A novel approach for the identification of protein-protein interaction with integral membrane proteins - PubMed (original) (raw)
A novel approach for the identification of protein-protein interaction with integral membrane proteins
M Hubsman et al. Nucleic Acids Res. 2001.
Abstract
Protein-protein interaction plays a major role in all biological processes. The currently available genetic methods such as the two-hybrid system and the protein recruitment system are relatively limited in their ability to identify interactions with integral membrane proteins. Here we describe the development of a reverse Ras recruitment system (reverse RRS), in which the bait used encodes a membrane protein. The bait is expressed in its natural environment, the membrane, whereas the protein partner (the prey) is fused to a cytoplasmic Ras mutant. Protein-protein interaction between the proteins encoded by the prey and the bait results in Ras membrane translocation and activation of a viability pathway in yeast. We devised the expression of the bait and prey proteins under the control of dual distinct inducible promoters, thus enabling a rapid selection of transformants in which growth is attributed solely to specific protein-protein interaction. The reverse RRS approach greatly extends the usefulness of the protein recruitment systems and the use of integral membrane proteins as baits. The system serves as an attractive approach to explore novel protein-protein interactions with high specificity and selectivity, where other methods fail.
Figures
Figure 1
Schematic diagram describing the reverse RRS approach. The membrane protein of interest is expressed in a Cdc25-2 yeast strain with no additional sequences. The bait is located at the membrane thus preserving its functional conformation. (A) The bait protein is not expected to induce cell growth of Cdc25-2 yeast cells at the restrictive temperature. (B) A known protein partner for the bait or a cDNA library is expressed as a membrane fusion with the cytoplasmic oncogenic Ras protein. Unless the cDNA encodes for a membrane protein or is associated with a membrane component it is not expected to result in Ras translocation to the membrane and cell growth at the restrictive temperature. (C) Upon protein–protein interaction between the membrane bait and the protein prey fused to Ras, Ras is translocated to the plasma membrane which allows cell growth at the restrictive temperature.
Figure 2
The regulation of the Gal1 and Met promoters. (A) Western blot analysis with anti-Myc antibodies. The regulation of the ADH (lanes 1–4), Met (lanes 5–7) and Gal1 (lanes 8–10) promoters was examined in three distinct media. The expression of myc-epitope Ras protein (myc–Ras; lanes 2–10) or Ras (lane 1) was detected with anti-Myc antibodies. Whole cell extract was derived from yeast transformants expressing the myc–Ras protein grown on either glucose (Glu)- or galactose (Gal)-containing medium in the presence or absence of methionine (GalΔM). The migration of myc–Ras and protein size markers are indicated. (B) Methionine-dependent growth of transformants expressing protein interacting pairs. Cdc25-2 transformants expressing the indicated protein pairs under the indicated promoter were grown on glucose medium at 24°C. The plate was replica plated onto galactose-containing plates in which methionine was either included or excluded from the medium. The plates were incubated at 36°C. Transformants expressing protein pairs under the control of Met and Gal1 promoters, respectively, are able to grow only on plates lacking methionine.
Figure 3
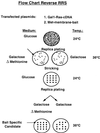
Flow chart for the reverse RRS. Cdc25-2 cells are cotransfected with the membrane bait and a cDNA library fused to Ras. The expression of the bait and Ras prey proteins are designed under the control of the Met and Gal1 promoters, respectively. Transformants are selected on glucose minimal plates lacking leucine and uracil at 24°C. After 5–7 days, plates are replica plated onto galactose-containing plates either containing or lacking methionine and incubated at 36°C. Transformants that exhibit efficient growth on the plates lacking methionine and no growth on the plate containing methionine are selected on a glucose plate and grown at 24°C. The selected transformants are retested for their methionine-dependent growth. Those transformants that pass this secondary methionine-dependent test are considered candidates and are further pursued. DNA is extracted from candidates and the library plasmid is identified by a restriction digest. The DNA is used to retransform Cdc25-2 cells with either the specific bait or non-specific bait.
Figure 4
Secondary methionine dependency test for the ChpAc.ΔN interacting candidates. Cdc25-2 cells were cotransfected with Met expression plasmid encoding for myc–ChpAc.ΔN protein bait and Yes expression plasmid encoding for oncogenic cytoplasmic Ras protein fused to rat pituitary cDNA library. 200 000 transformants were plated on 20 glucose plates lacking leucine and uracil and incubated at 24°C. Following the first replica plating ∼5% of the transformants exhibited efficient growth on galactose-containing medium at the restrictive temperature. Only 30 clones showed preferential growth on the galactose-containing plate lacking methionine. Subsequently, those transformants (only 20 are shown) were subjected to a secondary methionine test which resulted in the identification of two clones (indicated by *) that did not grow on the galactose plate containing methionine (right panel), but grew efficiently in the absence of methionine (left panel).
Figure 5
Specificity test for the library plasmids extracted from candidate clones #1 and #2. DNA plasmids isolated from candidate clones #1 and #2 were subjected to restriction digest and the library plasmid was identified. The plasmid was used to cotransfect Cdc25-2 cells with the Met expression vector encoding the original bait [i.e. ChpAc. deleted in its N-terminal domain (ChpAc.ΔN)], the Met expression vector (Met) and the Met expression vector encoding for full-length myristoylated Chp activated (Met–M–ChpAc.). Transformants were selected and used to replica plate onto appropriate plates containing galactose and lacking methionine incubated at 36°C.
Similar articles
- Adaptation of the Ras-recruitment system to the analysis of interactions between membrane-associated proteins.
Köhler F, Müller KM. Köhler F, et al. Nucleic Acids Res. 2003 Mar 15;31(6):e28. doi: 10.1093/nar/gng028. Nucleic Acids Res. 2003. PMID: 12626727 Free PMC article. - Analysis and identification of protein-protein interactions using protein recruitment systems.
Aronheim A, Karin M. Aronheim A, et al. Methods Enzymol. 2000;328:47-59. doi: 10.1016/s0076-6879(00)28389-4. Methods Enzymol. 2000. PMID: 11075337 No abstract available. - The Ras Recruitment System (RRS) for the Identification and Characterization of Protein-Protein Interactions.
Aronheim A. Aronheim A. Methods Mol Biol. 2018;1794:61-73. doi: 10.1007/978-1-4939-7871-7_5. Methods Mol Biol. 2018. PMID: 29855951 - Coregulation of starch degradation and dimorphism in the yeast Saccharomyces cerevisiae.
Vivier MA, Lambrechts MG, Pretorius IS. Vivier MA, et al. Crit Rev Biochem Mol Biol. 1997;32(5):405-35. doi: 10.3109/10409239709082675. Crit Rev Biochem Mol Biol. 1997. PMID: 9383611 Review. - [Ras proteins in Saccharomyces cerevisiae, their partners and their activation].
Jacquet M. Jacquet M. C R Seances Soc Biol Fil. 1997;191(2):221-35. C R Seances Soc Biol Fil. 1997. PMID: 9255349 Review. French.
Cited by
- RalA interacts with ZONAB in a cell density-dependent manner and regulates its transcriptional activity.
Frankel P, Aronheim A, Kavanagh E, Balda MS, Matter K, Bunney TD, Marshall CJ. Frankel P, et al. EMBO J. 2005 Jan 12;24(1):54-62. doi: 10.1038/sj.emboj.7600497. Epub 2004 Dec 9. EMBO J. 2005. PMID: 15592429 Free PMC article. - Functional characterization of SAMD9, a protein deficient in normophosphatemic familial tumoral calcinosis.
Hershkovitz D, Gross Y, Nahum S, Yehezkel S, Sarig O, Uitto J, Sprecher E. Hershkovitz D, et al. J Invest Dermatol. 2011 Mar;131(3):662-9. doi: 10.1038/jid.2010.387. Epub 2010 Dec 16. J Invest Dermatol. 2011. PMID: 21160498 Free PMC article. - The yeast split-ubiquitin membrane protein two-hybrid screen identifies BAP31 as a regulator of the turnover of endoplasmic reticulum-associated protein tyrosine phosphatase-like B.
Wang B, Pelletier J, Massaad MJ, Herscovics A, Shore GC. Wang B, et al. Mol Cell Biol. 2004 Apr;24(7):2767-78. doi: 10.1128/MCB.24.7.2767-2778.2004. Mol Cell Biol. 2004. PMID: 15024066 Free PMC article. - Examining protein protein interactions using endogenously tagged yeast arrays: the cross-and-capture system.
Suter B, Fetchko MJ, Imhof R, Graham CI, Stoffel-Studer I, Zbinden C, Raghavan M, Lopez L, Beneti L, Hort J, Fillingham J, Greenblatt JF, Giaever G, Nislow C, Stagljar I. Suter B, et al. Genome Res. 2007 Dec;17(12):1774-82. doi: 10.1101/gr.6667007. Epub 2007 Nov 7. Genome Res. 2007. PMID: 17989249 Free PMC article. - The Sos-recruitment system as a tool to analyze cellular localization of plant proteins: membrane localization of Arabidopsis thaliana PEPINO/PASTICCINO2.
Schönhofer-Merl S, Torres-Ruiz RA. Schönhofer-Merl S, et al. Mol Genet Genomics. 2010 May;283(5):439-49. doi: 10.1007/s00438-010-0528-5. Epub 2010 Mar 19. Mol Genet Genomics. 2010. PMID: 20300944
References
- Mendelsohn A.R. and Brent,R. (1999) Protein interaction methods – toward an endgame. Science, 284, 1948–1950. - PubMed
- Blackstock W.P. and Weir,M.P. (1999) Proteomics: quantitative and physical mapping of cellular proteins. Trends Biotechnol., 17, 121–127. - PubMed
- Dunham I., Shimizu,N., Roe,B.A., Chissoe,S., Hunt,A.R., Collins,J.E., Bruskiewich,R., Beare,D.M., Clamp,M., Smink,L.J. et al. (1999) The DNA sequence of human chromosome 22. Nature, 402, 489–495. - PubMed
- Goffeau A., Barrell,B.G., Bussey,H., Davis,R.W., Dujon,B., Feldmann,H., Galibert,F., Hoheisel,J.D., Jacq,C., Johnston,M. et al. (1996) Life with 6000 genes. Science, 274, 563–567. - PubMed
- Uetz P., Giot,L., Cagney,G., Mansfield,T.A., Judson,R.S., Knight,J.R., Lockshon,D., Narayan,V., Srinivasan,M., Pochart,P. et al. (2000) A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature, 403, 623–627. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases