Lactate-utilizing bacteria, isolated from human feces, that produce butyrate as a major fermentation product - PubMed (original) (raw)
Lactate-utilizing bacteria, isolated from human feces, that produce butyrate as a major fermentation product
Sylvia H Duncan et al. Appl Environ Microbiol. 2004 Oct.
Abstract
The microbial community of the human colon contains many bacteria that produce lactic acid, but lactate is normally detected only at low concentrations (<5 mM) in feces from healthy individuals. It is not clear, however, which bacteria are mainly responsible for lactate utilization in the human colon. Here, bacteria able to utilize lactate and produce butyrate were identified among isolates obtained from 10(-8) dilutions of fecal samples from five different subjects. Out of nine such strains identified, four were found to be related to Eubacterium hallii and two to Anaerostipes caccae, while the remaining three represent a new species within clostridial cluster XIVa based on their 16S rRNA sequences. Significant ability to utilize lactate was not detected in the butyrate-producing species Roseburia intestinalis, Eubacterium rectale, or Faecalibacterium prausnitzii. Whereas E. hallii and A. caccae strains used both D- and L-lactate, the remaining strains used only the d form. Addition of glucose to batch cultures prevented lactate utilization until the glucose became exhausted. However, when two E. hallii strains and one A. caccae strain were grown in separate cocultures with a starch-utilizing Bifidobacterium adolescentis isolate, with starch as the carbohydrate energy source, the L-lactate produced by B. adolescentis became undetectable and butyrate was formed. Such cross-feeding may help to explain the reported butyrogenic effect of certain dietary substrates, including resistant starch. The abundance of E. hallii in particular in the colonic ecosystem suggests that these bacteria play important roles in preventing lactate accumulation.
Figures
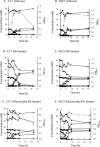
FIG. 1.
Time course of SCFA formation and growth in batch culture of E. hallii L2-7 (A, B, C) of the _C. indolis_-related strain SS2/1 (D, E, F) on media containing
dl
-lactate, glucose, or
dl
-lactate plus glucose. Millimolar concentrations of substrates utilized and products formed are shown for glucose (⧫),
l
-lactate (▪), formate (▴), acetate (×), butyrate ( ), and
), and
dl
-lactate (○). Growth (+) is measured as optical density at 650 nm (OD650). All values are the means of results from triplicate cultures. Standard deviations (not shown) were typically around 5% of the mean value.
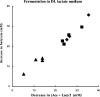
FIG. 2.
Relationship between butyrate formation and net removal of acetate and lactate from the medium during 24 h of growth on
dl
-lactate medium. Data are largely from Tables 2 and 3 for E. hallii strains L2-7, SM6/1 and SL6/1/1 (▪), A. caccae strains L1-92 and P2 (⧫), and _C. indolis_-related strains SSC/2 and SS2/1 (▴). Values from two independent experiments are included for L2-7, L1-92, SSC/2, and SS2/1.
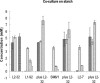
FIG. 3.
Coculture between lactate-utilizing bacteria and B. adolescentis L2-32. The concentrations of SCFA, formate ( ), acetate (
), acetate ( ), butyrate (
), butyrate ( ), and
), and
l
-lactate ( ) are shown after 24 h of growth in YCFA medium with 0.2% starch as an energy source (values for acetate, which was initially present at 33 mM in the medium in all cases, are shown on a 10-fold reduced scale). Butyrate production by A. caccae L1-92 and by E. hallii L2-7 and SM6/1 was stimulated by coculture with B. adolescentis L2-32, while
) are shown after 24 h of growth in YCFA medium with 0.2% starch as an energy source (values for acetate, which was initially present at 33 mM in the medium in all cases, are shown on a 10-fold reduced scale). Butyrate production by A. caccae L1-92 and by E. hallii L2-7 and SM6/1 was stimulated by coculture with B. adolescentis L2-32, while
l
-lactate disappeared from the cocultures. Means and standard deviations are shown based on results from triplicate cultures.
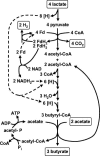
FIG. 4.
Scheme for butyrate formation from lactate in E. hallii and A. caccae.
References
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
- Atlas, R. M. 1997. Selenomonas selective medium (SS medium), p. 1245. In L. C. Parks (ed.), Handbook of microbiological media, 2nd ed. CRC Press, Cleveland, Ohio.
- Barcenilla, A. 1999. Diversity of the butyrate-producing microflora of the human gut. Ph.D. thesis. The Robert Gordon University, Aberdeen, United Kingdom.
- Benno, Y., K. Endo, H. Miyoshi, T. Okuda, H. Koishi, and T. Mitsuoka. 1989. Effect of rice fiber on human fecal microflora. Microbiol. Immunol. 33:435-440. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical