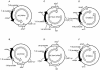Identification of factors influencing strand bias in oligonucleotide-mediated recombination in Escherichia coli - PubMed (original) (raw)
Identification of factors influencing strand bias in oligonucleotide-mediated recombination in Escherichia coli
Xin-tian Li et al. Nucleic Acids Res. 2003.
Abstract
Recombinogenic engineering methodology, also known as recombineering, utilizes homologous recombination to create targeted changes in cellular DNA with great specificity and flexibility. In Escherichia coli, the Red recombination system from bacteriophage lambda has been used successfully to modify both plasmid and chromosomal DNA in a highly efficient manner, using either a linear double-stranded DNA fragment or a synthetic single-stranded oligonucleotide (SSO). The current model for Red/SSO-mediated recombination involves the SSO first annealing to a transient, single-stranded region of DNA before being incorporated into the chromosome or plasmid target. It has been observed previously, in both eukaryotes and prokaryotes, that mutations in the two strands of the DNA double helix are 'corrected' by complementary SSOs with differing efficiencies. Here we investigate further the factors that influence the strand bias as well as the overall efficiency of Red/SSO-mediated recombination in E.coli. We show that the direction of DNA replication and the nature of the SSO-encoded mismatch are the main factors dictating the recombinational strand bias. However, the influence that the SSO-encoded mismatch exerts upon the recombinational strand bias is abolished in E.coli strains that are defective in mismatch repair (MMR). This reflects the fact that different base-base mispairs are corrected by the mutS/H/L-dependent MMR pathway with differing efficiencies. Furthermore, our data indicate that transcription has negligible influence on the strand bias. These results demonstrate for the first time that the interplay between DNA replication and MMR has a major effect on the efficiency and strand bias of Red/SSO-mediated recombination in E.coli.
Figures
Figure 1
Key features of the plasmids and chromosomes containing the mkan reporter gene. The solid black arrows indicate the orientation of the genes, while the solid grey arrows specify the direction of transcription (under the control of the Tn5 or Lac promoters). The open arrows inside the plasmid (A–D) or chromosome (E and F) denote the direction of replication from the ori. In (A), (C) and (E), the direction of transcription of the reporter genes is the same as that of DNA replication. In these reporter genes, the non-template strands (NT) are the same as the leading strands in replication. In (B), (D) and (F), the direction of transcription of the reporter gene is opposite to the direction of DNA replication, hence the NT strands are also the lagging strands in replication. In (A) and (B), and in (E) and (F), the mutated kanamycin (mkan) gene is under the control of the Tn5 promoter, whilst in (C) and (D) it is under the control of the inducible Lac promoter. oriC is the E.coli chromosomal replication origin. (G) Shows that the wild type kan gene sequence (TAT), the amber mutant (TAG), the sequences of the 12A, 12B, 12A-TAT, 12B-ATA oligonucleotides and the corresponding sequences (TAC) or (TAT) after recombination. Drawings are not to scale.
Figure 1
Key features of the plasmids and chromosomes containing the mkan reporter gene. The solid black arrows indicate the orientation of the genes, while the solid grey arrows specify the direction of transcription (under the control of the Tn5 or Lac promoters). The open arrows inside the plasmid (A–D) or chromosome (E and F) denote the direction of replication from the ori. In (A), (C) and (E), the direction of transcription of the reporter genes is the same as that of DNA replication. In these reporter genes, the non-template strands (NT) are the same as the leading strands in replication. In (B), (D) and (F), the direction of transcription of the reporter gene is opposite to the direction of DNA replication, hence the NT strands are also the lagging strands in replication. In (A) and (B), and in (E) and (F), the mutated kanamycin (mkan) gene is under the control of the Tn5 promoter, whilst in (C) and (D) it is under the control of the inducible Lac promoter. oriC is the E.coli chromosomal replication origin. (G) Shows that the wild type kan gene sequence (TAT), the amber mutant (TAG), the sequences of the 12A, 12B, 12A-TAT, 12B-ATA oligonucleotides and the corresponding sequences (TAC) or (TAT) after recombination. Drawings are not to scale.
Figure 2
The strand bias observed in SSO-mediated recombination with plasmid DNA. Plasmid (10 ng) and SSO DNA were co-transformed into recombination-competent DY380. Aliquots of the plasmid/SSO-cotransformed cells were plated onto LB + Amp or LB + Amp + Kan plates, and after incubation overnight at 32°C, the number of colonies was counted. The plasmid and SSO used in each experiment are noted at the bottom of each panel under ‘target’ and ‘SSO Sequence’, respectively. T (template) or NT (non-template) indicates that the SSO has the same sequence as the transcriptional template, or the transcript, respectively. Lag or Lead indicates that the SSO has the same sequence as the newly synthesized Okazaki fragment or the leading strand in replication, respectively. The _y_-axis represents the number of kan r colonies per 100 amp r colonies after co-transformation of SSO and reporter plasmid. At least six independent experiments were carried out for each set of conditions. In (A) and (B) the length of SSO was kept constant (91mer), but the concentration was varied from 10 to 1000 ng (per reaction). In (C) and (D) three lengths of oligonucleotide were used: 25 (6A and 6B), 73 (8A and 8B) and 91 nt (12A and 12B), with the concentration maintained at 100 ng per reaction. All SSOs in this figure introduce TAG to TAC or CTA to GTA substitutions, with this mismatch located at the centre of the oligo.
Figure 3
The strand bias observed in SSO-mediated recombination with chromosomal DNA. SSOs encoding for TAG to TAC or CTA to GTA substitutions were transformed into different DY380 and DY330 derived strains, as indicated at the bottom of each panel. Aliquots of the SSO-treated cells were spread onto LB + Amp or LB + Amp + Kan plates, with the number of colonies that appeared after overnight incubation at 32°C counted. The _y_-axis shows the number of kan r colonies per 10 000 colonies after transformation of SSO (i.e. a measure of the number of recombinants among cells that ‘survived’ electroporation with SSO). In (A) and (B) the length of the correction-SSO was kept constant (91mer), but the concentration was varied from 10 to 1000 ng. In (C) and (D) three lengths of oligo were used: 25 (6A and 6B), 73 (8A and 8B) and 91 nt (12A and 12B), with the concentration maintained at 100 ng per reaction. In (E) and (F), oligos 12A and 12B (100 ng per reaction) were transformed into DY330(+) and DY330(–) as indicated. For (A) through to (F), a minimum of six independent experiments were performed for each set of conditions.
Figure 3
The strand bias observed in SSO-mediated recombination with chromosomal DNA. SSOs encoding for TAG to TAC or CTA to GTA substitutions were transformed into different DY380 and DY330 derived strains, as indicated at the bottom of each panel. Aliquots of the SSO-treated cells were spread onto LB + Amp or LB + Amp + Kan plates, with the number of colonies that appeared after overnight incubation at 32°C counted. The _y_-axis shows the number of kan r colonies per 10 000 colonies after transformation of SSO (i.e. a measure of the number of recombinants among cells that ‘survived’ electroporation with SSO). In (A) and (B) the length of the correction-SSO was kept constant (91mer), but the concentration was varied from 10 to 1000 ng. In (C) and (D) three lengths of oligo were used: 25 (6A and 6B), 73 (8A and 8B) and 91 nt (12A and 12B), with the concentration maintained at 100 ng per reaction. In (E) and (F), oligos 12A and 12B (100 ng per reaction) were transformed into DY330(+) and DY330(–) as indicated. For (A) through to (F), a minimum of six independent experiments were performed for each set of conditions.
Figure 4
Recombinational strand bias in the absence of MMR. (A) SSOs (12A-TAT or 12B-ATA, 100 ng) and (B) SSOs (12A or 12B, 100 ng) were transformed into four different strains of recombination-competent E.coli, as indicated underneath each panel. Aliquots of the treated cells were spread onto LB + Amp or LB + Amp + Kan plates and after incubation overnight incubation at 32°C, the number of colonies that appeared was counted. The _y_-axis indicates the number of kan r colonies per 10 000 survived colonies after transformation of SSO. (C) Plasmids (10 ng) and SSOs (12A or 12B, 100 ng) were co-transformed into the indicated recombination-competent E.coli strains. Aliquots were spread onto LB + Amp or LB + Amp + Kan plates, and after overnight incubation at 32°C, the number of colonies that appeared was counted. The _y_-axis shows the number of kan r colonies per 100 amp r colonies after co-transformation of SSO and reporter plasmid.
Figure 4
Recombinational strand bias in the absence of MMR. (A) SSOs (12A-TAT or 12B-ATA, 100 ng) and (B) SSOs (12A or 12B, 100 ng) were transformed into four different strains of recombination-competent E.coli, as indicated underneath each panel. Aliquots of the treated cells were spread onto LB + Amp or LB + Amp + Kan plates and after incubation overnight incubation at 32°C, the number of colonies that appeared was counted. The _y_-axis indicates the number of kan r colonies per 10 000 survived colonies after transformation of SSO. (C) Plasmids (10 ng) and SSOs (12A or 12B, 100 ng) were co-transformed into the indicated recombination-competent E.coli strains. Aliquots were spread onto LB + Amp or LB + Amp + Kan plates, and after overnight incubation at 32°C, the number of colonies that appeared was counted. The _y_-axis shows the number of kan r colonies per 100 amp r colonies after co-transformation of SSO and reporter plasmid.
Figure 4
Recombinational strand bias in the absence of MMR. (A) SSOs (12A-TAT or 12B-ATA, 100 ng) and (B) SSOs (12A or 12B, 100 ng) were transformed into four different strains of recombination-competent E.coli, as indicated underneath each panel. Aliquots of the treated cells were spread onto LB + Amp or LB + Amp + Kan plates and after incubation overnight incubation at 32°C, the number of colonies that appeared was counted. The _y_-axis indicates the number of kan r colonies per 10 000 survived colonies after transformation of SSO. (C) Plasmids (10 ng) and SSOs (12A or 12B, 100 ng) were co-transformed into the indicated recombination-competent E.coli strains. Aliquots were spread onto LB + Amp or LB + Amp + Kan plates, and after overnight incubation at 32°C, the number of colonies that appeared was counted. The _y_-axis shows the number of kan r colonies per 100 amp r colonies after co-transformation of SSO and reporter plasmid.
Similar articles
- Highly efficient deletion method for the engineering of plasmid DNA with single-stranded oligonucleotides.
Lu LY, Huen MS, Tai AC, Liu DP, Cheah KS, Huang JD. Lu LY, et al. Biotechniques. 2008 Feb;44(2):217-20, 222, 224. doi: 10.2144/000112684. Biotechniques. 2008. PMID: 18330349 - In vitro and in vivo studies of MutS, MutL and MutH mutants: correlation of mismatch repair and DNA recombination.
Junop MS, Yang W, Funchain P, Clendenin W, Miller JH. Junop MS, et al. DNA Repair (Amst). 2003 Apr 2;2(4):387-405. doi: 10.1016/s1568-7864(02)00245-8. DNA Repair (Amst). 2003. PMID: 12606120 - DNA mismatch repair and mutation avoidance pathways.
Marti TM, Kunz C, Fleck O. Marti TM, et al. J Cell Physiol. 2002 Apr;191(1):28-41. doi: 10.1002/jcp.10077. J Cell Physiol. 2002. PMID: 11920679 Review. - Mechanisms underlying targeted gene correction using chimeric RNA/DNA and single-stranded DNA oligonucleotides.
Andersen MS, Sørensen CB, Bolund L, Jensen TG. Andersen MS, et al. J Mol Med (Berl). 2002 Dec;80(12):770-81. doi: 10.1007/s00109-002-0393-8. Epub 2002 Nov 22. J Mol Med (Berl). 2002. PMID: 12483462 Review.
Cited by
- Genome engineering.
Carr PA, Church GM. Carr PA, et al. Nat Biotechnol. 2009 Dec;27(12):1151-62. doi: 10.1038/nbt.1590. Nat Biotechnol. 2009. PMID: 20010598 Review. - Single-stranded heteroduplex intermediates in lambda Red homologous recombination.
Maresca M, Erler A, Fu J, Friedrich A, Zhang Y, Stewart AF. Maresca M, et al. BMC Mol Biol. 2010 Jul 29;11:54. doi: 10.1186/1471-2199-11-54. BMC Mol Biol. 2010. PMID: 20670401 Free PMC article. - The operator and early promoter region of the Shiga toxin type 2-encoding bacteriophage 933W and control of toxin expression.
Tyler JS, Mills MJ, Friedman DI. Tyler JS, et al. J Bacteriol. 2004 Nov;186(22):7670-9. doi: 10.1128/JB.186.22.7670-7679.2004. J Bacteriol. 2004. PMID: 15516581 Free PMC article. - LNA modification of single-stranded DNA oligonucleotides allows subtle gene modification in mismatch-repair-proficient cells.
van Ravesteyn TW, Dekker M, Fish A, Sixma TK, Wolters A, Dekker RJ, Te Riele HP. van Ravesteyn TW, et al. Proc Natl Acad Sci U S A. 2016 Apr 12;113(15):4122-7. doi: 10.1073/pnas.1513315113. Epub 2016 Mar 7. Proc Natl Acad Sci U S A. 2016. PMID: 26951689 Free PMC article. - Manipulating replisome dynamics to enhance lambda Red-mediated multiplex genome engineering.
Lajoie MJ, Gregg CJ, Mosberg JA, Washington GC, Church GM. Lajoie MJ, et al. Nucleic Acids Res. 2012 Dec;40(22):e170. doi: 10.1093/nar/gks751. Epub 2012 Aug 16. Nucleic Acids Res. 2012. PMID: 22904085 Free PMC article.
References
- Copeland N.G., Jenkins,N.A. and Court,D.L. (2001) Recombineering: a powerful new tool for mouse functional genomics. Nature Rev. Genet., 2, 769–779. - PubMed
- Court D.L., Sawitzke,J.A. and Thomason,L.C. (2002) Genetic engineering using homologous recombination. Annu. Rev. Genet., 36, 361–388. - PubMed
- Zhang Y., Buchholz,F., Muyrers,J.P.P. and Stewart,A.F. (1998) A new logic for DNA engineering using recombination in Escherichia coli. Nature Genet., 20, 123–128. - PubMed