Single nucleotide polymorphism-based validation of exonic splicing enhancers - PubMed (original) (raw)
Single nucleotide polymorphism-based validation of exonic splicing enhancers
William G Fairbrother et al. PLoS Biol. 2004 Sep.
Abstract
Because deleterious alleles arising from mutation are filtered by natural selection, mutations that create such alleles will be underrepresented in the set of common genetic variation existing in a population at any given time. Here, we describe an approach based on this idea called VERIFY (variant elimination reinforces functionality), which can be used to assess the extent of natural selection acting on an oligonucleotide motif or set of motifs predicted to have biological activity. As an application of this approach, we analyzed a set of 238 hexanucleotides previously predicted to have exonic splicing enhancer (ESE) activity in human exons using the relative enhancer and silencer classification by unanimous enrichment (RESCUE)-ESE method. Aligning the single nucleotide polymorphisms (SNPs) from the public human SNP database to the chimpanzee genome allowed inference of the direction of the mutations that created present-day SNPs. Analyzing the set of SNPs that overlap RESCUE-ESE hexamers, we conclude that nearly one-fifth of the mutations that disrupt predicted ESEs have been eliminated by natural selection (odds ratio = 0.82 +/- 0.05). This selection is strongest for the predicted ESEs that are located near splice sites. Our results demonstrate a novel approach for quantifying the extent of natural selection acting on candidate functional motifs and also suggest certain features of mutations/SNPs, such as proximity to the splice site and disruption or alteration of predicted ESEs, that should be useful in identifying variants that might cause a biological phenotype.
Conflict of interest statement
The authors have declared that no conflicts of interest exist.
Figures
Figure 1. Density of Predicted ESEs and SNPs along Human Exons
RESCUE-ESE hexamers were searched against a database of 121,000 internal human exons. ESE density (blue curve) was determined as the fraction of hexamers beginning at the given exon position in this dataset that were contained in the RESCUE-ESE set. SNP density (red curve) was determined analogously using SNPs from dbSNP mapped to the exon database. Both curves were smoothed by averaging the densities over a leading (3′ss) or lagging (5′ss) window of ten nucleotides.
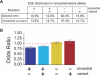
Figure 2. Analysis of the Effects of SNPs and Unselected Mutations on Predicted ESEs
(A) The percentages of the four prediction outcomes. ESE disruption (+ −), ESE alteration (+ +), ESE neutrality (− −), and ESE creation (− +) changes are listed for the set of 2,561 validated SNPs (selected) and for the set of 100,000 simulated (unselected) mutations. (B) Synonymous and nonsynonymous mutations were analyzed separately and then compared using the MH test for homogeneity. All outcomes passed the MH test for homogeneity (H0:Outcomesynon ≈ Outcomenonsynon; p < 0.05) and could, therefore, be combined into a summary OR (weighted combination of the ORs measured in the synonymous and nonsynonymous sets). The height of each bar can be interpreted as the odds that the listed outcome will occur in the evolutionarily selected set of mutations (SNPs) relative to the odds that the same outcome will occur in the unselected (simulated mutation) set. Error bars extend one standard deviation on either side of the calculated value.
Figure 3. Selection against Disruption of Predicted ESEs in Different Exon Regions
Summary ORs were calculated for mutations that disrupt RESCUE-ESEs as in Figure 2, for each of four regions spanning the length of a typical human internal exon. The heights of the blue bars represent the odds that an ESE will be disrupted by a mutation in the set of 2,561 validated SNPs (selected mutations) relative to the odds of disruption in the set of 100,000 simulated (unselected) mutations. Error bars extend one standard deviation on either side of the calculated value.
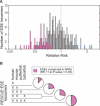
Figure 4. Measuring Selective Pressure on Each RESCUE-ESE Hexamer
Any point mutation alters six overlapping hexamers, and so a database of 8,408 SNP mutations alters a total of approximately 50,000 hexamers in the wild-type (ancestral) allele. In considering all 238 RESCUE-ESE hexamers, the frequency of each ESE hexamer in the total set of ancestral alleles was recorded for the database of SNPs and simulated mutations (8,408 SNP mutations and 100,000 simulated mutations). The ESE frequency in the SNP set was divided by the ESE frequency in the simulated set to calculate the RR for each of the 238 hexamers. (A) The distribution of RR for all 238 ESE hexamers is plotted on a logarithmic scale. A resampling strategy was used to identify 57 ESE hexamers that were significantly conserved (pink bars have an RR less than 1; p < 0.05) and also six ESE hexamers that were not conserved (blue bars have an RR greater than 1; p < 0.05). (B) The output of RESCUE-ESE was compared for several vertebrate genomes (human, mouse, pufferfish, and zebrafish). The set of 238 human RESCUE-ESE hexamers was divided into nonoverlapping subsets based on their conservation in the RESCUE-ESE output generated from other vertebrates. The proportion of ESEs that were significantly conserved in the SNP analysis (as described above in [A]) were recorded for each subset of RESCUE-ESE hexamers and are represented as pink sectors in the pie chart.
Similar articles
- Synonymous SNPs provide evidence for selective constraint on human exonic splicing enhancers.
Carlini DB, Genut JE. Carlini DB, et al. J Mol Evol. 2006 Jan;62(1):89-98. doi: 10.1007/s00239-005-0055-x. Epub 2005 Nov 30. J Mol Evol. 2006. PMID: 16320116 - Missense mutations in cancer suppressor gene TP53 are colocalized with exonic splicing enhancers (ESEs).
Gorlov IP, Gorlova OY, Frazier ML, Amos CI. Gorlov IP, et al. Mutat Res. 2004 Oct 4;554(1-2):175-83. doi: 10.1016/j.mrfmmm.2004.04.014. Mutat Res. 2004. PMID: 15450416 - Why Selection Might Be Stronger When Populations Are Small: Intron Size and Density Predict within and between-Species Usage of Exonic Splice Associated cis-Motifs.
Wu X, Hurst LD. Wu X, et al. Mol Biol Evol. 2015 Jul;32(7):1847-61. doi: 10.1093/molbev/msv069. Epub 2015 Mar 13. Mol Biol Evol. 2015. PMID: 25771198 Free PMC article. - Exonic splicing enhancers: mechanism of action, diversity and role in human genetic diseases.
Blencowe BJ. Blencowe BJ. Trends Biochem Sci. 2000 Mar;25(3):106-10. doi: 10.1016/s0968-0004(00)01549-8. Trends Biochem Sci. 2000. PMID: 10694877 Review. - [Pathogenic mutation or polymorphism? (How to find criteria)].
Kochański A. Kochański A. Med Wieku Rozwoj. 2006 Jan-Mar;10(1 Pt 2):335-41. Med Wieku Rozwoj. 2006. PMID: 17028397 Review. Polish.
Cited by
- Intronic splicing enhancers, cognate splicing factors and context-dependent regulation rules.
Wang Y, Ma M, Xiao X, Wang Z. Wang Y, et al. Nat Struct Mol Biol. 2012 Oct;19(10):1044-52. doi: 10.1038/nsmb.2377. Epub 2012 Sep 16. Nat Struct Mol Biol. 2012. PMID: 22983564 Free PMC article. - The etiology of Down syndrome: Maternal MCM9 polymorphisms increase risk of reduced recombination and nondisjunction of chromosome 21 during meiosis I within oocyte.
Pal U, Halder P, Ray A, Sarkar S, Datta S, Ghosh P, Ghosh S. Pal U, et al. PLoS Genet. 2021 Mar 22;17(3):e1009462. doi: 10.1371/journal.pgen.1009462. eCollection 2021 Mar. PLoS Genet. 2021. PMID: 33750944 Free PMC article. - Genome-wide analysis reveals the association between alternative splicing and DNA methylation across human solid tumors.
Sun X, Tian Y, Wang J, Sun Z, Zhu Y. Sun X, et al. BMC Med Genomics. 2020 Jan 6;13(1):4. doi: 10.1186/s12920-019-0654-9. BMC Med Genomics. 2020. PMID: 31906954 Free PMC article. - CYP2B6 non-coding variation associated with smoking cessation is also associated with differences in allelic expression, splicing, and nicotine metabolism independent of common amino-acid changes.
Bloom AJ, Martinez M, Chen LS, Bierut LJ, Murphy SE, Goate A. Bloom AJ, et al. PLoS One. 2013 Nov 15;8(11):e79700. doi: 10.1371/journal.pone.0079700. eCollection 2013. PLoS One. 2013. PMID: 24260284 Free PMC article. - Impairment of translation in neurons as a putative causative factor for autism.
Poliakov E, Koonin EV, Rogozin IB. Poliakov E, et al. Biol Direct. 2014 Jul 10;9:16. doi: 10.1186/1745-6150-9-16. Biol Direct. 2014. PMID: 25011470 Free PMC article.
References
- Altshuler D, Pollara VJ, Cowles CR, Van Etten WJ, Baldwin J, et al. An SNP map of the human genome generated by reduced representation shotgun sequencing. Nature. 2000;407:513–516. - PubMed
- Bailey JA, Gu Z, Clark RA, Reinert K, Samonte RV, et al. Recent segmental duplications in the human genome. Science. 2002;297:1003–1007. - PubMed
- Betticher DC, Thatcher N, Altermatt HJ, Hoban P, Ryder WD, et al. Alternate splicing produces a novel cyclin D1 transcript. Oncogene. 1995;11:1005–1011. - PubMed
- Blencowe BJ. Exonic splicing enhancers: Mechanism of action, diversity and role in human genetic diseases. Trends Biochem Sci. 2000;25:106–110. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials