Mps1p regulates meiotic spindle pole body duplication in addition to having novel roles during sporulation - PubMed (original) (raw)
Mps1p regulates meiotic spindle pole body duplication in addition to having novel roles during sporulation
P D Straight et al. Mol Biol Cell. 2000 Oct.
Free PMC article
Abstract
Sporulation in yeast requires that a modified form of chromosome segregation be coupled to the development of a specialized cell type, a process akin to gametogenesis. Mps1p is a dual-specificity protein kinase essential for spindle pole body (SPB) duplication and required for the spindle assembly checkpoint in mitotically dividing cells. Four conditional mutant alleles of MPS1 disrupt sporulation, producing two distinct phenotypic classes. Class I alleles of mps1 prevent SPB duplication at the restrictive temperature without affecting premeiotic DNA synthesis and recombination. Class II MPS1 alleles progress through both meiotic divisions in 30-50% of the population, but the asci are incapable of forming mature spores. Although mutations in many other genes block spore wall formation, the cells produce viable haploid progeny, whereas mps1 class II spores are unable to germinate. We have used fluorescently marked chromosomes to demonstrate that mps1 mutant cells have a dramatically increased frequency of chromosome missegregation, suggesting that loss of viability is due to a defect in spindle function. Overall, our cytological data suggest that MPS1 is required for meiotic SPB duplication, chromosome segregation, and spore wall formation.
Figures
Figure 1
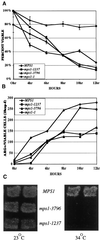
Return-to-growth assays were done to determine viability (A) and commitment to recombination (B) with the use of wild-type (YUMY3E3), mps1-1 (YUMY131 × YUMY1D1),mps1-1237 (YUMY3I1), and mps1-3796 (YUMY3I8) homozygous diploid mutant strains. Strains were synchronously sporulated, and aliquots of cells were plated to SC or SC-arginine medium at the times indicated. Viability was measured as the number of CFU at each time point as a percentage of CFU at 0 h. Commitment to recombinationwas determined with the use of_arg4_ heteroallelic diploids and plating to SC-arginine medium. Recombinant frequency is expressed as Arg+ colonies per CFU at each time point according to the method described by Esposito and Esposito (1974). Recombination assays were performed three times with triplicate samples. One representative example is shown. (C) Viability of cells sporulated at the restrictive temperature was assessed with the use of Can/Cyh resistance generated by haploidization of two recessive drug-resistance markers. Wild-type (TM002),mps1-3796 (TM019), and mps1-1237 (TM027) strains were sporulated at room temperature and 34°C. Patches were replica plated to medium containing cycloheximide and
l
-canavanine (see MATERIALS AND METHODS) and incubated at room temperature for 2–5 d to assay viable haploid cell production caused by haploidization of both can1-100 and_cyh_r markers.
Figure 2
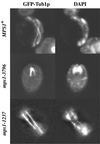
Cytological analysis of mps1 mutants during meiosis at the restrictive temperature. GFP–Tub1p and DAPI staining of DNA demonstrate allele-dependent phenotypes. Cultures were sporulated at 33°C for 8 h (MPS1, YUMY4B1;mps1-1237, YUMY4C6) or 12 h (mps1-3796, YUMY4C2), fixed with 3.7% formaldehyde for up to 1 h, and stained with 1 μg/ml DAPI for 5 min. Wild-type cells proceed through two rounds of spindle formation and chromosome segregation, as seen by GFP and DAPI, respectively. The_mps1-3796_ strain fails to form spindles over the course of meiosis and remains mononucleate. However, ∼30% of cells harboring the mps1-1237 allele successfully complete spindle formation and carry out chromosome segregation.
Figure 3
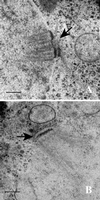
Thin-section electron microscopy of_MPS1_ (YUMY4F4) and mps1-3796 (YUMY070) strains during meiosis at the restrictive temperature reveals a SPB duplication defect in the mps1 mutant strain. (A)MPS1 strain after 6 h at 33°C in sporulation medium. Prophase meiotic nuclei have duplicated side-by-side SPBs (arrow) in the nuclear envelope. (B) mps1-3796 strain after 12 h at 33°C. Spindle pole bodies fail to duplicate during meiosis in both the mps1-3796 and mps1-1 (YUMY069) strains, leading to the class I phenotype. Bars, 0.2 μm.
Figure 4
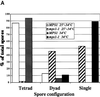
Timed temperature shift of an_mps1-1_ strain produces two-spored asci. Wild-type (YUMY4B1) and mps1-1 (YUMY4C9) strains were sporulated at 34 or 25°C for 6 h and then shifted to 34°C. Spore morphology was counted for 200 cells of each culture after 24 h of sporulation. White bars, wild-type, 25–34°C; striped bars,mps1-1, 25–34°C; gray bars, wild-type, 34°C; black bars, mps1-1, 34°C. The chart shows the percentages of cell morphology within the spore-forming population of the culture; n = 200 for each culture. Sporulation efficiency (two or more spores per ascus) was as follows: wild-type, 25–34°C = 91%;mps1-1, 25–34°C = 29.5%; wild- type, 34°C = 78%; mps1-1, 34°C = 2%.
Figure 5
DAPI staining and electron microscopy of wild-type and mps1 mutant strains reveals a spore wall defect. Cells were fixed for 15 min with 3.7% formaldehyde and stained with 1 μg/ml DAPI for <5 min (see MATERIALS AND METHODS). DAPI permeation into the interior of the spore is retarded by mature spore walls in wild-type MPS1 (YUMY4B1) (A). Lack of complete spore wall formation in the mps1 mutant strains allows DAPI to readily stain DNA within individual spores of mps1-1237 (YUMY4C6) (B), mps1-412 (YUMY4B5) (C),mps1-1 (YUMY4C9) (D), and mps1-3796 (YUMY4C2) (E). Note that in D and E, meiotic chromosome segregation fails as expected, and the cells build defective spore walls. Also note the fragmentation of DAPI-staining material within the mutant spores. Thin-section electron microscopy of MPS1 (YUMY4F4) (F),mps1-1237 (YUMY071) (G), and mps1-3796 (YUMY070) (H) strains displays the heterogeneous nature of the spore wall defect caused by mps1 mutation. (F) Wild-type spores have a four-layered structure that is relatively uniform in thickness. Mature spores are tightly associated in a tetrahedral array within a condensed ascus. Darkly staining material defines the outer, dityrosine-containing layer of the spore wall, and electron-dense material is seen adjoining individual spores (arrow). (G) Two to four spores are produced in mps1-1237 strains, and individual spores lack the organization of the spore wall seen in wild-type. Spore wall thickness is more variable than in wild- type, with component layers of spore wall material not organized as in wild-type. The electron-dense outer layer of the mutant spores is often missing, and the asci tend not to condense around tightly adherent spores. Note that the very electron-dense material connecting the MPS1 spores is disrupted in the mps1-1237 ascus (arrow). Up to 15% of cells containing the mps1-3796 allele eventually form an aberrant single spore body (H, arrowhead). The spore produced in these cells reveals similar structural defects as in a_mps1-1237_ strain. Bars, 0.7 μm
Figure 6
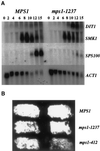
Northern analysis of sporulation-specific gene expression and dityrosine fluorescence in mps1 mutant strains. (A) MPS1 (YUMY125 × YUMY126) and_mps1-1237_ (YUMY119 × YUMY120) strains were synchronously sporulated, and aliquots were removed for RNA isolation. Filters were probed for expression of three midlate sporulation-specific genes, SMK1, SWM1, and DIT1, the late spore wall–specific gene_SPS100_, and the ACT1 gene as a loading control. Expression of SMK1 and SWM1 (our unpublished results) is unaffected in the mps1-1237 strain compared with wild type. The expression of the_SPS100_ gene is severely diminished in the mutant strains, whereas DIT1 expression is enhanced above wild-type levels. (B) Dityrosine fluorescence is detected in both_mps1_ mutant strains producing two- to four-spored asci. Patches of MPS1 (YUMY3E3), mps1-1237 (YUMY3I1), and mps1-412 (YUMY 1980) were sporulated at the restrictive temperature and then treated to activate dityrosine fluorescence. The presence of fluorescent material in wild-type and mps1 mutant strains indicates that insoluble fluorescent dityrosine is produced efficiently in both strain types.
Figure 7
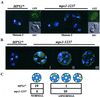
Loss of chromosome segregation fidelity in the class II mps1 mutant strains. Chromosome segregation was monitored in mps1-1237 and wild-type strains with the use of LacO_pr-marked centromere III (LEU2) visualized with a GFP–LacI repressor protein fusion (Straight_et al., 1996). (A) Chromosome segregation during meiosis I was monitored with the use of marked homologous chromosomes (four chromatids). Binucleate cells in a sporulating population were examined for segregation of paired homologues to opposite poles. Left panels, wild-type binucleate (GFP/DAPI) and four-spored (GFP/DIC) asci; right panels, examples of meiosis I missegregation in_mps1-1237_ binucleates (GFP/DAPI), which occurred in up to 80% of cells that entered meiosis I in a given experiment, and a four-spored ascus (GFP/DIC), showing no meiosis I or meiosis II segregation. (B and C) A single copy of chromosome III was marked to follow meiosis II (sister chromatid separation) specifically. Wild-type_MPS1_ strains show four DAPI-staining spore nuclei, two of which contain copies of the marked chromosome III (left panel). Right panels, mps1-1237 cells display a variety of aberrant segregation patterns. Three examples are shown. (I) Both marked centromeres reside in one of four detectable spore bodies. DAPI staining is seen within and outside of spore bodies. (II) A single GFP signal is associated with one of four DAPI-staining foci, suggesting that sister chromatids remain paired and unresolvable. (III) Many instances of DAPI staining outside of nascent spore bodies can be seen in mps1 cells. In some cases, one of the marked chromatids is found outside any of the four spores. To quantitate missegregation in these cells, we restricted our analysis to the few mutant cells with a single pair of marked chromatids and at least four visible DAPI foci. This restriction allowed us to unambiguously assign normal versus abnormal segregation, but it also resulted in a small number of scoreable cells due to the percentage of segregation in the population and weakened expression of GFP in sporulating (autofluorescent) cells. Nonetheless, the difference between_MPS1_ and mps1-1237 is statistically significant (C; p < 0.005 by χ2 analysis).
Similar articles
- The yeast protein kinase Mps1p is required for assembly of the integral spindle pole body component Spc42p.
Castillo AR, Meehl JB, Morgan G, Schutz-Geschwender A, Winey M. Castillo AR, et al. J Cell Biol. 2002 Feb 4;156(3):453-65. doi: 10.1083/jcb.200111025. Epub 2002 Feb 4. J Cell Biol. 2002. PMID: 11827982 Free PMC article. - New alleles of the yeast MPS1 gene reveal multiple requirements in spindle pole body duplication.
Schutz AR, Winey M. Schutz AR, et al. Mol Biol Cell. 1998 Apr;9(4):759-74. doi: 10.1091/mbc.9.4.759. Mol Biol Cell. 1998. PMID: 9529376 Free PMC article. - The Saccharomyces cerevisiae spindle pole body duplication gene MPS1 is part of a mitotic checkpoint.
Weiss E, Winey M. Weiss E, et al. J Cell Biol. 1996 Jan;132(1-2):111-23. doi: 10.1083/jcb.132.1.111. J Cell Biol. 1996. PMID: 8567717 Free PMC article. - Spindle pole body duplication: a model for centrosome duplication?
Adams IR, Kilmartin JV. Adams IR, et al. Trends Cell Biol. 2000 Aug;10(8):329-35. doi: 10.1016/s0962-8924(00)01798-0. Trends Cell Biol. 2000. PMID: 10884685 Review. - Genetic and biochemical approaches to spindle function and chromosome segregation in eukaryotic microorganisms.
Kilmartin JV. Kilmartin JV. Curr Opin Cell Biol. 1994 Feb;6(1):50-4. doi: 10.1016/0955-0674(94)90115-5. Curr Opin Cell Biol. 1994. PMID: 8167025 Review.
Cited by
- Mitotic Kinases and p53 Signaling.
Ha GH, Breuer EK. Ha GH, et al. Biochem Res Int. 2012;2012:195903. doi: 10.1155/2012/195903. Epub 2012 Jul 19. Biochem Res Int. 2012. PMID: 22852086 Free PMC article. - Mip1 associates with both the Mps1 kinase and actin, and is required for cell cortex stability and anaphase spindle positioning.
Mattison CP, Stumpff J, Wordeman L, Winey M. Mattison CP, et al. Cell Cycle. 2011 Mar 1;10(5):783-93. doi: 10.4161/cc.10.5.14955. Epub 2011 Mar 1. Cell Cycle. 2011. PMID: 21325884 Free PMC article. - Degradation of the human mitotic checkpoint kinase Mps1 is cell cycle-regulated by APC-cCdc20 and APC-cCdh1 ubiquitin ligases.
Cui Y, Cheng X, Zhang C, Zhang Y, Li S, Wang C, Guadagno TM. Cui Y, et al. J Biol Chem. 2010 Oct 22;285(43):32988-32998. doi: 10.1074/jbc.M110.140905. Epub 2010 Aug 20. J Biol Chem. 2010. PMID: 20729194 Free PMC article. - Role of E2Fs and mitotic regulators controlled by E2Fs in the epithelial to mesenchymal transition.
Jusino S, Saavedra HI. Jusino S, et al. Exp Biol Med (Maywood). 2019 Nov;244(16):1419-1429. doi: 10.1177/1535370219881360. Epub 2019 Oct 1. Exp Biol Med (Maywood). 2019. PMID: 31575294 Free PMC article. Review. - Ady3p links spindle pole body function to spore wall synthesis in Saccharomyces cerevisiae.
Nickas ME, Neiman AM. Nickas ME, et al. Genetics. 2002 Apr;160(4):1439-50. doi: 10.1093/genetics/160.4.1439. Genetics. 2002. PMID: 11973299 Free PMC article.
References
- Alani E, Padmore R, Kleckner N. Analysis of wild-type and rad50 mutants of yeast suggests an intimate relationship between meiotic chromosome synapsis and recombination. Cell. 1990;61:419–436. - PubMed
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K. Current Protocols in Molecular Biology. New York: John Wiley & Sons; 1997.
- Briza P, Winkler G, Kalchhauser H, Breitenbach M. Dityrosine is a prominent component of the yeast ascospore wall. J Biol Chem. 1986;261:4288–4294. - PubMed
- Byers B. Multiple roles of the spindle pole bodies in the life cycle of Saccharomyces cerevisiae. In: von Wettstein D, Stenderup A, Kielland-Brandt M, Friis J, editors. Molecular Genetics in Yeast: Alfred Benzon Symposia. Vol. 16. Copenhagen: Munksgaard; 1981. pp. 119–133.
- Chu S, DeRisi J, Eisen M, Mulholland J, Botstein D, Brown PO, Herskowitz I. The transcriptional program of sporulation in budding yeast. Science. 1998;282:699–705. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases