Prediction of effective genome size in metagenomic samples - PubMed (original) (raw)
Prediction of effective genome size in metagenomic samples
Jeroen Raes et al. Genome Biol. 2007.
Abstract
We introduce a novel computational approach to predict effective genome size (EGS; a measure that includes multiple plasmid copies, inserted sequences, and associated phages and viruses) from short sequencing reads of environmental genomics (or metagenomics) projects. We observe considerable EGS differences between environments and link this with ecologic complexity as well as species composition (for instance, the presence of eukaryotes). For example, we estimate EGS in a complex, organism-dense farm soil sample at about 6.3 megabases (Mb) whereas that of the bacteria therein is only 4.7 Mb; for bacteria in a nutrient-poor, organism-sparse ocean surface water sample, EGS is as low as 1.6 Mb. The method also permits evaluation of completion status and assembly bias in single-genome sequencing projects.
Figures
Figure 1
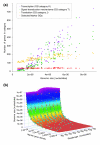
Predicting effective genome size from marker gene density. (a) Gene counts for various functional classes [50] and their relationship with genome size. Although counts of genes belonging to the categories T (signal transduction mechanisms), K (transcription) and J (translation, ribosome structure, and biogenesis) scale (to a greater or lesser extent) with genome size, the set of 35 universal, single-copy genes used in this study does not. (b) Calibration plane used to identify the relationship between marker gene density, read length, and genome size. The calibration was based on a simulated shotgun dataset of randomly extracted 'reads' from the sequenced genomes (see Materials and methods), because insufficient raw shotgun sequence data are currently available in the trace archives to allow a robust calibration based on 'real' data. Circles represent shotgun datasets. Circle fill color indicates the goodness of fit to the plane (blue = <1 standard deviation [SD], green = <2 SD, yellow = <3 SD, red = >3 SD). Circle border indicates position relative to plane (blue = above, red = below). OG, orthologous group.
Figure 2
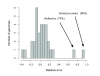
Prediction error and identification of sequencing artifacts. Distribution of the prediction error ([predicted - known genome size]/known genome size) of 32 complete genome shotgun datasets downloaded from the NCBI's trace archive (see Additional data file 9 for a list). The majority of predictions have an error estimate <20%, with a median value of about 9%. There are, however, two exceptions in which the error is significantly larger. The first is the Wolbachia endosymbiont of Drosophila melanogaster. The marker OG density in the simulated reads is considerably higher than in the real shotgun data, leading to a 70% difference in predicted genome size. After further investigation of the raw reads, we noticed that this difference was caused by an important contamination of the dataset by reads originating from the organism's host, Drosophila, that were filtered out during the assembly of the genome but that are still present in the shotgun data available at the trace archive. The second exception is the genome of the PCE-dechlorination bacterium Dehalococcoides ethenogenes. Also here, the marker OG density in the shotgun data is lower than in the simulated dataset. Mapping of the publicly available reads to the genome sequence showed a peak of read density in a region that was identified to be an integrated element that is believed to exist in variable copy numbers in different individuals but was only included once in the published genome sequence [51]. OG, orthologous group.
Figure 3
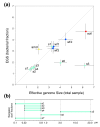
Predicted effective genome sizes for environments. (a) Comparison of predicted EGS for total samples versus the bacterial fraction. amd, acid mine drainage; wf, whale fall deep sea samples; s, Sargasso Sea samples. Error bars indicate standard deviation for total (horizontal) and bacteria-specific (vertical) estimates. (b) Overview of cell size in the different Sargasso Sea samples due to filtering during sampling.
References
- Gregory TR, DeSalle R. Comparative genomics in prokaryotes. In: Gregory TR, editor. The Evolution of the Genome. San Diego: Elsevier; 2005. pp. 585–675.
- Loferer-Krossbacher M, Witzel K-P, Psenner R. DNA content of aquatic bacteria measured by densitometric image analysis. Arch Hydrobiol Spec Issues Advanc Limnol. 1999;54:185–198.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous