Functional interaction between DNA-PKcs and telomerase in telomere length maintenance - PubMed (original) (raw)
Functional interaction between DNA-PKcs and telomerase in telomere length maintenance
Silvia Espejel et al. EMBO J. 2002.
Abstract
DNA-PKcs is the catalytic subunit of the DNA-dependent protein kinase (DNA-PK) complex that functions in the non-homologous end-joining of double-strand breaks, and it has been shown previously to have a role in telomere capping. In particular, DNA-PKcs deficiency leads to chromosome fusions involving telomeres produced by leading-strand synthesis. Here, by generating mice doubly deficient in DNA-PKcs and telomerase (Terc(-/-)/DNA-PKcs(-/-)), we demonstrate that DNA-PKcs also has a fundamental role in telomere length maintenance. In particular, Terc(-/-)/DNA-PKcs(-/-) mice displayed an accelerated rate of telomere shortening when compared with Terc(-/-) controls, suggesting a functional interaction between both activities in maintaining telomere length. In addition, we also provide direct demonstration that DNA-PKcs is essential for both end-to-end fusions and apoptosis triggered by critically short telomeres. Our data predict that, in telomerase-deficient cells, i.e. human somatic cells, DNA-PKcs abrogation may lead to a faster rate of telomere degradation and cell cycle arrest in the absence of increased apoptosis and/or fusion of telomere-exhausted chromosomes. These results suggest a critical role of DNA-PKcs in both cancer and aging.
Figures
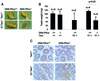
Fig. 1. Early infertility phenotype in G1 Terc–/–/DNA-PKcs–/– mice. (A) Representative images of testis derived from the indicated genotypes. Note the smaller testis size in the G1 Terc–/–/DNA-PKcs–/– mice. (B) Quantification of testis weight in the number of mice indicated from each genotype. Standard deviation bars are also shown. There was a statistically significant difference in testis weight between G1 Terc–/–/DNA-PKcs–/– and G1 Terc–/–/DNA-PKcs+/+ mice (P = 0.05). n = number of mice of the genotype indicated used in the study. (C) Representative images of testis sections from mice of the indicated genotypes. Arrows point to seminiferous tubules lacking meiotic cells in the G1 Terc–/–/DNA-PKcs–/– mice. (D) Quantification of BrdU-positive cells per meiotic tubule. For each mouse, >100 meiotic tubules were counted. n = number of mice of the genotype indicated used in the study. Bars represent average values for the number of mice indicated from each genotype. Standard deviation bars of the average values are shown. Statistical calculations are done using >100 values of BrdU+ cells/tubule for each individual mouse. G2 Terc–/–/DNA-PKcs+/+ mice showed significantly decreased BrdU incorporation compared with wild-type mice (P = 0.01). Similarly, G1 Terc–/–/DNA-PKcs–/– mice showed significantly decreased BrdU incorporation compared with the G1 Terc–/–/DNA-PKcs+/+ controls (P = 0.003). (E) Representative images of anti-BrdU-stained meiotic tubules from mice of the genotypes indicated. BrdU-positive cells are the ones with dark brown staining. (F) Quantification of TUNEL+ cells per 100 meiotic tubules. The total number of meiotic tubules analyzed from each mouse are shown in Table I (numbers in parenthesis). n = number of mice of the indicated genotype used in the study. Bars represent average values for the number of mice indicated from each genotype. Standard deviation bars of the average values are shown. Statistical calculations are done using between 339 and 2243 values of TUNEL+ cells/tubule for each genotype (Table I). G3 Terc–/–/DNA-PKcs+/+ mice showed significantly increased apoptosis compared with wild-type mice (P = 0.006). In contrast, G1 Terc–/–/DNA-PKcs–/– testis did not show increased apoptosis compared with the G1 Terc–/–/DNA-PKcs+/+ controls (P = 0.76). (G) Representative images of TUNEL+ cells in testis sections from mice of the genotypes indicated. Arrows indicate TUNEL+ cells that appear with dark brown staining. Note the low number of TUNEL+ cells in G1 Terc–/–/DNA-PKcs–/– testis.
Fig. 1. Early infertility phenotype in G1 Terc–/–/DNA-PKcs–/– mice. (A) Representative images of testis derived from the indicated genotypes. Note the smaller testis size in the G1 Terc–/–/DNA-PKcs–/– mice. (B) Quantification of testis weight in the number of mice indicated from each genotype. Standard deviation bars are also shown. There was a statistically significant difference in testis weight between G1 Terc–/–/DNA-PKcs–/– and G1 Terc–/–/DNA-PKcs+/+ mice (P = 0.05). n = number of mice of the genotype indicated used in the study. (C) Representative images of testis sections from mice of the indicated genotypes. Arrows point to seminiferous tubules lacking meiotic cells in the G1 Terc–/–/DNA-PKcs–/– mice. (D) Quantification of BrdU-positive cells per meiotic tubule. For each mouse, >100 meiotic tubules were counted. n = number of mice of the genotype indicated used in the study. Bars represent average values for the number of mice indicated from each genotype. Standard deviation bars of the average values are shown. Statistical calculations are done using >100 values of BrdU+ cells/tubule for each individual mouse. G2 Terc–/–/DNA-PKcs+/+ mice showed significantly decreased BrdU incorporation compared with wild-type mice (P = 0.01). Similarly, G1 Terc–/–/DNA-PKcs–/– mice showed significantly decreased BrdU incorporation compared with the G1 Terc–/–/DNA-PKcs+/+ controls (P = 0.003). (E) Representative images of anti-BrdU-stained meiotic tubules from mice of the genotypes indicated. BrdU-positive cells are the ones with dark brown staining. (F) Quantification of TUNEL+ cells per 100 meiotic tubules. The total number of meiotic tubules analyzed from each mouse are shown in Table I (numbers in parenthesis). n = number of mice of the indicated genotype used in the study. Bars represent average values for the number of mice indicated from each genotype. Standard deviation bars of the average values are shown. Statistical calculations are done using between 339 and 2243 values of TUNEL+ cells/tubule for each genotype (Table I). G3 Terc–/–/DNA-PKcs+/+ mice showed significantly increased apoptosis compared with wild-type mice (P = 0.006). In contrast, G1 Terc–/–/DNA-PKcs–/– testis did not show increased apoptosis compared with the G1 Terc–/–/DNA-PKcs+/+ controls (P = 0.76). (G) Representative images of TUNEL+ cells in testis sections from mice of the genotypes indicated. Arrows indicate TUNEL+ cells that appear with dark brown staining. Note the low number of TUNEL+ cells in G1 Terc–/–/DNA-PKcs–/– testis.
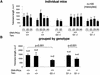
Fig. 2. Telomere length determination using Q-FISH on testis sections. (A) Average telomere fluorescence in arbitrary units (a.u.f.) of 100 meiotic cells from each mouse of the indicated genotype. Standard deviation, as well as the total number of meiocytes analyzed per mouse are indicated. (B) Average telomere fluorescence in arbitrary units (a.u.f.) of three or four mice grouped per genotype. Standard deviation, as well as the total number of mice of each genotype, are indicated. There was a significant decrease in average telomere fluorescence in G1 Terc–/–/DNA-PKcs–/– meiotic cells compared with G1 Terc–/–/DNA-PKcs+/+ controls (P < 0.001).
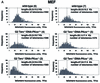
Fig. 3. (A) Telomere length distribution in primary MEFs from the indicated genotype. One telomere fluorescence unit (TFU) corresponds to 1 kb of TTAGGG repeats (Zijlmans et al., 1997). Average telomere length and the standard deviation, as well as the total number of telomeres analyzed, are indicated. The distribution of the telomere length frequencies for each MEF is an indication of the standard deviation. Notice the marked shift towards short telomeres in Terc–/–/DNA-PKcs–/– MEFs compared with the corresponding Terc–/–/DNA-PKcs+/+ controls. (B) Telomere length distribution in primary splenocytes from the indicated genotype. One TFU corresponds to 1 kb of TTAGGG repeats (Zijlmans et al., 1997). Average telomere length and the standard deviation, as well as the total number of telomeres analyzed, are indicated. The distribution of the telomere length frequencies for each splenocyte culture is an indication of the standard deviation. Notice the marked shift towards short telomeres in Terc–/–/DNA-PKcs–/– splenocytes compared with the corresponding Terc–/–/DNA-PKcs+/+ controls. (C) G-strand overhangs using liver nuclei visualized in native gel after hybridization with a (CCCTAA)4 probe. Notice that upon treatment with 100 U of mung bean nuclease (Mbn) the G-strand-specific signal decreases. Note that the G-strand overhang signal was present at lower molecular weights in the G3 and G4 Terc–/–/DNA-PKcs–/– liver cells compared with the corresponding G3 and G4 Terc–/–/DNA-PKcs+/+ controls, again indicating the presence of shorter telomeres in Terc–/–/DNA-PKcs–/– cells. Mice numbers 116 and 117 are littermates. Quantification of G-strand signals is also shown (see Materials and methods for details).
Fig. 3. (A) Telomere length distribution in primary MEFs from the indicated genotype. One telomere fluorescence unit (TFU) corresponds to 1 kb of TTAGGG repeats (Zijlmans et al., 1997). Average telomere length and the standard deviation, as well as the total number of telomeres analyzed, are indicated. The distribution of the telomere length frequencies for each MEF is an indication of the standard deviation. Notice the marked shift towards short telomeres in Terc–/–/DNA-PKcs–/– MEFs compared with the corresponding Terc–/–/DNA-PKcs+/+ controls. (B) Telomere length distribution in primary splenocytes from the indicated genotype. One TFU corresponds to 1 kb of TTAGGG repeats (Zijlmans et al., 1997). Average telomere length and the standard deviation, as well as the total number of telomeres analyzed, are indicated. The distribution of the telomere length frequencies for each splenocyte culture is an indication of the standard deviation. Notice the marked shift towards short telomeres in Terc–/–/DNA-PKcs–/– splenocytes compared with the corresponding Terc–/–/DNA-PKcs+/+ controls. (C) G-strand overhangs using liver nuclei visualized in native gel after hybridization with a (CCCTAA)4 probe. Notice that upon treatment with 100 U of mung bean nuclease (Mbn) the G-strand-specific signal decreases. Note that the G-strand overhang signal was present at lower molecular weights in the G3 and G4 Terc–/–/DNA-PKcs–/– liver cells compared with the corresponding G3 and G4 Terc–/–/DNA-PKcs+/+ controls, again indicating the presence of shorter telomeres in Terc–/–/DNA-PKcs–/– cells. Mice numbers 116 and 117 are littermates. Quantification of G-strand signals is also shown (see Materials and methods for details).
Fig. 3. (A) Telomere length distribution in primary MEFs from the indicated genotype. One telomere fluorescence unit (TFU) corresponds to 1 kb of TTAGGG repeats (Zijlmans et al., 1997). Average telomere length and the standard deviation, as well as the total number of telomeres analyzed, are indicated. The distribution of the telomere length frequencies for each MEF is an indication of the standard deviation. Notice the marked shift towards short telomeres in Terc–/–/DNA-PKcs–/– MEFs compared with the corresponding Terc–/–/DNA-PKcs+/+ controls. (B) Telomere length distribution in primary splenocytes from the indicated genotype. One TFU corresponds to 1 kb of TTAGGG repeats (Zijlmans et al., 1997). Average telomere length and the standard deviation, as well as the total number of telomeres analyzed, are indicated. The distribution of the telomere length frequencies for each splenocyte culture is an indication of the standard deviation. Notice the marked shift towards short telomeres in Terc–/–/DNA-PKcs–/– splenocytes compared with the corresponding Terc–/–/DNA-PKcs+/+ controls. (C) G-strand overhangs using liver nuclei visualized in native gel after hybridization with a (CCCTAA)4 probe. Notice that upon treatment with 100 U of mung bean nuclease (Mbn) the G-strand-specific signal decreases. Note that the G-strand overhang signal was present at lower molecular weights in the G3 and G4 Terc–/–/DNA-PKcs–/– liver cells compared with the corresponding G3 and G4 Terc–/–/DNA-PKcs+/+ controls, again indicating the presence of shorter telomeres in Terc–/–/DNA-PKcs–/– cells. Mice numbers 116 and 117 are littermates. Quantification of G-strand signals is also shown (see Materials and methods for details).
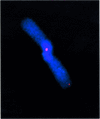
Fig. 4. Representative CO-FISH image of a chromatid-type dicentric leading-strand telomere fusion present in G2 Terc–/–/DNA-PKcs–/– cells. Blue, DAPI; red, TTAGGG signal.
Fig. 5. Working model showing that DNA-PKcs is essential for both telomere length maintenance (as shown here for Terc–/–/DNA-PKcs–/– mice) and for proper telomere processing (as shown here, as well as elsewhere; Bailey et al., 2001). In addition, the results shown here also demonstrate that DNA-PKcs mediates both end-to-end fusion and apoptosis produced by critically short telomeres. In the absence of both Terc and DNA-PKcs, telomeres suffer a faster rate of shortening; however, these prematurely shortened telomeres do not result in end-to-end fusions or apoptosis.
Similar articles
- Impact of telomerase ablation on organismal viability, aging, and tumorigenesis in mice lacking the DNA repair proteins PARP-1, Ku86, or DNA-PKcs.
Espejel S, Klatt P, Ménissier-de Murcia J, Martín-Caballero J, Flores JM, Taccioli G, de Murcia G, Blasco MA. Espejel S, et al. J Cell Biol. 2004 Nov 22;167(4):627-38. doi: 10.1083/jcb.200407178. Epub 2004 Nov 15. J Cell Biol. 2004. PMID: 15545322 Free PMC article. - DNA-dependent protein kinase catalytic subunit is not required for dysfunctional telomere fusion and checkpoint response in the telomerase-deficient mouse.
Maser RS, Wong KK, Sahin E, Xia H, Naylor M, Hedberg HM, Artandi SE, DePinho RA. Maser RS, et al. Mol Cell Biol. 2007 Mar;27(6):2253-65. doi: 10.1128/MCB.01354-06. Epub 2006 Dec 4. Mol Cell Biol. 2007. PMID: 17145779 Free PMC article. - Chromosomal end-to-end fusions in immortalized mouse embryonic fibroblasts deficient in the DNA-dependent protein kinase catalytic subunit.
Rebuzzini P, Lisa A, Giulotto E, Mondello C. Rebuzzini P, et al. Cancer Lett. 2004 Jan 8;203(1):79-86. doi: 10.1016/j.canlet.2003.08.028. Cancer Lett. 2004. PMID: 14670620 - DNA and telomeres: beginnings and endings.
Bailey SM, Goodwin EH. Bailey SM, et al. Cytogenet Genome Res. 2004;104(1-4):109-15. doi: 10.1159/000077474. Cytogenet Genome Res. 2004. PMID: 15162023 Review. - DNA repair factors and telomere-chromosome integrity in mammalian cells.
Hande MP. Hande MP. Cytogenet Genome Res. 2004;104(1-4):116-22. doi: 10.1159/000077475. Cytogenet Genome Res. 2004. PMID: 15162024 Review.
Cited by
- Shorter telomeres, accelerated ageing and increased lymphoma in DNA-PKcs-deficient mice.
Espejel S, Martín M, Klatt P, Martín-Caballero J, Flores JM, Blasco MA. Espejel S, et al. EMBO Rep. 2004 May;5(5):503-9. doi: 10.1038/sj.embor.7400127. Epub 2004 Apr 23. EMBO Rep. 2004. PMID: 15105825 Free PMC article. - HTLV-1 Tax oncoprotein subverts the cellular DNA damage response via binding to DNA-dependent protein kinase.
Durkin SS, Guo X, Fryrear KA, Mihaylova VT, Gupta SK, Belgnaoui SM, Haoudi A, Kupfer GM, Semmes OJ. Durkin SS, et al. J Biol Chem. 2008 Dec 26;283(52):36311-20. doi: 10.1074/jbc.M804931200. Epub 2008 Oct 27. J Biol Chem. 2008. PMID: 18957425 Free PMC article. - An increase in telomere sister chromatid exchange in murine embryonic stem cells possessing critically shortened telomeres.
Wang Y, Erdmann N, Giannone RJ, Wu J, Gomez M, Liu Y. Wang Y, et al. Proc Natl Acad Sci U S A. 2005 Jul 19;102(29):10256-60. doi: 10.1073/pnas.0504635102. Epub 2005 Jul 6. Proc Natl Acad Sci U S A. 2005. PMID: 16000404 Free PMC article. - Role of mammalian Rad54 in telomere length maintenance.
Jaco I, Muñoz P, Goytisolo F, Wesoly J, Bailey S, Taccioli G, Blasco MA. Jaco I, et al. Mol Cell Biol. 2003 Aug;23(16):5572-80. doi: 10.1128/MCB.23.16.5572-5580.2003. Mol Cell Biol. 2003. PMID: 12897131 Free PMC article. - The human telomerase RNA component, hTR, activates the DNA-dependent protein kinase to phosphorylate heterogeneous nuclear ribonucleoprotein A1.
Ting NS, Pohorelic B, Yu Y, Lees-Miller SP, Beattie TL. Ting NS, et al. Nucleic Acids Res. 2009 Oct;37(18):6105-15. doi: 10.1093/nar/gkp636. Epub 2009 Aug 5. Nucleic Acids Res. 2009. PMID: 19656952 Free PMC article.
References
- Bailey S.M., Conforth,M.N., Kurimasa,A., Chen,D.J. and Goodwin,E.H. (2001) Strand-specific postreplicative processing of mammalian telomeres. Science, 293, 2462–2465. - PubMed
- Bianchi A. and de Lange,T. (1999) Ku binds telomeric DNA in vitro. J. Biol. Chem., 274, 21223–21227. - PubMed
- Blackburn E.H. (2001) Switching and signaling at the telomere. Cell, 106, 661–673. - PubMed
- Blasco M.A., Lee,H.-W., Hande,P., Samper,E., Lansdorp,P., DePinho,R. and Greider,C.W. (1997) Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell, 91, 25–34. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases