Functions of eIF3 downstream of 48S assembly impact AUG recognition and GCN4 translational control - PubMed (original) (raw)
Functions of eIF3 downstream of 48S assembly impact AUG recognition and GCN4 translational control
Klaus H Nielsen et al. EMBO J. 2004.
Abstract
The binding of eIF2-GTP-tRNA(i)(Met) ternary complex (TC) to 40S subunits is impaired in yeast prt1-1 (eIF3b) mutant extracts, but evidence is lacking that TC recruitment is a critical function of eIF3 in vivo. If TC binding was rate-limiting in prt1-1 cells, overexpressing TC should suppress the temperature-sensitive phenotype and GCN4 translation should be strongly derepressed in this mutant, but neither was observed. Rather, GCN4 translation is noninducible in prt1-1 cells, and genetic analysis indicates defective ribosomal scanning between the upstream open reading frames that mediate translational control. prt1-1 cells also show reduced utilization of a near-cognate start codon, implicating eIF3 in AUG selection. Using in vivo cross-linking, we observed accumulation of TC and mRNA/eIF4G on 40S subunits and a 48S 'halfmer' in prt1-1 cells. Genetic evidence suggests that 40S-60S subunit joining is not rate-limiting in the prt1-1 mutant. Thus, eIF3b functions between 48S assembly and subunit joining to influence AUG recognition and reinitiation on GCN4 mRNA. Other mutations that disrupt eIF2-eIF3 contacts in the multifactor complex (MFC) diminished 40S-bound TC, indicating that MFC formation enhances 43S assembly in vivo.
Figures
Figure 1
eIF2 remains bound to 40S subunits in extracts of prt1-1 cells incubated at the nonpermissive temperature. (A) Isogenic PRT1 (H2879) and prt1-1 (H1676) cells were grown in YPD medium at 25°C and treated for 20 min at 37°C. Cyclohexamide was added to 50 μg/ml just prior to harvesting and WCEs prepared with heparin (200 μg/ml) in the breaking buffer were separated on a 4.5–45% sucrose gradient by centrifugation at 39 000 r.p.m. for 2.5 h. The gradients were collected and scanned at 254 nm to visualize the ribosomal species. (B) WCEs described in (A) were separated on a 7.5–30% sucrose gradient by centrifugation at 41 000 r.p.m. for 5 h. Proteins were subjected to Western analysis using antibodies against the proteins listed between the blots. An aliquot of each WCE was analyzed in parallel (In, input).
Figure 2
TC and mRNA remain bound to 40S subunits in HCHO-treated prt1-1 cells at the nonpermissive temperature. PRT1 (H2879) and prt1-1 (H1676) cells were grown in YPD at 25°C, heat-treated for 20 min at 37°C, and cross-linked with HCHO for 15 min (A) or 1 h (B, C). (A) WCEs were separated and analyzed as in Figure 1A. (B, C) WCEs were separated and treated as in Figure 1B, except that each fraction was divided and analyzed by Western and Northern blotting.
Figure 3
sui3-td and cdc33-1 mutations reduce the binding of TC and mRNA to 40S subunits in vivo. (A–C) sui3-td (YAJ18-3) cells were grown in SC-raffinose in the presence of 0.1 M copper sulfate at 25°C and shifted to SC-raffinose+galactose in the absence of copper and grown overnight at 37°C. Cells were cross-linked with HCHO and analyzed as described in Figure 2A–C, except that eIF2βtd was detected in (B) using HA antibody. (D) Strain F324 was transformed with YEplac195-CDC33-URA3 (CDC33) or empty vector (cdc33-1) and the resulting transformants were grown in SC-Ura medium at 25°C, heat treated for 2 h at 37°C, and cross-linked with HCHO. WCEs were prepared and analyzed as (B–C).
Figure 4
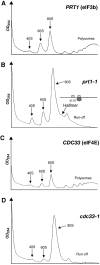
The prt1-1 mutant displays a halfmer phenotype at the nonpermissive temperature. Polysome profiles of (A) PRT1 (H2879) and (B) prt1-1 (H1676) cells after 20 min at 37°C. The halfmer shoulder on the 80S peak is indicated along with an explanatory schematic of a 1½-mer (see text). Polysome profiles of the (C) CDC33 and (D) cdc33-1 transformants described in Figure 3D after 2 h at 37°C.
Figure 5
The prt1-1 strain has a strong Gcn− phenotype and its Ts− phenotype is not suppressed by TC overexpression. (A) prt1-1 (H1676) and gcd1-502 (H70) cells were transformed with hc plasmid p3000 encoding hc TC, or empty vector, streaked on SC-Ura plates and incubated for 4 (left panels) or 2 days (right panels) at 33°C. (B) prt1-1 (YKHN60) cells were transformed with p3000 (hc TC) or empty vector, grown overnight in SC-Ura, and 10-fold serial dilutions were spotted in rows 1 and 2 on SC-Ura plates or SC-Ura-His plates containing 40 mM leucine with 30 mM 3-AT and incubated for 7 and 10 days, respectively, at 33°C. PRT1 GCN2 (H2879) and _PRT1 gcn2_Δ (H2881) cells transformed with empty vector were analyzed in parallel in rows 3 and 4. (C) PRT1 (H2879) and prt1-1 (H1676) cells were transformed with p3000 (hc TC) or empty vector and analyzed essentially as in (B). (D) ssu2-1 (F708) cells were transformed with empty vector or p3342 encoding SSU2 (TIF5), and analyzed in rows 1 and 2 as in (B), except using SC-Ura and SC-Ura-Ile-Val+1 μg/μl SM plates and incubating for 2 or 3 days (row 1) and 1 or 2 days (rows 2–4) at 30°C. _SSU2 gcn2_Δ (H2881) and SSU2 GCN2 (H2879) cells, transformed with empty vector, were analyzed in parallel in rows 3 and 4. (E) prt1-1 (H1676) cells were transformed with p3993 encoding SUI5-R31G or p3992 encoding SUI5 (TIF5) and analyzed essentially as in (B), except using SC-Leu plates or SC-Leu-His plates containing 10 mM 3-AT and incubating for 5 (row 1) or 3 days (row 2) at 33°C. _PRT1 gcn2_Δ (H2881) and PRT1 GCN2 (H2879) cells, transformed with empty vector, were analyzed in parallel in rows 3 and 4, incubating for 2 days. (F) tif5-7A GCN2 (YKHN206) cells were transformed with p3927 encoding TIF32_Δ_6–His or empty vector or p3342 encoding TIF5 and analyzed essentially as in (B), except that plates were incubated at 30°C for 2 or 3 days (rows 2–5) and 3 or 4 days (row 1) due to the synthetic growth defect of the tif5-7A TIF32_Δ_6–His strain. TIF5 GCN2 (YKHN205) and _TIF5 gcn2_Δ (H2898), transformed with empty vector, were analyzed in parallel in rows 4 and 5 and incubated for 2 or 3 days.
Figure 6
The prt1-1 mutation impairs GCN4 translational control, leads to hyperaccurate start codon selection, and does not show a synthetic growth defect with depletion of 60S subunit protein RPL11A. (A) prt1-1 (H1676), PRT1 (H2879), YEF3 (F1006) or yef3 (F650S) (F1007) cells were transformed with p180 containing the GCN4–lacZ fusion with all four uORFs and grown in SC-Ura in the presence or absence of 10 mM 3-AT, or 0.06 μg/μl SM, as shown, at the indicated temperatures. In row 1, the prt1-1 strain was also transformed with hc LEU2 plasmid (p832) containing GCN2 (hcGCN2) or empty vector, and the PRT1 strain also harbored an empty vector. β-Galactosidase activities were measured in WCEs and expressed in units of nmol of _o_-nitrophenyl-β-D-galactopyranoside hydrolyzed per min per mg of protein. The mean activities and standard errors obtained from independent transformants are indicated. (B) The prt1-1 (H1676) and PRT1 (H2879) strains were transformed with plasmids p180, pM199, pM226 or p226 (rows 1–4), respectively, and contained the GCN4–lacZ constructs shown schematically to the right and analyzed as in (A). Row 1 contains the same data as in row 4 of (A) shown for comparison. (C) prt1-1 (H1676), PRT1 (H2879), YEF3 (F1006) or yef3 ts (F1007) cells were transformed with p367 or p391 containing a HIS4–lacZ reporter harboring AUG or UUG start codons, respectively. The PRT1 and prt1-1 transformants were grown at 34°C, while the YEF3 and yef3 ts transformants were grown at 35°C, in SC-Ura medium, and β-galactosidase was assayed in WCEs. The graph shows the mean ratios of expression from the UUG to the AUG reporter measured for four independent transformants of each strain (for each reporter), with standard errors indicated as error bars. (D) Growth of prt1-1 (H1676) and _prt1-1 rpl11a_Δ (H2925) strains in YPD medium for 3 days at 25°C (upper panel) or 4 days at 33°C (lower panel).
Figure 7
Redistribution of eIF2 from the 40S-bound to the 40S-free state in the hc-TIF32_Δ_6 tif5-7A mutant (A–D) and schematic model to explain the differing effects of mutations in MFC components on GCN4 translational control (E). (A) WT strain H2898 transformed with empty vector and (C) tif5-7A strain H2899 transformed with hc plasmid p3927 containing TIF32_Δ_6–His were grown in SC-Ura medium and cross-linked with HCHO. WCEs were separated and analyzed as in Figure 2B. (B) Schematic representation of the defective MFC in hc-TIF32_Δ_6 tif5-7A cells lacking the contact between eIF2β and TIF32–CTD and that between eIF2β and NIP1–NTD bridged by eIF5 (see text). (D) The amounts of eIF2γ (eIF2) or PRT1 (eIF3) in fractions 1–4 (top), 5–7 (MFC), and 10–11 (40S) were quantified with a PhosphorImager or by videodensitometry using NIH image 1.63 software, and the resulting values for the hc-TIF32_Δ_6 tif5-7A strain were normalized to the corresponding WT values. The results from four independent experiments were averaged and the mean normalized values and standard errors were plotted, with 100% corresponding to the WT values. The assignment of the MFC to fractions 5–7 was based on our previous analysis of mutant MFC complexes (Valášek et al, 2003). (E) Mutations in MFC components (prt1-1, TIF32_Δ_6 or tif5-7A) that decrease the rate of TC binding to 40S subunits scanning downstream from uORF1 should allow a fraction of 40S subunits to bypass uORF4 and reinitiate at GCN4, even in the absence of eIF2α phosphorylation in _gcn2_Δ cells where TC levels are high (Gcd− phenotype). (F) The rate-limiting defects conferred by mutations in MFC components reduce the rate of scanning by 40S subunits following uORF1 translation. This compensates for the reduced rate of TC recruitment caused by these mutations and thereby suppresses the Gcd− phenotype predicted from the recruitment defects.
Similar articles
- Study of translational control of eukaryotic gene expression using yeast.
Hinnebusch AG, Asano K, Olsen DS, Phan L, Nielsen KH, Valásek L. Hinnebusch AG, et al. Ann N Y Acad Sci. 2004 Dec;1038:60-74. doi: 10.1196/annals.1315.012. Ann N Y Acad Sci. 2004. PMID: 15838098 - Interactions of eukaryotic translation initiation factor 3 (eIF3) subunit NIP1/c with eIF1 and eIF5 promote preinitiation complex assembly and regulate start codon selection.
Valásek L, Nielsen KH, Zhang F, Fekete CA, Hinnebusch AG. Valásek L, et al. Mol Cell Biol. 2004 Nov;24(21):9437-55. doi: 10.1128/MCB.24.21.9437-9455.2004. Mol Cell Biol. 2004. PMID: 15485912 Free PMC article. - A subcomplex of three eIF3 subunits binds eIF1 and eIF5 and stimulates ribosome binding of mRNA and tRNA(i)Met.
Phan L, Schoenfeld LW, Valásek L, Nielsen KH, Hinnebusch AG. Phan L, et al. EMBO J. 2001 Jun 1;20(11):2954-65. doi: 10.1093/emboj/20.11.2954. EMBO J. 2001. PMID: 11387228 Free PMC article. - A multifactor complex of eIF1, eIF2, eIF3, eIF5, and tRNA(i)Met promotes initiation complex assembly and couples GTP hydrolysis to AUG recognition.
Asano K, Phan L, Valásek L, Schoenfeld LW, Shalev A, Clayton J, Nielsen K, Donahue TF, Hinnebusch AG. Asano K, et al. Cold Spring Harb Symp Quant Biol. 2001;66:403-15. doi: 10.1101/sqb.2001.66.403. Cold Spring Harb Symp Quant Biol. 2001. PMID: 12762043 Review. No abstract available. - Gene-specific translational control of the yeast GCN4 gene by phosphorylation of eukaryotic initiation factor 2.
Hinnebusch AG. Hinnebusch AG. Mol Microbiol. 1993 Oct;10(2):215-23. doi: 10.1111/j.1365-2958.1993.tb01947.x. Mol Microbiol. 1993. PMID: 7934812 Review.
Cited by
- Cotranslational assembly of the yeast SET1C histone methyltransferase complex.
Halbach A, Zhang H, Wengi A, Jablonska Z, Gruber IM, Halbeisen RE, Dehé PM, Kemmeren P, Holstege F, Géli V, Gerber AP, Dichtl B. Halbach A, et al. EMBO J. 2009 Oct 7;28(19):2959-70. doi: 10.1038/emboj.2009.240. Epub 2009 Aug 27. EMBO J. 2009. PMID: 19713935 Free PMC article. - Reprogramming mRNA Expression in Response to Defect in RNA Polymerase III Assembly in the Yeast Saccharomyces cerevisiae.
Rudzińska I, Cieśla M, Turowski TW, Armatowska A, Leśniewska E, Boguta M. Rudzińska I, et al. Int J Mol Sci. 2021 Jul 7;22(14):7298. doi: 10.3390/ijms22147298. Int J Mol Sci. 2021. PMID: 34298922 Free PMC article. - eIF4F complex dynamics are important for the activation of the integrated stress response.
Kim KQ, Nanjaraj Urs AN, Lasehinde V, Greenlaw AC, Hudson BH, Zaher HS. Kim KQ, et al. Mol Cell. 2024 Jun 6;84(11):2135-2151.e7. doi: 10.1016/j.molcel.2024.04.016. Mol Cell. 2024. PMID: 38848692 Free PMC article. - Dissociation of eIF1 from the 40S ribosomal subunit is a key step in start codon selection in vivo.
Cheung YN, Maag D, Mitchell SF, Fekete CA, Algire MA, Takacs JE, Shirokikh N, Pestova T, Lorsch JR, Hinnebusch AG. Cheung YN, et al. Genes Dev. 2007 May 15;21(10):1217-30. doi: 10.1101/gad.1528307. Genes Dev. 2007. PMID: 17504939 Free PMC article. - Structural analysis of an eIF3 subcomplex reveals conserved interactions required for a stable and proper translation pre-initiation complex assembly.
Herrmannová A, Daujotyte D, Yang JC, Cuchalová L, Gorrec F, Wagner S, Dányi I, Lukavsky PJ, Valásek LS. Herrmannová A, et al. Nucleic Acids Res. 2012 Mar;40(5):2294-311. doi: 10.1093/nar/gkr765. Epub 2011 Nov 15. Nucleic Acids Res. 2012. PMID: 22090426 Free PMC article.
References
- Anand M, Chakraburtty K, Marton MJ, Hinnebusch AG, Kinzy TG (2003) Functional interactions between yeast translation eukaryotic elongation factor (eEF) 1A and eEF3. J Biol Chem 278: 6985–6991 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous