pH-dependent entry of severe acute respiratory syndrome coronavirus is mediated by the spike glycoprotein and enhanced by dendritic cell transfer through DC-SIGN - PubMed (original) (raw)
pH-dependent entry of severe acute respiratory syndrome coronavirus is mediated by the spike glycoprotein and enhanced by dendritic cell transfer through DC-SIGN
Zhi-Yong Yang et al. J Virol. 2004 Jun.
Abstract
The severe acute respiratory syndrome coronavirus (SARS-CoV) synthesizes several putative viral envelope proteins, including the spike (S), membrane (M), and small envelope (E) glycoproteins. Although these proteins likely are essential for viral replication, their specific roles in SARS-CoV entry have not been defined. In this report, we show that the SARS-CoV S glycoprotein mediates viral entry through pH-dependent endocytosis. Further, we define its cellular tropism and demonstrate that virus transmission occurs through cell-mediated transfer by dendritic cells. The S glycoprotein was used successfully to pseudotype replication-defective retroviral and lentiviral vectors that readily infected Vero cells as well as primary pulmonary and renal epithelial cells from human, nonhuman primate, and, to a lesser extent, feline species. The tropism of this reporter virus was similar to that of wild-type, replication-competent SARS-CoV, and binding of purified S to susceptible target cells was demonstrated by flow cytometry. Although myeloid dendritic cells were able to interact with S and to bind virus, these cells could not be infected by SARS-CoV. However, these cells were able to transfer the virus to susceptible target cells through a synapse-like structure. Both cell-mediated infection and direct infection were inhibited by anti-S antisera, indicating that strategies directed toward this gene product are likely to confer a therapeutic benefit for antiviral drugs or the development of a SARS vaccine.
Figures
FIG. 1.
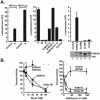
Infection of Vero cells by S-pseudotyped retroviral and lentiviral vectors. Amphotropic (Ampho), S, and no-envelope control vectors were prepared as described in Materials and Methods. Viruses were used at similar multiplicities of infection, standardized by p24 protein levels. Viral pseudotypes were prepared by cotransfection of the indicated combinations of S, M, and E (center). 293T cell supernatants were used to infect Vero cell lines, and luciferase activity was analyzed as previously described (45). (A) (Left) Infection of the Vero cell line with the S-pseudotyped lentiviral or retroviral vector expressing luciferase (42). MLV, murine leukemia virus. (Center) The S glycoprotein, but not the M and E glycoproteins, mediates viral entry by the S-pseudotyped lentiviral vector. (Right) (Top) The requirement for the cytoplasmic domain of S was analyzed by generation of pseudotyped virus using full-length S proteins (S) or S proteins from which the COOH-terminal end was deleted. (Bottom) Expression of these S variants was confirmed by Western blot analysis. (B) pH-dependent entry of SARS-CoV S-pseudotyped lentiviral vectors. Pseudolentiviruses were incubated in the presence of increasing amounts of ammonium chloride (left) or bafilomycin (Sigma) (right). The experiment was performed in triplicate. Data are presented as the percentage of activity at the indicated dose relative to activity with no drug treatment. GP, glycoprotein.
FIG. 2.
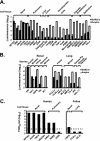
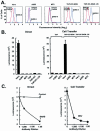
Tropism of a SARS-CoV S-pseudotyped lentiviral vector for human and animal cells and correlation with SARS-CoV infectibility. (A) Tropism of a SARS-CoV S pseudolentivirus for different types of human cells. All infections were performed in triplicate. Data are presented as averages ± standard deviations. Results from one of two independent experiments are shown. (B) Infectibility of renal cells from different species. Cells were infected and analyzed in triplicate. Data are presented as averages ± standard deviations. Results from one of two independent experiments are shown. (C) Infection of selected susceptible and resistant cells from panel A by SARS-CoV strain Urbani, titered as previously described (38) on Vero cells. Dashed line indicates the detection limit of infectivity of SARS-CoV.
FIG. 3.
DC-SIGN-dependent uptake of the SARS-CoV S-pseudotyped lentiviral vector, and cell-mediated transfer and infection of target cells. (A) Binding of purified SARS-CoV S glycoprotein to cell lines. A total of 106 African green monkey kidney cells (Vero), human T-cell leukemia cells (A3R5 and MT2), or THP-1 myelomonocytic leukemia cells expressing wild-type or mutant forms of DC-SIGN (THP-DC-SIGN or THP-DC-SIGNΔ35, respectively) were incubated with purified S(1190)-Myc-His glycoprotein for 20 min on ice. Binding of S protein to the cells was detected by using an FITC-labeled anti-His (COOH-terminal) antibody (blue) (dilution, 1:100; Invitrogen). A FITC-labeled IgG isotype was used as a control (red). Data were analyzed by flow cytometry. (B) Direct viral entry (left) and cell-mediated virus transfer (right) of the SARS-CoVS-pseudotyped lentiviral vector from THP-1, THP-DC-SIGN, and THP-DC-SIGNΔ35 cells. (Left) Susceptibilities of Vero, A3R5, MT2, THP-1, THP-DC-SIGN, and THP-DC-SIGNΔ35 cells to SARS-CoV S-pseudotyped lentiviral vector infection were measured after transduction by use of the luciferase reporter. (Right) Cell-mediated pseudoviral transfer by THP-1, THP-DC-SIGN, or THP-DC-SIGNΔ35 cells (3 × 104) was also assessed by incubating the cells with the SARS-CoV S-pseudotyped lentiviral vector for 2 h at 37°C, followed by three washes before addition of the respective cells to the indicated target Vero cells at a 1:1 ratio. Cells were collected 72 h later for luciferase assay. (C) Inhibition of direct infection and cell-mediated transfer of SARS-CoV S pseudolentivirus by a mouse anti-SARS-CoV S protein antiserum. (Left) The SARS-CoV S-pseudotyped lentiviral vector was exposed to a mouse control or anti-S specific antiserum at the indicated dilutions for 60 min at 37°C before being added to Vero cells. (Right) For cell-mediated transfer, THP-DC-SIGN cells were incubated with pseudoviruses as described in the legend to panel B, followed by incubation with Vero cells in the presence of a control or anti-SARS-CoV S specific mouse antiserum for 48 h. After 48 h, cells were collected for luciferase assays.
FIG. 4.
Uptake and transfer of a GFP-Vpr-labeled SARS-CoV S-pseudotyped lentiviral vector and SARS-CoV by mature human mDC. (A) Uptake of a GFP-Vpr-labeled SARS-CoV S-pseudotyped lentiviral vector by mature mDC and subsequent transfer to renal epithelial cells by mDC, as detected by confocal microscopy. mDC were infected with a GFP-Vpr-labeled SARS-CoV S-pseudotyped lentivirus for 30 min at 37°C and were then added to human renal epithelial cells (786-O; 3 × 104 cells/well, plated 1 day before) in 8-well coverslip slides (Nalge Nunc) at a 1:1 ratio. Uptake, polarization, and transfer were assessed by confocal microscopy with representative cells. Arrow indicates transfer of labeled virus from DC to 786-O cells. (B) Human mature mDC are not directly infected by SARS-CoV (strain Urbani) but instead promote cell-mediated infection of susceptible target cells. (Left) Vero cells, immature mDC, or mature mDC were infected with SARS-CoV (strain Urbani) for 1 h in 96-well-dishes (2 × 104 cells/well), washed three times, and maintained in cell culture medium. (Right) Mature mDC were also infected for 1 h with SARS-CoV, washed, detached with trypsin, and replated onto 96-well-dishes with Vero cells (2 × 104 cells/well; 1:1 ratio) in the presence of a control or anti-SARS-CoV S specific mouse antiserum at a dilution of 1:100. Cell culture supernatants were collected 72 h later, and viral titers were measured as described previously (38). Virus yield is expressed as 50% tissue culture infective doses (TCID50) per milliliter. Dashed line indicates the detection limit of SARS-CoV.
Similar articles
- DC-SIGN and DC-SIGNR interact with the glycoprotein of Marburg virus and the S protein of severe acute respiratory syndrome coronavirus.
Marzi A, Gramberg T, Simmons G, Möller P, Rennekamp AJ, Krumbiegel M, Geier M, Eisemann J, Turza N, Saunier B, Steinkasserer A, Becker S, Bates P, Hofmann H, Pöhlmann S. Marzi A, et al. J Virol. 2004 Nov;78(21):12090-5. doi: 10.1128/JVI.78.21.12090-12095.2004. J Virol. 2004. PMID: 15479853 Free PMC article. - Identifying epitopes responsible for neutralizing antibody and DC-SIGN binding on the spike glycoprotein of the severe acute respiratory syndrome coronavirus.
Shih YP, Chen CY, Liu SJ, Chen KH, Lee YM, Chao YC, Chen YM. Shih YP, et al. J Virol. 2006 Nov;80(21):10315-24. doi: 10.1128/JVI.01138-06. J Virol. 2006. PMID: 17041212 Free PMC article. - Aromatic amino acids in the juxtamembrane domain of severe acute respiratory syndrome coronavirus spike glycoprotein are important for receptor-dependent virus entry and cell-cell fusion.
Howard MW, Travanty EA, Jeffers SA, Smith MK, Wennier ST, Thackray LB, Holmes KV. Howard MW, et al. J Virol. 2008 Mar;82(6):2883-94. doi: 10.1128/JVI.01805-07. Epub 2008 Jan 16. J Virol. 2008. PMID: 18199653 Free PMC article. - The SARS-CoV S glycoprotein.
Xiao X, Dimitrov DS. Xiao X, et al. Cell Mol Life Sci. 2004 Oct;61(19-20):2428-30. doi: 10.1007/s00018-004-4257-y. Cell Mol Life Sci. 2004. PMID: 15526150 Free PMC article. Review. - Hosting the severe acute respiratory syndrome coronavirus: specific cell factors required for infection.
de Haan CA, Rottier PJ. de Haan CA, et al. Cell Microbiol. 2006 Aug;8(8):1211-8. doi: 10.1111/j.1462-5822.2006.00744.x. Epub 2006 Jun 27. Cell Microbiol. 2006. PMID: 16803585 Free PMC article. Review.
Cited by
- COVID-19 Therapeutic Options Under Investigation.
Kaddoura M, AlIbrahim M, Hijazi G, Soudani N, Audi A, Alkalamouni H, Haddad S, Eid A, Zaraket H. Kaddoura M, et al. Front Pharmacol. 2020 Aug 6;11:1196. doi: 10.3389/fphar.2020.01196. eCollection 2020. Front Pharmacol. 2020. PMID: 32848795 Free PMC article. Review. - Immunotherapeutic approaches to curtail COVID-19.
Owji H, Negahdaripour M, Hajighahramani N. Owji H, et al. Int Immunopharmacol. 2020 Nov;88:106924. doi: 10.1016/j.intimp.2020.106924. Epub 2020 Aug 21. Int Immunopharmacol. 2020. PMID: 32877828 Free PMC article. Review. - Possible Therapeutic Use of Natural Compounds Against COVID-19.
Khan N, Chen X, Geiger JD. Khan N, et al. J Cell Signal. 2021;2(1):63-79. J Cell Signal. 2021. PMID: 33768214 Free PMC article. - Leukocyte-specific protein 1 interacts with DC-SIGN and mediates transport of HIV to the proteasome in dendritic cells.
Smith AL, Ganesh L, Leung K, Jongstra-Bilen J, Jongstra J, Nabel GJ. Smith AL, et al. J Exp Med. 2007 Feb 19;204(2):421-30. doi: 10.1084/jem.20061604. Epub 2007 Feb 12. J Exp Med. 2007. PMID: 17296787 Free PMC article. - Modeling the early events of severe acute respiratory syndrome coronavirus infection in vitro.
Yen YT, Liao F, Hsiao CH, Kao CL, Chen YC, Wu-Hsieh BA. Yen YT, et al. J Virol. 2006 Mar;80(6):2684-93. doi: 10.1128/JVI.80.6.2684-2693.2006. J Virol. 2006. PMID: 16501078 Free PMC article.
References
- Abrahamsen, T. G., C. S. Carter, E. J. Read, M. Rubin, H. G. Goetzman, E. F. Lizzio, Y. L. Lee, P. A. Pizzo, and T. Hoffman. 1991. Stimulatory effect of counterflow centrifugal elutriation in large-scale separation of peripheral blood monocytes can be reversed by storing the cells at 37 degrees C. J. Clin. Apheresis 6:48-53. - PubMed
- Bullough, P. A., F. M. Hughson, J. J. Skehel, and D. C. Wiley. 1994. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature 371:37-43. - PubMed
- Burgard, M., M. J. Mayaux, S. Blanche, A. Ferroni, M. L. Guihard-Moscato, M. C. Allemon, N. Ciraru-Vigneron, G. Firtion, C. Floch, F. Guillot, et al. 1992. The use of viral culture and p24 antigen testing to diagnose human immunodeficiency virus infection in neonates. N. Engl. J. Med. 327:1192-1197. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous