Emergence of Multidrug-Resistant Klebsiella pneumoniae Isolates Producing VIM-4 Metallo-β-Lactamase, CTX-M-15 Extended-Spectrum β-Lactamase, and CMY-4 AmpC β-Lactamase in a Tunisian University Hospital (original) (raw)
Abstract
Klebsiella pneumoniae clinical isolates resistant to carbapenems were recovered from 11 patients in the hospital of Sfax, Tunisia. The isolates were closely related as shown by pulsed-field gel electrophoresis, and they produced VIM-4 metallo-enzyme, CTX-M-15 extended-spectrum β-lactamase, and CMY-4 AmpC enzyme. The _bla_VIM-4 gene is part of a class 1 integron.
During the last decade, acquired metallo-β-lactamases (MBLs) emerged among Pseudomonas aeruginosa isolates and other gram-negative nosocomial pathogens (2, 6, 7, 14, 15, 20, 27). The most frequently acquired MBLs are the IMP and VIM types (15, 20). Three other types of acquired MBLs in P. aeruginosa isolates from Brazil (SPM-1) (28) and Germany (GIM-1) (3) and in Acinetobacter baumannii isolates from Korea (SIM-1) (13) have recently been described.
The first member of the VIM family of determinants, VIM-1, was identified from a clinical isolate of P. aeruginosa in Verona, Italy (11). Over the past few years, studies have reported the dissemination of VIM-type MBLs in Enterobacteriaceae (30), suggesting the ongoing spread of these resistance determinants among more pathogens. Outbreaks of Klebsiella pneumoniae strains producing VIM-type MBLs have been reported recently in Greece (8), France (10), and Italy (16).
We report the emergence of a multidrug-resistant K. pneumoniae isolate that produces the metallo-β-lactamase VIM-4, extended-spectrum β-lactamase (ESBL) CTX-M-15, AmpC β-lactamase CMY-4, and class A β-lactamase TEM-1 in a Tunisian university hospital.
Between May and July 2005, 20 imipenem-resistant strains of K. pneumoniae were recovered from 11 patients from different wards. The index case was a 50-year-old woman who underwent placement of an indwelling double ureteral stent for acute purulent calculous pyelonephritis and received 12 days of treatment with cefotaxime. One month later, the patient developed a stent-associated infection. Carbapenem-resistant K. pneumoniae was recovered from her urine and blood. After treatment with colimycin and imipenem over 4 weeks and removal of the ureteral stent, the patient recovered. This strain was subsequently recovered from 10 other patients (Table 1). All infections were acquired in the hospital. Ten out of the 11 patients had received some kind of surgery, implying that the K. pneumoniae isolate could have been acquired in operating theaters, but no common source was identified.
TABLE 1.
Origins of _bla_VIM-4-containing K. pneumoniae isolates and clinical characteristics of the 11 infected patients
| Patient | Age (yr)/sexa | Hospital ward(s)b | Collection date(s) (day/mo/yr) (source) | Therapy | Status (type of infection) | Outcome |
|---|---|---|---|---|---|---|
| 1 | 45/F | Internal medicine | 07/05/05 (urine), 13/05/05 (blood) | Imipenem + colimycin | Urinary infection | Recovered |
| 2 | 67/F | Internal medicine | 14/05/05 (urine) | Imipenem + colimycin | Urinary infection | Recovered |
| 3 | 36/M | ICU | 17/05/05 (blood) | Imipenem + colimycin | Bacteremia (contamination) | Death |
| 4 | 22/M | ICU, orthopedics | 14/06/05 (wound), 16/06/05 (wound) | Imipenem + colimycin | Wound infection | Death |
| 5 | 36/F | ICU | 21/06/05 (catheter), 21/06/05 (sputum), 27/06/05 (blood) | Vancomycin | Bacteremia | Death |
| 6 | 87/M | ICU | 20/06/05 (blood) | Bacteremia (contamination) | Recovered | |
| 7 | 40/M | Urology | 24/06/05 (urine) | Ciprofloxacin | Colonization | Recovered |
| 8 | 74/F | ICU | 13/07/05 (urine) | Vancomycin | Colonization | Death |
| 9 | 34/M | ICU | 13/07/05 (urine), 13/07/05 (blood), 13/07/05 (blood), 14/07/05 (catheter), 24/07/05 (blood), 24/07/05 (blood) | Imipenem + colimycin | Bacteremia | Death |
| 10 | 48/F | Neurosurgery, ICU | 13/07/05 (cerebral abscess) | Imipenem + colimycin + ciprofloxacin | Cerebral abscess | Death |
| 11 | 65/M | ICU | 13/07/05 (urine) | Colonization | Death |
Six of the 11 study patients were infected with a carbapenem-resistant isolate, and 4 of these died during their stay in the intensive care unit, with the K. pneumoniae infection being causative or contributory. The two patients with urinary tract infections were successfully treated with colimycin and imipenem.
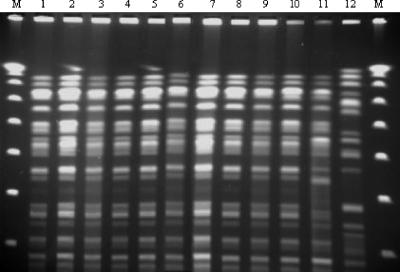
The first isolate for each patient was included in this study. Susceptibility testing using the disk diffusion method showed that all isolates were highly resistant to all β-lactams and exhibited resistance to most non-β-lactam antimicrobials tested (including aminoglycosides and ciprofloxacin), except for colistin. As all the K. pneumoniae isolates had similar antimicrobial susceptibility patterns, we investigated the clonal relationship of these strains by pulsed-field gel electrophoresis (PFGE) of SpeI-restricted genomic DNA. The PFGE results revealed that all K. pneumoniae strains isolated were genetically identical (Fig. 1) (28) and were different from the profiles obtained for VIM-1-producing K. pneumoniae strains K1, K5, and K8 from Greece used as controls (8).
FIG. 1.
PFGE fingerprints of Klebsiella pneumoniae isolates after digestion with SpeI. Lanes: M, lambda ladder (molecular size marker; Bio-Rad); 1 to 11, PFGE patterns of imipenem-resistant Klebsiella pneumoniae isolates from Sfax University Hospital; 12, PFGE pattern of Klebsiella pneumoniae K1 isolate from Greece.
All isolates were resistant to both aztreonam and imipenem. These isolates were positive by the EDTA disk synergy test, suggesting the presence of a class B enzyme (MBL), but this could not explain the high level of resistance to aztreonam. Thus, we investigated the presence of β-lactamases by PCR, using specific primers for _bla_TEM, _bla_SHV, _bla_CTX-M, _bla_CMY, _bla_ACC, _bla_VIM, and _bla_IMP (Table 2), and by sequencing. This screening showed the presence of VIM-4 class B enzyme, CTX-M-15 ESBL, CMY-4 AmpC enzyme, and TEM-1 class A β-lactamase. The _bla_SHV-1 gene that was detected probably corresponded to the chromosome-encoded enzyme.
TABLE 2.
PCR primers used in the analysis of the _bla_VIM gene and other sequences
| Product of target gene | Designation | Sequence (5′→ 3′) | Amplicon's expected size (bp) | Annealing temp (°C) | Source or reference |
|---|---|---|---|---|---|
| TEM | OT3 | ATG AGT ATT CAA CAT TTC CG | 850 | 55 | 5 |
| OT4 | CCA ATG CTT AAT CAG TGA GG | ||||
| CTX-M-1 (group M1-M3) | M13 up | GGT TAA AAA ATC ACT GCG TC | 840 | 55 | 5 |
| M13 low | TTG GTG ACG ATT TTA GCC GC | ||||
| CTX-M-2 (group M2-M5) | M25 up | ATG ATG ACT CAG AGC ATT CG | 850 | 55 | 5 |
| M25 low | TGG GTT ACG ATT TTC GCC GC | ||||
| CTX-M-9 (group M9) | M9 up | ATG GTG ACA AAG AGA GTG CA | 850 | 55 | 5 |
| M9 low | CCC TTC GGC GAT GAT TCT C | ||||
| CTX-M consensus | MA-1 | SCS ATG TGC AGY ACC AGT AA | 450 | 55 | 5 |
| MA-2 | CCG CRA TAT GRT TGG TGG TG | ||||
| CMY-2 type | CF1 | ATG ATG AAA AAA TCG TTA TGC | 1,200 | 55 | 5 |
| CF2 | TTG CAG CTT TTC AAG AAT GCG C | ||||
| ACC | ACC-U1 | GAC ACC GTT GAT GAC CTG AT | 700 | 55 | 19 |
| ACC-L1 | CAC CGA AGC CGT TAG TTG AT | ||||
| VIM | VIM-DIA/f | CAG ATT GCC GAT GGT GTT TGG | 523 | 55 | 4 |
| VIM-DIA/r | AGG TGG GCC ATT CAG CCA GA | ||||
| IMP-1 | IMP-1 Upper 5′ | CTA CCG CAG CAG AGT CTT TG | 587 | 58 | 24 |
| IMP-1 Lower 5′ | AAC CAG TTT TGC CTT ACC AT | ||||
| IMP-2 | IMP-2 Upper 5′ | GTT TTA TGT GTA TGC TTC C | 678 | 52 | 25 |
| IMP-2 Lower 5′ | AGC CTG TTC CCA TGT AC |
In view of the results of β-lactamase screening, we determined the MICs of cefotaxime, ceftazidime, cefepime, aztreonam, imipenem, and meropenem, with and without inhibitors (EDTA or clavulanic acid alone and in association), by agar dilution. Table 3 shows the MICs for the index case. The remainder of the strains were phenotypically identical. In the presence of EDTA, the MICs of imipenem decreased to 0.25 μg/ml and those of meropenem to 0.06 μg/ml, indicating the production of the metallo-β-lactamase. Moreover, the MICs of extended-spectrum cephalosporins were reduced fourfold to eightfold, whereas the MICs of aztreonam were not significantly reduced. The ESBL phenotype was thus verified by the significant reduction of the MICs of aztreonam in the presence of clavulanic acid. Only the combination of two inhibitors, EDTA and clavulanic acid, restored the activities of the broad-spectrum cephalosporins. The decrease in the MICs was more important for cefepime than for cefotaxime and ceftazidime, indicating the presence of an additional class C β-lactamase (Table 3).
TABLE 3.
MICs of various ß-lactams for the Klebsiella pneumoniae isolate recovered from patient 1
| Inhibitor(s)a | MIC (μg/ml) | |||||
|---|---|---|---|---|---|---|
| Cefotaxime | Ceftazidime | Cefepime | Aztreonam | Imipenem | Meropenem | |
| None | 256 | 256 | 128 | 128 | 32 | 2 |
| EDTA | 64 | 64 | 32 | 128 | 0.25 | 0.06 |
| CA | 64 | 128 | 8 | 8 | 32 | 2 |
| EDTA and CA | 4 | 2 | 0.06 | 8 | 0.25 | 0.06 |
To analyze the genetic support of these various β-lactamase genes, conjugational transfers were done with Escherichia coli J53-2 Rifr as the recipient and with selection on aztreonam (4 μg/ml), cefotaxime (4 μg/ml), or imipenem (2 μg/ml) and rifampin (250 μg/ml). Two different antimicrobial resistance phenotypes were obtained, the first on aztreonam suggesting the presence of an ESBL and the second on cefotaxime or imipenem suggesting the presence of the metallo-β-lactamase. Plasmid extraction showed the presence of two large plasmids (>130 kb) in the K. pneumoniae isolate (data not shown). By PCR, the smallest encoded both CMY-2-type and VIM-type enzymes in the transconjugants obtained on imipenem or cefotaxime, whereas the largest (transferred on aztreonam) encoded both CTX-M-type and TEM-type β-lactamases (data not shown).
To confirm the presence and the sequences of the three β-lactamases (VIM-4, CTX-M-15, and CMY-4), we did cloning experiments. DNA fragments obtained from genomic DNA partially digested with Sau3A were ligated into the vector pACYC184 digested with BamHI. E. coli DH10B (Invitrogen SARL, Cergy-Pontoise, France) transformants were selected on Mueller-Hinton agar supplemented with 50 μg/ml of chloramphenicol and 2 μg/ml of ceftazidime. Three different antimicrobial resistance phenotypes were obtained and were consistent with the production of MBL enzyme, ESBL, and cephalosporinase. The identification of these β-lactamase-encoding genes confirmed the presence of VIM-4, CTX-M-15, and CMY-4 enzymes. Further nucleotide sequence analysis of the MBL determinant revealed that _bla_VIM-4 was part of a class 1 integron as previously described (12, 22, 26, 29). The cassette region contained (from 5′ to 3′) _bla_VIM-4, aacA7, dhfrA1, and aadA1 genes. This structure was similar to those reported for VIM-1-producing P. aeruginosa, E. coli, and K. pneumoniae (8-11, 17).
This is the first report of MBLs in Tunisia (1). The simultaneous production of three β-lactamases (VIM-4, CMY-4, and CTX-M15) by K. pneumoniae clinical isolates is noteworthy. The coexistence of two enzymes, an MBL and a non-MBL extended-spectrum β-lactamase, in the same strain has been previously documented for Enterobacteriaceae, with both VIM-1 and a CTX-M-type β-lactamase (23), VIM-1 and SHV-5 (10), SHV-12 and VIM-4 (16), VIM-2 and IBC-1 (7), IMP-1 and CTX-M-2 (15), VIM-12 and a CMY-type cephalosporinase (21), and VIM-1 and CMY-13 (18).
The emergence of the _bla_VIM-4 gene indicates the wide circulation of MBL-encoding genes and poses challenges for the treatment of hospital infections due to gram-negative bacteria. The outbreak of imipenem-resistant K. pneumoniae occurred in our hospital over a 3-month period. From July 2005 until July 2006, only two other K. pneumoniae isolates producing MBLs were recovered from two patients. Although the _bla_VIM-positive isolates were still confined to these units and spread at a low rate in our hospital, strict infection control measures against such isolates should be implemented to prevent their further dissemination.
Nucleotide sequence accession number.
The nucleotide sequences reported in this work have been deposited in the EMBL nucleotide sequence database under accession number AM181293.
Acknowledgments
This study was done with the financial support of the Ministry of Scientific Research Technology and Competence Development of Tunisia. This work was also financed by grants from the Faculté de Médecine Pierre et Marie Curie (Saint-Antoine site), Université Paris VI, Paris, France, and from the European Community, contract LSHM-CT 2003-503335.
Footnotes
▿
Published ahead of print on 2 October 2006.
REFERENCES
- 1.Ben Redjeb, S., and I. Boutiba Ben Boubaker. 2005. L'antibio-resistance en Tunisie: données 1999-2003. Laboratoire Résistance aux Antibiotiques, LAB MDT-03, Ministère de la Recherche Scientifique, de la Technologie et du Développement des Compétences, Tunis, République Tunisienne.
- 2.Bush, K. 2001. New β-lactamases in gram-negative bacteria: diversity and impact on the selection of antimicrobial therapy. Clin. Infect. Dis. 32**:**1085-1089. [DOI] [PubMed] [Google Scholar]
- 3.Castanheira, M., M. A. Toleman, R. N. Jones, F. J. Schmidt, and T. R. Walsh. 2004. Molecular characterization of a β-lactamase gene, _bla_GIM-1, encoding a new subclass of metallo-β-lactamase. Antimicrob. Agents Chemother. 48**:**4654-4661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Docquier, J. D., F. Luzzaro, G. Amicosante, A. Toniolo, and G. M. Rossolini. 2001. Multidrug-resistant Pseudomonas aeruginosa producing PER-1 extended-spectrum serine-beta-lactamase and VIM-2 metallo-beta-lactamase. Emerg. Infect. Dis. 7**:**910-911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Eckert, C., V. Gautier, and G. Arlet. 2006. DNA sequence analysis of the genetic environment of various _bla_CTX-M genes. J. Antimicrob. Chemother. 57**:**14-23. [DOI] [PubMed] [Google Scholar]
- 6.Fiett, J., A. Baraniak, A. Mréwka, M. Fleischer, Z. Drulis-Kawa, £. Naumiuk, A. Samet, W. Hryniewicz, and M. Gniadkowski. 2006. Molecular epidemiology of acquired-metallo-β-lactamase-producing bacteria in Poland. Antimicrob. Agents Chemother. 50**:**880-886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Galani, I., M. Souli, Z. Chryssouli, D. Katsala, and H. Giamarellou. 2004. First identification of an Escherichia coli clinical isolate producing both metallo-β-lactamase VIM-2 and extended-spectrum β-lactamase IBC-1. Clin. Microbiol. Infect. 10**:**757-760. [DOI] [PubMed] [Google Scholar]
- 8.Giakkoupi, P., A. Xanthaki, M. Kanelopoulou, A. Vlahaki, V. Miriagou, S. Kontou, E. Papafraggas, H. Malamou-Lada, L. S. Tzouvelekis, N. J. Legakis, and A. C. Vatopoulos. 2003. VIM-1 metallo-β-lactamase-producing Klebsiella pneumoniae strains in Greek hospitals. J. Clin. Microbiol. 41**:**3893-3896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ikonomidis, A., D. Tokatlidou, I. Kristo, D. Sofianou, A. Tsakris, P. Mantzana, S. Pournaras, and A. N. Maniatis. 2005. Outbreaks in distinct regions due to a single Klebsiella pneumoniae clone carrying a _bla_VIM-1 metallo-β-lactamase gene. J. Clin. Microbiol. 43**:**5344-5347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kassis-Chikhani, N., D. Decre, V. Gautier, B. Burghoffer, F. Saliba, D. Mathieu, D. Samuel, D. Castaing, J. C. Petit, E. Dussaix, and G. Arlet. 2006. First outbreak of multidrug-resistant Klebsiella pneumoniae carrying _bla_VIM-1 and _bla_SHV-5 in a French university hospital. J. Antimicrob. Chemother. 57**:**142-145. [DOI] [PubMed] [Google Scholar]
- 11.Lauretti, L., M. L. Riccio, A. Mazzariol, G. Cornaglia, G. Amicosante, R. Fontana, and G. M. Rossolini. 1999. Cloning and characterization of _bla_VIM, a new integron-borne metallo-β-lactamase gene from a Pseudomonas aeruginosa clinical isolate. Antimicrob. Agents Chemother. 43**:**1584-1590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee, K., J. B. Lim, J. H. Yum, D. Yong, Y. Chong, J. M. Kim, and D. M. Livermore. 2002. _bla_VIM-2 cassette-containing novel integrons in metallo-β-lactamase-producing Pseudomonas aeruginosa and Pseudomonas putida isolates disseminated in a Korean hospital. Antimicrob. Agents Chemother. 46**:**1053-1058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lee, K., J. H. Yum, D. Yong, H. M. Lee, H. D. Kim, J.-D. Docquier, G. M. Rossolini, and Y. Chong. 2005. Novel acquired metallo-β-lactamase gene, _bla_SIM-1, in a class 1 integron from Acinetobacter baumannii clinical isolates from Korea. Antimicrob. Agents Chemother. 49**:**4485-4491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lincopan, N., J. A. McCulloch, C. Reinert, V. C. Cassettari, A. C. Gales, and E. M. Mamizuka. 2005. First isolation of metallo-β-lactamase-producing multiresistant Klebsiella pneumoniae from a patient in Brazil. J. Clin. Microbiol. 43**:**516-519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Livermore, D. M., and N. Woodford. 2000. Carbapenemases: a problem in waiting? Curr. Opin. Microbiol. 3**:**489-495. [DOI] [PubMed] [Google Scholar]
- 16.Luzzaro, F., J.-D. Docquier, C. Colinon, A. Endimiani, G. Lombardi, G. Amicosante, G. M. Rossolini, and A. Toniolo. 2004. Emergence in Klebsiella pneumoniae and Enterobacter cloacae clinical isolates of the VIM-4 metallo-β-lactamase encoded by a conjugative plasmid. Antimicrob. Agents Chemother. 48**:**648-650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Miriagou, V., E. Tzelepi, D. Gianneli, and L. S. Tzouvelekis. 2003. Escherichia coli with a self-transferable, multiresistant plasmid coding for metallo-β-lactamase VIM-1. Antimicrob. Agents Chemother. 47**:**395-397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Miriagou, V., L. S. Tzouvelekis, L. Villa, E. Lebessi, A. C. Vatopoulos, A. Carattoli, and E. Tzelepi. 2004. CMY-13, a novel inducible cephalosporinase encoded by an Escherichia coli plasmid. Antimicrob. Agents Chemother. 48**:**3172-3174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nadjar, D., R. Labia, C. Cerceau, C. Bizet, A. Philippon, and G. Arlet. 2000. Outbreak of Klebsiella pneumoniae producing transferable AmpC-type beta-lactamase (ACC-1) originating from Hafnia alvei. FEMS Microbiol. Lett. 187**:**35-40. [DOI] [PubMed] [Google Scholar]
- 20.Nordmann, P., and L. Poirel. 2002. Emerging carbapenemases in gram-negative aerobes. Clin. Microbiol. Infect. 8**:**321-331. [DOI] [PubMed] [Google Scholar]
- 21.Pournaras, S., A. Ikonomidis, L. S. Tzouvelekis, D. Tokatlidou, N. Spanakis, A. N. Maniatis, N. J. Legakis, and A. Tsakris. 2005. VIM-12, a novel plasmid-mediated metallo-β-lactamase from Klebsiella pneumoniae that resembles a VIM-1/VIM-2 hybrid. Antimicrob. Agents Chemother. 49**:**5153-5156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Riccio, M. L., L. Pallecchi, J.-D. Docquier, S. Cresti, M. R. Catania, L. Pagani, C. Lagatolla, G. Cornaglia, R. Fontana, and G. M. Rossolini. 2005. Clonal relatedness and conserved integron structures in epidemiologically unrelated Pseudomonas aeruginosa strains producing the VIM-1 metallo-β-lactamase from different Italian hospitals. Antimicrob. Agents Chemother. 49**:**104-110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Scoulica, E. V., I. K. Neonakis, A. I. Gikas, and Y. J. Tselentis. 2004. Spread of _bla_VIM-1 producing E. coli in a university hospital in Greece. Genetic analysis of the integron carrying the _bla_VIM-1 metallo-beta-lactamase gene. Diagn. Microbiol. Infect. Dis. 48**:**167-172. [DOI] [PubMed] [Google Scholar]
- 24.Senda, K., Y. Arakawa, S. Ichiyama, K. Nakashima, H. Ito, S. Ohsuka, K. Shimokata, N. Kato, and M. Ohta. 1996. PCR detection of metallo-β-lactamase gene (_bla_IMP) in gram-negative rods resistant to broad-spectrum β-lactams. J. Clin. Microbiol. 34**:**2909-2913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Shibata, N., Y. Doi, K. Yamane, T. Yagi, H. Kurokawa, K. Shibayama, H. Kato, K. Kai, and Y. Arakawa. 2003. PCR typing of genetic determinants for metallo-β-lactamases and integrases carried by gram-negative bacteria isolated in Japan, with focus on the class 3 integron. J. Clin. Microbiol. 41**:**5407-5413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tenover, F. C., R. D. Arbeit, R. V. Goering, P. A. Mickelsen, B. E. Murray, D. H. Persing, and B. Swaminathan. 1995. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J. Clin. Microbiol. 33**:**2233-2239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Toleman, M. A., D. Biedenbach, D. M. Bennett, R. N. Jones, and T. R. Walsh. 2005. Italian metallo-beta-lactamases: a national problem? Report from the SENTRY Antimicrobial Surveillance Programme. J. Antimicrob. Chemother. 55**:**61-70. [DOI] [PubMed] [Google Scholar]
- 28.Toleman, M. A., A. M. Simm, T. A. Murphy, A. C. Gales, D. J. Biedenbach, R. N. Jones, and T. R. Walsh. 2002. Molecular characterization of SPM-1, a novel metallo-beta-lactamase isolated in Latin America: report from the SENTRY antimicrobial surveillance programme. J. Antimicrob. Chemother. 50**:**673-679. [DOI] [PubMed] [Google Scholar]
- 29.Tórtola, M. T., S. Lavilla, E. Miró, J. J. González, N. Larrosa, M. Sabaté, F. Navarro, and G. Prats. 2005. First detection of a carbapenem-hydrolyzing metalloenzyme in two Enterobacteriaceae isolates in Spain. Antimicrob. Agents Chemother. 49**:**3492-3494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Walsh, T. R., M. A. Toleman, L. Poirel, and P. Nordmann. 2005. Metallo-beta-lactamases: the quiet before the storm? Clin. Microbiol. Rev. 18**:**306-325. [DOI] [PMC free article] [PubMed] [Google Scholar]