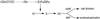Control of proliferation, endoreduplication and differentiation by the Arabidopsis E2Fa-DPa transcription factor - PubMed (original) (raw)
Control of proliferation, endoreduplication and differentiation by the Arabidopsis E2Fa-DPa transcription factor
Lieven De Veylder et al. EMBO J. 2002.
Abstract
New plant cells arise at the meristems, where they divide a few times before they leave the cell-cycle program and start to differentiate. Here we show that the E2Fa-DPa transcription factor of Arabidopsis thaliana is a key regulator determining the proliferative status of plant cells. Ectopic expression of E2Fa induced sustained cell proliferation in normally differentiated cotyledon and hypocotyl cells. The phenotype was enhanced strongly by the co-expression of E2Fa with its dimerization partner, DPa. In endoreduplicating cells, E2Fa--DPa also caused extra DNA replication that was correlated with transcriptional induction of S phase genes. Because E2Fa--DPa transgenic plants arrested early in development, we argue that controlled exit of the cell cycle is a prerequisite for normal plant development.
Figures
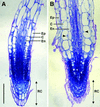
Fig. 1. In situ localization of E2Fa and DPa mRNA. Hybridization signals are seen as red dots. (A and B) Expression of E2Fa and DPa in the shoot apex of 2-month-old Arabidopsis plants, respectively. (C and D) E2Fa and DPa expression in the longitudinal section through a root meristem of Arabidopsis, respectively. (E and F) E2Fa expression in a longitudinal section through a hypocotyl of 5-day-old light- and dark-grown seedlings of Arabidopsis, respectively. C, cortex; Ep, epidermis; EZ, elongation zone; L, maturing leaf; LP, leaf primordia; Me, mesophyll; P, pith; RM, root meristem; S, stipule; SAM, shoot apical meristem; VT, vascular tissue. Bars = 100 µm (A, B and F) and 50 µm (C–E).
Fig. 2. Molecular analysis of _E2Fa_- and _DPa_-overexpressing Arabidopsis plants. (A and B) RNA gel blots of independent CaMV35S-E2Fa and CaMV35S-DPa transgenic plants, respectively. (C) Linkage of the observed growth arrest with the presence of both CaMV35S-E2Fa and CaMV35S-DPa transgenes. 1, wild-type plant; 2, cross between a CaMV35S-E2Fa plant and a control plant; 3, cross between a CaMV35S-DPa plant and a control plant; 4–13, individual siblings of a cross between a homozygous CaMV35S-E2Fa plant and a heterozygous CaMV35S-DPa. Lines marked with an asterisk had curled leaves and cotyledons, and were arrested at the seedling stage. Presence of transgenes was tested by PCR. (D) Transcript levels of S phase genes determined by semi-quantitative RT–PCR in control and CaMV35S-E2Fa–DPa plants harvested seven days after sowing.
Fig. 3. Phenotype of _E2Fa_- and _DPa_-overexpressing 12-day-old seedlings. (A) Untransformed control. (B) _E2Fa_- and (C) _E2Fa–DPa_-overexpressing plant. All plants were photographed at the same magnification. Bar = 2.5 mm.
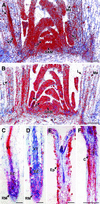
Fig. 4. Microscopic analysis of _E2Fa_- and _E2Fa–DPa_-overexpressing Arabidopsis plants. (A and E) Abaxial epidermis of cotyledons of a 5-day-old and 3-week-old control plants, respectively; (B and F) abaxial epidermis of cotyledons of a 5-day-old and 3-week-old E2Fa plant, respectively; (C and D) palisade parenchyma of a 5-day-old control and E2Fa plant, respectively; (G and H) hypocotyl of a 12-day-old control and E2Fa plant, respectively; (I and J) hypocotyl of a 12-day-old and 3-week-old E2Fa–DPa plant, respectively and (K and L) scanning micrographs of (G) and (I). Arrowheads in (D) and (F) point to novel synthesized cell walls; asterisks in (G) and (H) indicate cell file in which stomata are formed. Bars = 100 µm (A–D, K and L, same magnification) and 50 µm (E–J, same magnification).
Fig. 5. Microscopic analysis of a mature cotyledon of control and _E2Fa–DPa_-overexpressing plants. (A and B) Transverse section through the central part of a cotyledon of a control and E2Fa–DPa plant, respectively. (C) Group of small non-vacuolated cells with dense cytoplasm located at the epidermis of a cotyledon of an _E2Fa–DPa_-overexpressing plant. (D and E) Detail of cotelydon palisade parenchyma cells of wild-type and E2Fa–DPa plants, respectively. (F) E2Fa–DPa cotyledon palisade parenchyma cell containing two giant nuclei. Arrowheads point to nuclei (D–F). Scale bars: 500 µm (A and B, same magnification) and 50 µm (C and D; D–F, same magnification).
Fig. 6. Microscopic analysis of root tissue. Median, longitudinal section through a 3-week-old control (A) and E2Fa–DPa plant (B). Arrowheads point to nuclei. C, cortex; En, endodermis; Ep, epidermis; RC, root cap. Scale bar = 100 µm (A and B, same magnification).
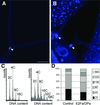
Fig. 7. DNA ploidy level in control and CaMV35S-E2Fa–DPa transgenic plants. (A and B) Trichome of control and E2Fa–DPa transgenic plant, respectively. Arrowheads point to the nucleus. (C) Ploidy distribution of control (left) and E2Fa–DPa transgenic seedlings (right) harvested 12 days after germination. (D) Quantification of the results shown in (C). Bar = 50 µm (A and B, same magnification).
Fig. 8. Model explaining the different phenotypes seen in plants with sustained E2Fa–DPa activity. MIF, mitosis-inducing factor.
Similar articles
- The plant-specific cyclin-dependent kinase CDKB1;1 and transcription factor E2Fa-DPa control the balance of mitotically dividing and endoreduplicating cells in Arabidopsis.
Boudolf V, Vlieghe K, Beemster GT, Magyar Z, Torres Acosta JA, Maes S, Van Der Schueren E, Inzé D, De Veylder L. Boudolf V, et al. Plant Cell. 2004 Oct;16(10):2683-92. doi: 10.1105/tpc.104.024398. Epub 2004 Sep 17. Plant Cell. 2004. PMID: 15377755 Free PMC article. - The DP-E2F-like gene DEL1 controls the endocycle in Arabidopsis thaliana.
Vlieghe K, Boudolf V, Beemster GT, Maes S, Magyar Z, Atanassova A, de Almeida Engler J, De Groodt R, Inzé D, De Veylder L. Vlieghe K, et al. Curr Biol. 2005 Jan 11;15(1):59-63. doi: 10.1016/j.cub.2004.12.038. Curr Biol. 2005. PMID: 15649366 - Microarray analysis of E2Fa-DPa-overexpressing plants uncovers a cross-talking genetic network between DNA replication and nitrogen assimilation.
Vlieghe K, Vuylsteke M, Florquin K, Rombauts S, Maes S, Ormenese S, Van Hummelen P, Van de Peer Y, Inze D, De Veylder L. Vlieghe K, et al. J Cell Sci. 2003 Oct 15;116(Pt 20):4249-59. doi: 10.1242/jcs.00715. Epub 2003 Sep 2. J Cell Sci. 2003. PMID: 12953064 - Constitutive E2F expression in tobacco plants exhibits altered cell cycle control and morphological change in a cell type-specific manner.
Kosugi S, Ohashi Y. Kosugi S, et al. Plant Physiol. 2003 Aug;132(4):2012-22. doi: 10.1104/pp.103.025080. Plant Physiol. 2003. PMID: 12913157 Free PMC article. - Genome-wide identification of potential plant E2F target genes.
Vandepoele K, Vlieghe K, Florquin K, Hennig L, Beemster GT, Gruissem W, Van de Peer Y, Inzé D, De Veylder L. Vandepoele K, et al. Plant Physiol. 2005 Sep;139(1):316-28. doi: 10.1104/pp.105.066290. Epub 2005 Aug 26. Plant Physiol. 2005. PMID: 16126853 Free PMC article.
Cited by
- Enhancement of secondary xylem cell proliferation by Arabidopsis cyclin D overexpression in tobacco plants.
Fujii T, Sato K, Matsui N, Furuichi T, Takenouchi S, Nishikubo N, Suzuki Y, Kawai S, Demura T, Kajita S, Katayama Y. Fujii T, et al. Plant Cell Rep. 2012 Sep;31(9):1573-80. doi: 10.1007/s00299-012-1271-7. Epub 2012 May 1. Plant Cell Rep. 2012. PMID: 22547095 - The E2FC-DPB Transcription Factor Controls Cell Division, Endoreplication and Lateral Root Formation in a SCF-Dependent Manner.
Del Pozo JC, Diaz-Trivino S, Cisneros N, Gutierrez C. Del Pozo JC, et al. Plant Signal Behav. 2007 Jul;2(4):273-4. doi: 10.4161/psb.2.4.3897. Plant Signal Behav. 2007. PMID: 19704635 Free PMC article. - Combining growth-promoting genes leads to positive epistasis in Arabidopsis thaliana.
Vanhaeren H, Gonzalez N, Coppens F, De Milde L, Van Daele T, Vermeersch M, Eloy NB, Storme V, Inzé D. Vanhaeren H, et al. Elife. 2014 Apr 29;3:e02252. doi: 10.7554/eLife.02252. Elife. 2014. PMID: 24843021 Free PMC article. - A mutation of casein kinase 2 α4 subunit affects multiple developmental processes in Arabidopsis.
Wang WS, Zhu J, Zhang KX, Lü YT, Xu HH. Wang WS, et al. Plant Cell Rep. 2016 May;35(5):1071-80. doi: 10.1007/s00299-016-1939-5. Epub 2016 Feb 16. Plant Cell Rep. 2016. PMID: 26883224 - Development of a rapamycin-inducible protein-knockdown system in the unicellular red alga Cyanidioschyzon merolae.
Fujiwara T, Hirooka S, Yamashita S, Yagisawa F, Miyagishima SY. Fujiwara T, et al. Plant Physiol. 2024 Sep 2;196(1):77-94. doi: 10.1093/plphys/kiae316. Plant Physiol. 2024. PMID: 38833589 Free PMC article.
References
- Albani D., Mariconti,L., Ricagno,S., Pitto,L., Moroni,C., Helin,K. and Cella,R. (2000) DcE2F, a functional plant E2F-like transcriptional activator from Daucus carota. J. Biol. Chem., 275, 19258–19267. - PubMed
- Beeckman T. and Viane,R. (2000) Embedding thin plant specimens for oriented sectioning. Biotech. Histochem., 75, 23–26. - PubMed
- Berger F., Linstead,P., Dolan,L. and Haseloff,J. (1998) Stomata patterning on the hypocotyl of Arabidopsis thaliana is controlled by genes involved in the control of root epidermis patterning. Dev. Biol., 194, 226–234. - PubMed
- Brehm A., Miska,E.A., McCance,D.J., Reid,J.L., Bannister,A.J. and Kouzarides,T. (1998) Retinoblastoma protein recruits histone deacetylase to repress transcription. Nature, 391, 597–601. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases