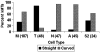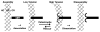Kinetochores use a novel mechanism for coordinating the dynamics of individual microtubules - PubMed (original) (raw)
Kinetochores use a novel mechanism for coordinating the dynamics of individual microtubules
Kristin J VandenBeldt et al. Curr Biol. 2006.
Abstract
Chromosome alignment during mitosis is frequently accompanied by a dynamic switching between elongation and shortening of kinetochore fibers (K-fibers) that connect kinetochores and spindle poles . In higher eukaryotes, mature K-fibers consist of 10-30 kinetochore microtubules (kMTs) whose plus ends are embedded in the kinetochore . A critical and long-standing question is how the dynamics of individual kMTs within the K-fiber are coordinated . We have addressed this question by using electron tomography to determine the polymerization/depolymerization status of individual kMTs in the K-fibers of PtK1 and Drosophila S2 cells. Surprisingly, we find that the plus ends of two-thirds of kMTs are in a depolymerizing state, even when the K-fiber exhibits net tubulin incorporation at the plus end . Furthermore, almost all individual K-fibers examined had a mixture of kMTs in the polymerizing and depolymerizing states. Therefore, although K-fibers elongate and shrink as a unit, the dynamics of individual kMTs within a K-fiber are not coordinated at any given moment. Our results suggest a novel control mechanism through which attachment to the kinetochore outer plate prevents shrinkage of kMTs. We discuss the ramifications of this new model on the regulation of chromosome movement and the stability of K-fibers.
Figures
Figure 1
Illustration of the Use of Electron Tomography to Identify the Plus-End Conformations of kMTs in PtK1 Cells (A) EM image of a 200 nm thick plastic section containing a metaphase chromosome from a PtK1 cell prepared by high-pressure freezing and freeze substitution. Scale bar = 1 μm. (B) Higher magnification image of the boxed area in (A), with the chromosome, kinetochore, and kMTs labeled. Scale bar = 200 nm. (C) A 1.6 nm thick slice from the tomographic reconstruction of the same area represented in (B). The kinetochore outer plate and two kMTs displaying the curved conformation are indicated. Scale bar = 200 nm. (D and E) Examples of the range of straight (D) and curved (E) conformations that were detected for the plus ends of kMTs by electron tomography. Underneath each image is the percentage of kMT plus ends in untreated metaphase PtK1 cells having a similar conformation. Scale bars represent 25 nm.
Figure 2
Bar Graph of kMT Plus-End Conformations The percentage of kMT plus ends classified as straight and curved are plotted for: M, untreated metaphase; T, taxol-treated metaphase; N, nocodazole-treated metaphase; A, untreated anaphase PtK1 cells; and S2, untreated metaphase Drosophila S2 cells. The sample size (number of kMT plus ends classified) is indicated in parentheses on the abscissa.
Figure 3
Plus-End Conformations of the kMTs in Individual K-Fibers from Metaphase PtK1 and S2 Cells (A) A plot of the number of kMT plus ends classified in the straight and curved conformations for ten K-fibers from four different cells. On average, K-fibers in PtK1 cells contain 23 MTs [3], but for technical reasons, we only analyzed 8–16 plus ends per kinetochore as a random sampling of the total. The ordering of kinetochores along the abscissa is by increasing percentage of kMTs found in the curved conformation. (B) Histogram of the percentage of kMTs exhibiting the curved conformation in 27 K-fibers, for which 3–16 kMTs each were analyzed. The broad distribution with a weak secondary peak at a low percentage of curved plus ends indicates that the kMT plus-end conformations within a given K-fiber are not completely random. (C) A plot of the number of kMT plus ends classified in the straight and curved conformations for K-fibers from three pairs of sister kinetochores in PTK metaphase cells. (D) A plot of the number of kMT plus ends classified in the straight and curved conformations for six K-fibers from a S2 cell.
Figure 4
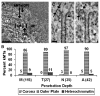
Depth that kMTs Penetrated into the Centromere (A) A single 10 nm thick slice from a tomographic reconstruction, with regional delineations for the corona (CR), outer plate (OP), and heterochromatin (HC). Two kMTs terminating in the outer plate are indicated (kMTs [OP]). Scale bar = 200 nm. (B) Bar graph of kMT penetration. The percentages of kMTs terminating in the three regions delineated in (A) are indicated above the bars for: M, untreated; T, taxol treated; N, nocodazole-treated metaphase PtK1 cells; and A, untreated anaphase PtK1 cells. The sample sizes are indicated in parentheses on the abscissa. Measurements were limited to kinetochores for which a clearly defined outer plate was visible. (C) Curved plus ends showing the kinks (arrows) and tight-radius curvatures (arrowhead) that were frequently observed. Scale bar = 25 nm.
Figure 5
Model for kMT Dynamic Instability on PtK1 and Drosophila S2 Kinetochores A single kMT is shown undergoing a conformational change from straight (left side) to curved (right side). A key feature of the model is that attachment to the outer plate prevents kMT disassembly that is not coupled to chromosome motion. This enables kMTs in the disassembly phase to be rescued, which in turn results in a continual cycling of individual kMTs that are in the growing phase. The cycling of kMTs between the growing and shrinking phases permits net incorporation of tubulin into the plus end of the K-fiber, even though the majority of its kMTs are in the disassembly phase. The catastrophe rate is shown as twice the rescue rate because that generates a 1:2 ratio of straight to curved conformations. Dissociation from the outer plate is low in PtK1 cells but is much higher in cells with a high kMT turnover rate. In the latter case, dynamic cycles of MT growth and shrinkage would occur primarily in the spindle, rather than on the kinetochore outer plate. However, the principle of cycling between which individual kMTs are contributing to K-fiber growth would be the same. The heterochromatin (HC), outer plate (OP), and corona (CR) designations are the same as those given in Figure 4A.
Comment in
- Mitosis: disorderly conduct at the kinetochore.
Compton DA. Compton DA. Curr Biol. 2006 Jul 11;16(13):R494-6. doi: 10.1016/j.cub.2006.06.010. Curr Biol. 2006. PMID: 16824907
Similar articles
- The outer plate in vertebrate kinetochores is a flexible network with multiple microtubule interactions.
Dong Y, Vanden Beldt KJ, Meng X, Khodjakov A, McEwen BF. Dong Y, et al. Nat Cell Biol. 2007 May;9(5):516-22. doi: 10.1038/ncb1576. Epub 2007 Apr 15. Nat Cell Biol. 2007. PMID: 17435749 Free PMC article. - Direct kinetochore-spindle pole connections are not required for chromosome segregation.
Sikirzhytski V, Magidson V, Steinman JB, He J, Le Berre M, Tikhonenko I, Ault JG, McEwen BF, Chen JK, Sui H, Piel M, Kapoor TM, Khodjakov A. Sikirzhytski V, et al. J Cell Biol. 2014 Jul 21;206(2):231-43. doi: 10.1083/jcb.201401090. Epub 2014 Jul 14. J Cell Biol. 2014. PMID: 25023516 Free PMC article. - Kinetochore-driven formation of kinetochore fibers contributes to spindle assembly during animal mitosis.
Maiato H, Rieder CL, Khodjakov A. Maiato H, et al. J Cell Biol. 2004 Dec 6;167(5):831-40. doi: 10.1083/jcb.200407090. Epub 2004 Nov 29. J Cell Biol. 2004. PMID: 15569709 Free PMC article. - Dynamic regulation of kinetochore-microtubule interaction during mitosis.
Tanaka K. Tanaka K. J Biochem. 2012 Nov;152(5):415-24. doi: 10.1093/jb/mvs109. Epub 2012 Sep 20. J Biochem. 2012. PMID: 22995986 Review. - The KMN protein network--chief conductors of the kinetochore orchestra.
Varma D, Salmon ED. Varma D, et al. J Cell Sci. 2012 Dec 15;125(Pt 24):5927-36. doi: 10.1242/jcs.093724. Epub 2013 Feb 15. J Cell Sci. 2012. PMID: 23418356 Free PMC article. Review.
Cited by
- Mechanical coupling coordinates microtubule growth.
Leeds BK, Kostello KF, Liu YY, Nelson CR, Biggins S, Asbury CL. Leeds BK, et al. Elife. 2023 Dec 27;12:RP89467. doi: 10.7554/eLife.89467. Elife. 2023. PMID: 38150374 Free PMC article. - Mechanical coupling coordinates microtubule growth.
Leeds BK, Kostello KF, Liu YY, Nelson CR, Biggins S, Asbury CL. Leeds BK, et al. bioRxiv [Preprint]. 2023 Oct 17:2023.06.29.547092. doi: 10.1101/2023.06.29.547092. bioRxiv. 2023. PMID: 37905093 Free PMC article. Updated. Preprint. - Mechanisms of microtubule dynamics and force generation examined with computational modeling and electron cryotomography.
Gudimchuk NB, Ulyanov EV, O'Toole E, Page CL, Vinogradov DS, Morgan G, Li G, Moore JK, Szczesna E, Roll-Mecak A, Ataullakhanov FI, Richard McIntosh J. Gudimchuk NB, et al. Nat Commun. 2020 Jul 28;11(1):3765. doi: 10.1038/s41467-020-17553-2. Nat Commun. 2020. PMID: 32724196 Free PMC article. - The kinesin-13 proteins Kif2a, Kif2b, and Kif2c/MCAK have distinct roles during mitosis in human cells.
Manning AL, Ganem NJ, Bakhoum SF, Wagenbach M, Wordeman L, Compton DA. Manning AL, et al. Mol Biol Cell. 2007 Aug;18(8):2970-9. doi: 10.1091/mbc.e07-02-0110. Epub 2007 May 30. Mol Biol Cell. 2007. PMID: 17538014 Free PMC article. - Mitotic spindle: kinetochore fibers hold on tight to interpolar bundles.
Tolić IM. Tolić IM. Eur Biophys J. 2018 Apr;47(3):191-203. doi: 10.1007/s00249-017-1244-4. Epub 2017 Jul 19. Eur Biophys J. 2018. PMID: 28725997 Free PMC article. Review.
References
- Maiato H, Deluca J, Salmon ED, Earnshaw WC. The dynamic kinetochore-microtubule interface. J Cell Sci. 2004;117:5461–5477. - PubMed
- McIntosh JR, Grishchuk EL, West RR. Chromosome-microtubule interactions during mitosis. Annu Rev Cell Dev Biol. 2002;18:193–219. - PubMed
- Kline-Smith SL, Walczak CE. Mitotic spindle assembly and chromosome segregation: refocusing on microtubule dynamics. Mol Cell. 2004;15:317–327. - PubMed
Publication types
MeSH terms
Grants and funding
- P41 RR01219/RR/NCRR NIH HHS/United States
- R01 GM06627/GM/NIGMS NIH HHS/United States
- P41 RR001219/RR/NCRR NIH HHS/United States
- R01 GM040198/GM/NIGMS NIH HHS/United States
- R01 GM066270/GM/NIGMS NIH HHS/United States
- R01 GM40198/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous