Fanconi anemia proteins are required to prevent accumulation of replication-associated DNA double-strand breaks - PubMed (original) (raw)
doi: 10.1128/MCB.26.2.425-437.2006.
Stacie Stone, Vincenzo Costanzo, Bendert de Graaf, Tanja Reuter, Johan de Winter, Michael Wallisch, Yassmine Akkari, Susan Olson, Weidong Wang, Hans Joenje, Jan L Christian, Patrick J Lupardus, Karlene A Cimprich, Jean Gautier, Maureen E Hoatlin
Affiliations
- PMID: 16382135
- PMCID: PMC1346898
- DOI: 10.1128/MCB.26.2.425-437.2006
Fanconi anemia proteins are required to prevent accumulation of replication-associated DNA double-strand breaks
Alexandra Sobeck et al. Mol Cell Biol. 2006 Jan.
Abstract
Fanconi anemia (FA) is a multigene cancer susceptibility disorder characterized by cellular hypersensitivity to DNA interstrand cross-linking agents such as mitomycin C (MMC). FA proteins are suspected to function at the interface between cell cycle checkpoints, DNA repair, and DNA replication. Using replicating extracts from Xenopus eggs, we developed cell-free assays for FA proteins (xFA). Recruitment of the xFA core complex and xFANCD2 to chromatin is strictly dependent on replication initiation, even in the presence of MMC indicating specific recruitment to DNA lesions encountered by the replication machinery. The increase in xFA chromatin binding following treatment with MMC is part of a caffeine-sensitive S-phase checkpoint that is controlled by xATR. Recruitment of xFANCD2, but not xFANCA, is dependent on the xATR-xATR-interacting protein (xATRIP) complex. Immunodepletion of either xFANCA or xFANCD2 from egg extracts results in accumulation of chromosomal DNA breaks during replicative synthesis. Our results suggest coordinated chromatin recruitment of xFA proteins in response to replication-associated DNA lesions and indicate that xFA proteins function to prevent the accumulation of DNA breaks that arise during unperturbed replication.
Figures
FIG. 1.
Homology between Xenopus and human FANCD2. Full-length Xenopus FANCD2 (xFANCD2) was aligned with full-length human FANCD2 (hFANCD2) with MacVector 7.1.1 software. Amino acid identity is indicated by dark shading; similar amino acids are indicated by light shading. Two known critical residues, S222 and K561 (indicated by arrows), are conserved between human and Xenopus FANCD2.
FIG. 2.
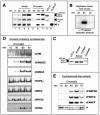
xFA proteins accumulate in nuclei and associate with chromatin in a replication-dependent manner. (A) Nuclear accumulation and chromatin binding of FA proteins during replication. Sperm chromatin was added to S-phase egg extracts and reisolated at indicated time points during replication. Nuclear or chromatin fractions were analyzed by SDS-PAGE and immunoblotting. Xenopus XTC-2 cell lysates (see the supplementary material) were used as a control for xFANCD2-S (short form), xFANCD2-L (long form), and xFANCA; a lysate containing Flag-tagged xFANCF (overexpressed in 293 EBNA cells) was used as a positive control. The suffix “-chr” indicates the chromatin-bound protein isoforms from egg extracts as described in the text. (B) Replication assay. Throughout the experimental procedure described in the legend to panel A, semiconservative replication was monitored by pulsing replicating extract aliquots with [α-32P]GTP at time windows 0 to 30, 30 to 60, and 60 to 90 min. (C) Isoforms of xFANCD2 detectable in M and S phase and in chromatin fractions from replicating nuclei. An isoform of xFANCD2 with relatively slower mobility associates with chromatin during replication. Electrophoretic mobility was compared between xFANCD2 isolated from XTC-2 cell lysates, DNA-free mitotic egg extracts (M phase), and DNA-free S-phase egg extracts (S phase) and from nuclear sperm chromatin replicating in S-phase egg extracts (chromatin). (D) Comparison of chromatin binding patterns between xFA proteins and other DNA replication-associated proteins. Sperm chromatin was added to cycloheximide-containing S-phase egg extracts, and chromatin fractions were reisolated at the times shown and analyzed for the indicated proteins by immunoblotting. (E) xFA proteins dissociate from chromatin in nonarrested extracts following replication. Sperm chromatin was added to S-phase egg extracts in the absence of cycloheximide. Chromatin fractions were reisolated at the indicated time points and analyzed for xFANCA, xFANCD2, and xFANCF by immunoblotting. A nonspecific band indicated by an asterisk was used as a loading control.
FIG.3.
Chromatin recruitment of xFA proteins in response to replication fork stalling and checkpoint activation. (A, top left) S-phase extracts were treated as indicated and supplemented with sperm chromatin. Following replication (60 min), chromatin was reisolated and the same blot was assayed in series for the presence of the xFA proteins indicated. A nonspecific band, indicated by an asterisk, was used as a loading control. (Top right) MMC-treated extracts activate a caffeine-sensitive S-phase checkpoint. A replication assay was performed to monitor incorporation of [α-32P]GTP during one round of replication in extracts treated as indicated. Total amounts of replication products at 60 min following addition of sperm chromatin to nontreated or treated extracts were analyzed by agarose gel electrophoresis and exposure to X-ray film. Band density percentages were determined using a Bio-Rad Molecular PhosphorImager Fx (Kodak screens, Quantity-1 software). (Bottom) Rescue of dose-dependent, MMC-induced replication reduction by neutralizing xATR. A total of 1,000 sperm pronuclei/μl were added to replicating extracts containing different concentrations of MMC (lanes 3 to 9). Extracts were otherwise untreated or contained caffeine (lanes 6 and 7) or a neutralizing anti-xATR antibody (lanes 8 and 9). The MMC concentration was 150 μM (lanes 3, 6, and 8), 50 μM (lanes 4, 7, and 9), or 5 μM (lane 5). (B) Stalling of replication forks induces recruitment of xFA proteins to chromatin. Sperm chromatin was added to S-phase egg extracts. Extracts were either untreated (−) or aphidicolin was added (+) at the indicated time points following addition of sperm DNA: 0 min, prenuclear membrane formation; 30 min, beginning of replication; 60 min, midreplication; 90 min, late or postreplication; and 120 min, postreplication. All chromatin fractions were reisolated at 135 min following the addition of sperm DNA, analyzed by SDS-PAGE, and immunoblotted as indicated. (C) Chromatin binding of xFANCD2 is xATRIP dependent. Egg extracts were treated with beads coupled to preimmune serum (lanes 2, 3, and 5) or beads coupled to anti-xATRIP serum (lanes 1, 4, and 6). Sperm chromatin was added to preimmune- or xATRIP-depleted extracts and allowed to replicate. Extracts were either untreated or supplemented with aphidicolin (added 50 min after the addition of sperm DNA). Chromatin fractions were reisolated at 70 min and assayed by immunoblotting for the presence of xFANCD2, xFANCA, xATRIP, and xORC-2. (D) Chromatin binding of xATRIP is xFANCA independent. Egg extracts were treated with beads coupled to preimmune serum (lanes 1 and 3) or beads coupled to anti-xFANCA serum (lanes 2 and 4). Sperm chromatin was added to preimmune- or xFANCA-depleted extracts and allowed to replicate. Extracts were either untreated or supplemented with aphidicolin (added 50 min after the addition of sperm DNA). Chromatin fractions were reisolated at 70 min and assayed for the presence of xFANCA, xATRIP, and xORC-2 by immunoblotting.
FIG. 4.
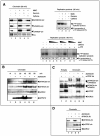
Replication-associated chromatin binding of xFANCD2 requires the presence of xFANCA. (A) S-phase egg extracts were incubated either with bead-coupled anti-xFANCA antibody (xA) or with bead-coupled preimmune serum (Pre) and assayed for presence of xFANCA by immunoblotting. Right lane, xFANCA binds to anti-xFANCA beads during the depletion process. (B) Sperm chromatin was added to xA- or predepleted extracts, reisolated following replication, and assayed for the presence of xFANCD2-chr by immunoblotting. (C and D) Replication-associated chromatin binding of xFANCA does not require the presence of xFANCD2. (C) S-phase egg extracts were incubated either with bead-coupled anti-xFANCD2 antibody (xD2) or with bead-coupled preimmune serum (Pre) and assayed for presence of xFANCD2 by immunoblotting. Right lane, xFANCD2 binds to anti-xFANCD2-beads during the depletion process. (D) Sperm chromatin was added to xD2- or predepleted extracts, reisolated following replication, and assayed for the presence of xFANCA-chr by immunoblotting.
FIG. 5.
xFA proteins are required to prevent accumulation of DNA breaks during unperturbed replication. (A and B) xFANCA and xFANCD2 are not essential for DNA replication. Genomic DNA replication was monitored by 30-min pulses of [α-32P]GTP. (A) Nucleotide incorporation into replicating chromatin in extracts incubated with protein A beads bound to preimmune serum (xA preimmune serum-depleted extracts) or extract incubated with protein A beads bound to anti-xFANCA antibodies (xA-depleted extracts). (B) Incorporation into replicating chromatin in extracts incubated with protein A beads bound to preimmune serum (xD2 preimmune serum-depleted extracts) or extract incubated with protein A beads bound to anti-xFANCD2 antibodies (xD2-depleted extracts). (C and D) Charted averages (including error bars) of the incorporation of radiolabeled nucleotides by band density from three identical replication assays (indicated) including the assays from panels A and B. (E and F) Depletion of xFANCA or xFANCD2 causes DNA breaks during normal replication. (E) Egg extracts were treated with beads coupled to xFANCA-preimmune serum (lane 1) or beads coupled to anti-xFANCA serum (lane 2). (F) Likewise, extracts were treated with either beads coupled to xFANCD2 preimmune serum (lane 1) or beads coupled to anti-xFANCD2 serum (lane 2). Sperm chromatin was added to treated extracts; after 120 min, TUNEL assays were performed. Samples were subjected to agarose gel electrophoresis and phosphorimaging.
Similar articles
- DNA structure-induced recruitment and activation of the Fanconi anemia pathway protein FANCD2.
Sobeck A, Stone S, Hoatlin ME. Sobeck A, et al. Mol Cell Biol. 2007 Jun;27(12):4283-92. doi: 10.1128/MCB.02196-06. Epub 2007 Apr 9. Mol Cell Biol. 2007. PMID: 17420278 Free PMC article. - The Fanconi anemia protein FANCM is controlled by FANCD2 and the ATR/ATM pathways.
Sobeck A, Stone S, Landais I, de Graaf B, Hoatlin ME. Sobeck A, et al. J Biol Chem. 2009 Sep 18;284(38):25560-8. doi: 10.1074/jbc.M109.007690. Epub 2009 Jul 24. J Biol Chem. 2009. PMID: 19633289 Free PMC article. - Fanconi anemia proteins stabilize replication forks.
Wang LC, Stone S, Hoatlin ME, Gautier J. Wang LC, et al. DNA Repair (Amst). 2008 Dec 1;7(12):1973-81. doi: 10.1016/j.dnarep.2008.08.005. Epub 2008 Sep 25. DNA Repair (Amst). 2008. PMID: 18786657 Free PMC article. - The interplay of Fanconi anemia proteins in the DNA damage response.
Wang X, D'Andrea AD. Wang X, et al. DNA Repair (Amst). 2004 Aug-Sep;3(8-9):1063-9. doi: 10.1016/j.dnarep.2004.04.005. DNA Repair (Amst). 2004. PMID: 15279794 Review. - The Fanconi anemia ID2 complex: dueling saxes at the crossroads.
Boisvert RA, Howlett NG. Boisvert RA, et al. Cell Cycle. 2014;13(19):2999-3015. doi: 10.4161/15384101.2014.956475. Cell Cycle. 2014. PMID: 25486561 Free PMC article. Review.
Cited by
- Fanconi DNA repair pathway is required for survival and long-term maintenance of neural progenitors.
Sii-Felice K, Etienne O, Hoffschir F, Mathieu C, Riou L, Barroca V, Haton C, Arwert F, Fouchet P, Boussin FD, Mouthon MA. Sii-Felice K, et al. EMBO J. 2008 Mar 5;27(5):770-81. doi: 10.1038/emboj.2008.14. Epub 2008 Jan 31. EMBO J. 2008. PMID: 18239686 Free PMC article. - Cellular and molecular consequences of defective Fanconi anemia proteins in replication-coupled DNA repair: mechanistic insights.
Thompson LH, Hinz JM. Thompson LH, et al. Mutat Res. 2009 Jul 31;668(1-2):54-72. doi: 10.1016/j.mrfmmm.2009.02.003. Epub 2009 Feb 21. Mutat Res. 2009. PMID: 19622404 Free PMC article. Review. - Kip3-ing kinetochores clustered.
Ohi R. Ohi R. Cell Cycle. 2010 Jul 1;9(13):2497. doi: 10.4161/cc.9.13.12274. Cell Cycle. 2010. PMID: 20647749 Free PMC article. No abstract available. - Repair of DNA interstrand cross-links during S phase of the mammalian cell cycle.
Legerski RJ. Legerski RJ. Environ Mol Mutagen. 2010 Jul;51(6):540-51. doi: 10.1002/em.20566. Environ Mol Mutagen. 2010. PMID: 20658646 Free PMC article. Review. - Fanconi anaemia and cancer: an intricate relationship.
Nalepa G, Clapp DW. Nalepa G, et al. Nat Rev Cancer. 2018 Mar;18(3):168-185. doi: 10.1038/nrc.2017.116. Epub 2018 Jan 29. Nat Rev Cancer. 2018. PMID: 29376519 Review.
References
- Bartek, J., and J. Lukas. 2001. Mammalian G1- and S-phase checkpoints in response to DNA damage. Curr. Opin. Cell Biol. 13:738-747. - PubMed
- Bessho, T. 2003. Induction of DNA replication-mediated double strand breaks by psoralen DNA interstrand cross-links. J. Biol. Chem. 278:5250-5254. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM062193/GM/NIGMS NIH HHS/United States
- CA092245/CA/NCI NIH HHS/United States
- R01 CA092245/CA/NCI NIH HHS/United States
- R03 HD050242/HD/NICHD NIH HHS/United States
- GM62193/GM/NIGMS NIH HHS/United States
- R01 CA112775/CA/NCI NIH HHS/United States
- CA112775/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases