar]
- Manolios N, Collier S, Taylor J, Pollard J, Harrison LC, Bender V. T-cell antigen receptor transmembrane peptides modulate T-cell function and T cell-mediated disease. Nat Med. 1997;3:84–88. doi: 10.1038/nm0197-84. [DOI] [PubMed] [Google Scholar]
- Mattoon D, Gupta K, Doyon J, Loll PJ, DiMaio D. Identification of the transmembrane dimer interface of the bovine papillomavirus E5 protein. Oncogene. 2001;20:3824–3834. doi: 10.1038/sj.onc.1204523. [DOI] [PubMed] [Google Scholar]
- Meij IC, Koenderink JB, van Bokhoven H, Assink KFH, Groenestege WT, de Pont J, Bindels RJM, Monnens LAH, van den Heuvel L, Knoers N. Dominant isolated renal magnesium loss is caused by misrouting of the Na+,K+-ATPase γ-subunit. Nat Genet. 2000;26:265–266. doi: 10.1038/81543. [DOI] [PubMed] [Google Scholar]
- Metcalf DG, Law PB, DeGrado WF. Mutagenesis data in the automated prediction of transmembrane helix dimers. Proteins. 2007;67:375–384. doi: 10.1002/prot.21265. [DOI] [PubMed] [Google Scholar]
- North B, Cristian L, Fu Stowell X, Lear JD, Saven JG, DeGrado WF. Characterization of a membrane protein folding motif, the Ser zipper, using designed peptides. J Mol Biol. 2006;359:930–939. doi: 10.1016/j.jmb.2006.04.001. [DOI] [PubMed] [Google Scholar]
- O’Shea EK, Klemm JD, Kim PS, Alber TA. X-ray structure of the GCN4 leucine zipper, a two-stranded coiled coil. Science. 1991;254:539–544. doi: 10.1126/science.1948029. [DOI] [PubMed] [Google Scholar]
- Ohsawa Y, Toko H, Katsura M, Morimoto K, Yamada H, Ichikawa Y, Murakami T, Ohkuma S, Komuro I, Sunada Y. Overexpression of P104L mutant caveolin-3 in mice develops hypertrophic cardiomyopathy with enhanced contractility in association with increased endothelial nitric oxide synthase activity. Hum Mol Genet. 2004;13:151–157. doi: 10.1093/hmg/ddh014. [DOI] [PubMed] [Google Scholar]
- Oxenoid K, Chou JJ. The structure of phospholamban pentamer reveals a channel-like architecture in membranes. Proc Natl Acad Sci USA. 2005;102:10870–10875. doi: 10.1073/pnas.0504920102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paloneva J, Manninen T, Christman G, Hovanes K, Mandelin J, Adolfsson R, Bianchin M, Bird T, Miranda R, Salmaggi A, et al. Mutations in two genes encoding different subunits of a receptor signaling complex result in an identical disease phenotype. Am J Hum Genet. 2002;71:656–662. doi: 10.1086/342259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Partridge AW, Liu S, Kim S, Bowie JU, Ginsberg MH. Transmembrane domain helix packing stabilizes integrin α IIbβ 3 in the low affinity state. J Biol Chem. 2004;280:7294–7300. doi: 10.1074/jbc.M412701200. [DOI] [PubMed] [Google Scholar]
- Pikman Y, Lee BH, Mercher T, McDowell E, Ebert BL, Gozo M, Cuker A, Wernig G, Moore S, Galinsky I, et al. MPLW515L is a novel somatic activating mutation in myelofibrosis with myeloid metaplasia. PLoS Med. 2006;3:1140–1151. doi: 10.1371/journal.pmed.0030270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pleiman CM, Chien NC, Cambier JC. Point mutations define a mIgM transmembrane region motif that determines intersubunit signal transduction in the antigen receptor. J Immunol. 1994;152:2837–2844. [PubMed] [Google Scholar]
- Qiu JX, Petersson EJ, Matthews EE, Schepartz A. Toward β-amino acid proteins: a cooperatively folded β-peptide quaternary structure. J Am Chem Soc. 2006;128:11338–11339. doi: 10.1021/ja063164+. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quintana FJ, Gerber D, Bloch I, Cohen IR, Shai Y. A structurally altered D,L-amino acid TCR α transmembrane peptide interacts with the TCR α and inhibits T-cell activation in vitro and in an animal model. Biochemistry. 2007;46:2317–2325. doi: 10.1021/bi061849g. [DOI] [PubMed] [Google Scholar]
- Ruan W, Becker V, Klingmuller U, Langosch D. The interface between self-assembling erythropoietin receptor transmembrane segments corresponds to a membrane-spanning leucine zipper. J Biol Chem. 2004a;279:3273–3279. doi: 10.1074/jbc.M309311200. [DOI] [PubMed] [Google Scholar]
- Ruan W, Lindner E, Langosch D. The interface of a membrane-spanning leucine zipper mapped by asparagine-scanning mutagenesis. Protein Sci. 2004b;13:555–559. doi: 10.1110/ps.03357404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salzer U, Chapel HM, Webster ADB, Pan-Hammarstrom Q, Schmitt-Graeff A, Schlesier M, Peter HH, Rockstroh JK, Schneider P, Schaffer AA, et al. Mutations in TNFRSF13B encoding TACI are associated with common variable immunodeficiency in humans. Nat Genet. 2005;37:820–828. doi: 10.1038/ng1600. [DOI] [PubMed] [Google Scholar]
- Schnell JR, Chou JJ. Structure and mechanism of the M2 proton channel of influenza A virus. Nature. 2008;451:591–595. doi: 10.1038/nature06531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Senes A, Gerstein M, Engelman DM. Statistical analysis of amino acid patterns in transmembrane helices: the GxxxG motif occurs frequently and in association with β-branched residues at neighboring positions. J Mol Biol. 2000;296:921–936. doi: 10.1006/jmbi.1999.3488. [DOI] [PubMed] [Google Scholar]
- Senes A, Ubarretxena-Belandia I, Engelman DM. The Cα—H…O hydrogen bond: a determinant of stability and specificity in transmembrane helix interactions. Proc Natl Acad Sci USA. 2001;98:9056–9061. doi: 10.1073/pnas.161280798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Senes A, Engel DE, DeGrado WF. Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr Opin Struct Biol. 2004;14:465–479. doi: 10.1016/j.sbi.2004.07.007. [DOI] [PubMed] [Google Scholar]
- Seubert N, Royer Y, Staerk J, Kubatzky KF, Moucadel V, Krishnakumar S, Smith SO, Constantinescu SN. Active and inactive orientations of the transmembrane and cytosolic domains of the erythropoietin receptor dimer. Mol Cell. 2003;12:1239–1250. doi: 10.1016/s1097-2765(03)00389-7. [DOI] [PubMed] [Google Scholar]
- Shy ME, Jani A, Krajewski K, Grandis M, Lewis RA, Li J, Shy RR, Balsamo J, Lilien J, Garbern JY, et al. Phenotypic clustering in MPZ mutations. Brain. 2004;127:371–384. doi: 10.1093/brain/awh048. [DOI] [PubMed] [Google Scholar]
- Simmerman HK, Kobayashi YM, Autry JM, Jones LR. A leucine zipper stabilizes the pentameric membrane domain of phospholamban and forms a coiled-coil pore structure. J Biol Chem. 1996;271:5941–5946. doi: 10.1074/jbc.271.10.5941. [DOI] [PubMed] [Google Scholar]
- Stevens TL, Blum JH, Foy SP, Matsuuchi L, DeFranco AL. A mutation of the μ transmembrane that disrupts endoplasmic reticulum retention: effects on association with accessory proteins and signal transduction. J Immunol. 1994;152:4397–4406. [PubMed] [Google Scholar]
- Stokes DL, Pomfret AJ, Rice WJ, Glaves JP, Young HS. Interactions between Ca2+-ATPase and the pentameric form of phospholamban in two-dimensional co-crystals. Biophys J. 2006;90:4213–4223. doi: 10.1529/biophysj.105.079640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stouffer AL, Acharya R, Salom D, Levine AS, Di Costanzo L, Soto CS, Tereshko V, Nanda V, Stayrook S, DeGrado WF. Structural basis for the function and inhibition of an influenza virus proton channel. Nature. 2008;451:596–599. doi: 10.1038/nature06528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sulistijo ES, MacKenzie KR. Sequence dependence of BNIP3 transmembrane domain dimerization implicates side-chain hydrogen bonding and a tandem GxxxG motif in specific helix-helix interactions. J Mol Biol. 2006;364:974–990. doi: 10.1016/j.jmb.2006.09.065. [DOI] [PubMed] [Google Scholar]
- Tatko CD, Nanda V, Lear JD, Degrado WF. Polar networks control oligomeric assembly in membranes. J Am Chem Soc. 2006;128:4170–4171. doi: 10.1021/ja055561a. [DOI] [PubMed] [Google Scholar]
- Unterreitmeier S, Fuchs A, Schaffler T, Heym RG, Frishman D, Langosch D. Phenylalanine promotes interaction of transmembrane domains via GxxxG motifs. J Mol Biol. 2007;374:705–718. doi: 10.1016/j.jmb.2007.09.056. [DOI] [PubMed] [Google Scholar]
- Vincent MC, Biancalana V, Ginisty D, Mandel JL, Calvas P. Mutational spectrum of the ED1 gene in X-linked hypohidrotic ectodermal dysplasia. Eur J Hum Genet. 2001;9:355–363. doi: 10.1038/sj.ejhg.5200635. [DOI] [PubMed] [Google Scholar]
- Walters RF, DeGrado WF. Helix-packing motifs in membrane proteins. Proc Natl Acad Sci USA. 2006;103:13658–13663. doi: 10.1073/pnas.0605878103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Zhao X, Duan W, Fu JX, Lu M, Wang G, Bai X, Ruan C. A novel mutation in the transmembrane region of glycoprotein IX associated with Bernard-Soulier syndrome. Thromb Haemost. 2004;92:606–613. doi: 10.1160/TH04-04-0240. [DOI] [PubMed] [Google Scholar]
- Wohlfahrt G. Analysis of pH-dependent elements in proteins: geometry and properties of pairs of hydrogen-bonded carboxylic acid side-chains. Proteins. 2005;58:396–406. doi: 10.1002/prot.20328. [DOI] [PubMed] [Google Scholar]
- Wu Y, Shih SC, Goto NK. Probing the structure of the Ff bacteriophage major coat protein transmembrane helix dimer by solution NMR. Biochim Biophys Acta. 2007;1768:3206–3215. doi: 10.1016/j.bbamem.2007.08.010. [DOI] [PubMed] [Google Scholar]
- Yin H, Litvinov RI, Vilaire G, Zhu H, Li W, Caputo GA, Moore DT, Lear JD, Weisel JW, DeGrado WF, et al. Activation of platelet α IIb β 3 by an exogenous peptide corresponding to the transmembrane domain of α IIb. J Biol Chem. 2006;281:36732–36741. doi: 10.1074/jbc.M605877200. [DOI] [PubMed] [Google Scholar]
- Yin H, Slusky JS, Berger BW, Walters RS, Vilaire G, Litvinov RI, Lear JD, Caputo GA, Bennett JS, DeGrado WF. Computational design of peptides that target transmembrane helices. Science. 2007;315:1817–1822. doi: 10.1126/science.1136782. [DOI] [PubMed] [Google Scholar]
- Yohannan S, Yang D, Faham S, Boulting G, Whitelegge J, Bowie JU. Proline substitutions are not easily accommodated in a membrane protein. J Mol Biol. 2004;341:1–6. doi: 10.1016/j.jmb.2004.06.025. [DOI] [PubMed] [Google Scholar]
- Zhou FX, Cocco MJ, Russ WP, Brunger AT, Engelman DM. Interhelical hydrogen bonding drives strong interactions in membrane proteins. Nat Struct Biol. 2000;7:154–160. doi: 10.1038/72430. [DOI] [PubMed] [Google Scholar]
- Zhou FX, Merianos HJ, Brunger AT, Engelman DM. Polar residues drive association of polyleucine transmembrane helices. Proc Natl Acad Sci USA. 2001;98:2250–2255. doi: 10.1073/pnas.041593698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zouzoulas A, Therien AG, Scanzano R, Deber CM, Blostein R. Modulation of Na,K-ATPase by the γ subunit: studies with transfected cells and transmembrane mimetic peptides. J Biol Chem. 2003;278:40437–40441. doi: 10.1074/jbc.M308610200. [DOI] [PubMed] [Google Scholar]