Support for the homeobox transcription factor gene ENGRAILED 2 as an autism spectrum disorder susceptibility locus - PubMed (original) (raw)
doi: 10.1086/497705. Epub 2005 Oct 5.
Neda Gharani, Ian Rossman, Vincent Mancuso, Gloria Lazar, Silky Kamdar, Shannon E Bruse, Samuel Tischfield, Brett J Smith, Raymond A Zimmerman, Emanuel Dicicco-Bloom, Linda M Brzustowicz, James H Millonig
Affiliations
- PMID: 16252243
- PMCID: PMC1271392
- DOI: 10.1086/497705
Support for the homeobox transcription factor gene ENGRAILED 2 as an autism spectrum disorder susceptibility locus
Rym Benayed et al. Am J Hum Genet. 2005 Nov.
Abstract
Our previous research involving 167 nuclear families from the Autism Genetic Resource Exchange (AGRE) demonstrated that two intronic SNPs, rs1861972 and rs1861973, in the homeodomain transcription factor gene ENGRAILED 2 (EN2) are significantly associated with autism spectrum disorder (ASD). In this study, significant replication of association for rs1861972 and rs1861973 is reported for two additional data sets: an independent set of 222 AGRE families (rs1861972-rs1861973 haplotype, P=.0016) and a separate sample of 129 National Institutes of Mental Health families (rs1861972-rs1861973 haplotype, P=.0431). Association analysis of the haplotype in the combined sample of both AGRE data sets (389 families) produced a P value of .0000033, whereas combining all three data sets (518 families) produced a P value of .00000035. Population-attributable risk calculations for the associated haplotype, performed using the entire sample of 518 families, determined that the risk allele contributes to as many as 40% of ASD cases in the general population. Linkage disequilibrium (LD) mapping with the use of polymorphisms distributed throughout the gene has shown that only intronic SNPs are in strong LD with rs1861972 and rs1861973. Resequencing and association analysis of all intronic SNPs have identified alleles associated with ASD, which makes them candidates for future functional analysis. Finally, to begin defining the function of EN2 during development, mouse En2 was ectopically expressed in cortical precursors. Fewer En2-transfected cells than controls displayed a differentiated phenotype. Together, these data provide further genetic evidence that EN2 might act as an ASD susceptibility locus, and they suggest that a risk allele that perturbs the spatial/temporal expression of EN2 could significantly alter normal brain development.
Figures
Figure 1
Schematic representation of the human EN2 gene, with the location, name, and MAF of, as well as the genotyping method used for, the polymorphisms tested for association with ASD.
Figure 2
Intermarker LD values. A, Tabular representation of the pairwise LD (_D_′) values between the 18 polymorphisms tested for association in the AGRE I data set. The LD relationships of each polymorphism with rs1861972 and rs1861973 are highlighted, with the degree of shading corresponding to the strength of LD: black indicates strong LD (_D_′ > 0.72); gray indicates intermediate/weak LD (_D_′ range 0.024–0.632). B, The EN2 gene is illustrated with the position of all 18 tested polymorphisms demarcated by arrows. The decay of LD of each polymorphism with respect to rs1861972 and rs1861973 is represented by a gradient bar drawn below the gene. The degree of shading corresponds to the strength of LD, as described above. Only intronic polymorphisms are in strong LD with rs1861972 and rs1861973.
Figure 3
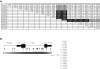
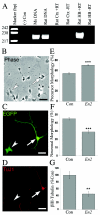
Effects of En2 misexpression on neurogenesis. A, No En2 expression in rat cerebral cortex. En2 expression was detected by RT-PCR of RNAs obtained from freshly dissected E14.5 rat cortex (Ctx) and hindbrain (HB). The primers amplified the same 220-bp region of the En2 3′ UTR from both genomic mouse (Ms) and rat DNAs, consistent with the high degree of nucleotide homology. No expression was found in rat cortex, whereas a high level was detected in the hindbrain, recapitulating the known expression pattern of En2. +/− RT = with/without reverse transcriptase; Con = control. B–D, Transfected cortical cells exhibiting morphologies of undifferentiated precursors and differentiated neurons. The arrowheads denote cells with precursor morphology, whereas the arrows identify cells with neuronal morphology. Magnification 40×; scale bar = 30 μm. B, Phase photomicrograph demonstrating cells with both precursor and neuronal morphologies. Note the large, flat cell bodies and the thick, tapering, and short processes of the precursors (arrowhead). In contrast, neuron-like cells exhibit round cell bodies with long, thin, and uniform-diameter processes (arrow). C, Examples of both precursors and neurons transfected with the En2 vectors and identified for gene expression by immunostaining for EGFP marker protein. D, Cultures that were double labeled to detect EGFP and neuron-specific cytoskeletal protein βIII-tubulin (TuJ1), an early marker of neuronal differentiation. TuJ1 expression was only detected in cells exhibiting neuronal morphologies, a finding consistent with our previous studies (Nicot and DiCicco-Bloom 2001). E, Increase in the proportion of GFP+ precursor cells, noted 24 h after En2 transfection. F, Corresponding decrease in the proportion of _En2_-transfected cells exhibiting neuronal morphology. G, En2 transfection elicited a >50% reduction in TuJ1-immunoreactive cells, a change that parallels the decrease in neuronal morphology. **
P<.0005
,
_N_=9
; ***
P<4×10-8
,
_N_=15
.
Similar articles
- Autism-associated haplotype affects the regulation of the homeobox gene, ENGRAILED 2.
Benayed R, Choi J, Matteson PG, Gharani N, Kamdar S, Brzustowicz LM, Millonig JH. Benayed R, et al. Biol Psychiatry. 2009 Nov 15;66(10):911-7. doi: 10.1016/j.biopsych.2009.05.027. Epub 2009 Jul 17. Biol Psychiatry. 2009. PMID: 19615670 Free PMC article. - Association of the homeobox transcription factor, ENGRAILED 2, 3, with autism spectrum disorder.
Gharani N, Benayed R, Mancuso V, Brzustowicz LM, Millonig JH. Gharani N, et al. Mol Psychiatry. 2004 May;9(5):474-84. doi: 10.1038/sj.mp.4001498. Mol Psychiatry. 2004. PMID: 15024396 - Intronic single nucleotide polymorphisms of engrailed homeobox 2 modulate the disease vulnerability of autism in a han chinese population.
Yang P, Shu BC, Hallmayer JF, Lung FW. Yang P, et al. Neuropsychobiology. 2010;62(2):104-15. doi: 10.1159/000315441. Epub 2010 Jun 3. Neuropsychobiology. 2010. PMID: 20523082 - Autism associated gene, engrailed2, and flanking gene levels are altered in post-mortem cerebellum.
Choi J, Ababon MR, Soliman M, Lin Y, Brzustowicz LM, Matteson PG, Millonig JH. Choi J, et al. PLoS One. 2014 Feb 10;9(2):e87208. doi: 10.1371/journal.pone.0087208. eCollection 2014. PLoS One. 2014. PMID: 24520327 Free PMC article. - EN2 in Prostate Cancer.
McGrath SE, Michael A, Morgan R, Pandha H. McGrath SE, et al. Adv Clin Chem. 2015;71:47-76. doi: 10.1016/bs.acc.2015.06.002. Epub 2015 Jul 21. Adv Clin Chem. 2015. PMID: 26411411 Review.
Cited by
- GABAergic System Dysfunction in Autism Spectrum Disorders.
Zhao H, Mao X, Zhu C, Zou X, Peng F, Yang W, Li B, Li G, Ge T, Cui R. Zhao H, et al. Front Cell Dev Biol. 2022 Feb 7;9:781327. doi: 10.3389/fcell.2021.781327. eCollection 2021. Front Cell Dev Biol. 2022. PMID: 35198562 Free PMC article. Review. - Implication of Hippocampal Neurogenesis in Autism Spectrum Disorder: Pathogenesis and Therapeutic Implications.
Liu C, Liu J, Gong H, Liu T, Li X, Fan X. Liu C, et al. Curr Neuropharmacol. 2023;21(11):2266-2282. doi: 10.2174/1570159X21666221220155455. Curr Neuropharmacol. 2023. PMID: 36545727 Free PMC article. Review. - The Wnt Signaling Pathway and Related Therapeutic Drugs in Autism Spectrum Disorder.
Bae SM, Hong JY. Bae SM, et al. Clin Psychopharmacol Neurosci. 2018 May 31;16(2):129-135. doi: 10.9758/cpn.2018.16.2.129. Clin Psychopharmacol Neurosci. 2018. PMID: 29739125 Free PMC article. Review. - NEuronMOrphological analysis tool: open-source software for quantitative morphometrics.
Billeci L, Magliaro C, Pioggia G, Ahluwalia A. Billeci L, et al. Front Neuroinform. 2013 Feb 14;7:2. doi: 10.3389/fninf.2013.00002. eCollection 2013. Front Neuroinform. 2013. PMID: 23420185 Free PMC article. - Clinical and Neurobiological Relevance of Current Animal Models of Autism Spectrum Disorders.
Kim KC, Gonzales EL, Lázaro MT, Choi CS, Bahn GH, Yoo HJ, Shin CY. Kim KC, et al. Biomol Ther (Seoul). 2016 May 1;24(3):207-43. doi: 10.4062/biomolther.2016.061. Biomol Ther (Seoul). 2016. PMID: 27133257 Free PMC article. Review.
References
Web Resources
- Autism Genetic Resource Exchange (AGRE), http://www.agre.org/
- National Institute of Mental Health (NIMH) Data Set, http://zork.wustl.edu/nimh/NIMH_initiative/NIMH_initiative_link.html
- NIMH Center for Collaborative Genetic Studies, http://zork.wustl.edu/nimh/home/d_autism.html
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for ASD and EN2) - PubMed
References
- Abecasis GR, Cookson WO (2000) GOLD—graphical overview of linkage disequilibrium. Bioinformatics 16:182–183 - PubMed
- Ahmadian A, Gharizadeh B, Gustafsson AC, Sterky F, Nyren P, Uhlen M, Lundeberg J (2000) Single nucleotide polymorphism analysis by pyrosequencing. Anal Biochem 280:103–110 - PubMed
- Akshoomoff NA, Courchesne E, Townsend J (1997) Attention coordination and anticipatory control. Int Rev Neurobiol 41:575–598 - PubMed
- Allen G, Buxton RB, Wong EC, Courchesne E (1997) Attentional activation of the cerebellum independent of motor involvement. Science 275:1940–1943 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 MH70366/MH/NIMH NIH HHS/United States
- P30 ES005022/ES/NIEHS NIH HHS/United States
- MH52708/MH/NIMH NIH HHS/United States
- P01 ES11256/ES/NIEHS NIH HHS/United States
- 1 F30 NS48649-01/NS/NINDS NIH HHS/United States
- P01 ES011256/ES/NIEHS NIH HHS/United States
- MH00219/MH/NIMH NIH HHS/United States
- F30 NS048649/NS/NINDS NIH HHS/United States
- R01 MH070366/MH/NIMH NIH HHS/United States
- U24 MH068457/MH/NIMH NIH HHS/United States
- T32 MH019957/MH/NIMH NIH HHS/United States
- MH39437/MH/NIMH NIH HHS/United States
- MH068457/MH/NIMH NIH HHS/United States
- MH/AG-19957/MH/NIMH NIH HHS/United States
- MH00980/MH/NIMH NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials