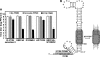In vitro resistance selection and in vivo efficacy of morpholino oligomers against West Nile virus - PubMed (original) (raw)
In vitro resistance selection and in vivo efficacy of morpholino oligomers against West Nile virus
Tia S Deas et al. Antimicrob Agents Chemother. 2007 Jul.
Abstract
We characterize in vitro resistance to and demonstrate the in vivo efficacy of two antisense phosphorodiamidate morpholino oligomers (PMOs) against West Nile virus (WNV). Both PMOs were conjugated with an Arg-rich peptide. One peptide-conjugated PMO (PPMO) binds to the 5' terminus of the viral genome (5'-end PPMO); the other targets an essential 3' RNA element required for genome cyclization (3' conserved sequence I [3' CSI] PPMO). The 3' CSI PPMO displayed a broad spectrum of antiflavivirus activity, suppressing WNV, Japanese encephalitis virus, and St. Louis encephalitis virus, as demonstrated by reductions in viral titers of 3 to 5 logs in cell cultures, likely due to the absolute conservation of the 3' CSI PPMO-targeted sequences among these viruses. The selection and sequencing of PPMO-resistant WNV showed that the 5'-end-PPMO-resistant viruses contained two to three mismatches within the PPMO-binding site whereas the 3' CSI PPMO-resistant viruses accumulated mutations outside the PPMO-targeted region. The mutagenesis of a WNV infectious clone demonstrated that the mismatches within the PPMO-binding site were responsible for the 5'-end PPMO resistance. In contrast, a U insertion or a G deletion located within the 3'-terminal stem-loop of the viral genome was the determinant of the 3' CSI PPMO resistance. In a mouse model, both the 5'-end and 3' CSI PPMOs (administered at 100 or 200 microg/day) partially protected mice from WNV disease, with minimal to no PPMO-mediated toxicity. A higher treatment dose (300 microg/day) caused toxicity. Unconjugated PMOs (3 mg/day) showed neither efficacy nor toxicity, suggesting the importance of the peptide conjugate for efficacy. The results suggest that a modification of the peptide conjugate composition to reduce its toxicity yet maintain its ability to effectively deliver PMO into cells may improve PMO-mediated therapy.
Figures
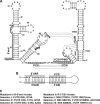
FIG. 1.
Sequences and structures of the PPMO-targeted WNV RNA regions. (A) Terminal stem-loop structures and potential RNA-RNA interactions within the WNV genome. Secondary structures formed by the 5′- and 3′-terminal sequences of the WNV genome are depicted. Two PPMOs, the 5′-end PPMO and the 3′ CSI PPMO, are indicated as thick lines with filled circles (representing an Arg-rich peptide [11]) at the 5′ ends. The AUG initiation codon of the open reading frame is shaded in gray. Three RNA interactions are denoted by dashed lines, the 5′ UAR-3′ UAR and 5′ CS-3′ CSI interactions and a pseudoknot base pairing (located at the 3′-terminal stem-loops). The sequences involved in the 5′ UAR-3′ UAR and 5′ CS-3′ CSI interactions are indicated by thin lines. Nucleotide positions are numbered based on the full-length sequence of the WNV genome. Gppp indicates a gap structure of viral genome. (B) Potential genome cyclization of WNV through the 5′ UAR-3′ UAR and 5′ CS-3′ CSI base pairings. (C) Adaptive mutations derived from viruses resistant to the 5′-end PPMO (left panel) or the 3′ CSI PPMO (right panel).
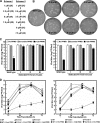
FIG. 2.
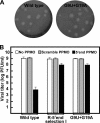
Selection and characterization of PPMO-resistant WNV. (A) Two schemes for the selection of PPMO-resistant viruses. (B) Plaque morphologies of wild-type and PPMO-resistant viruses. Plaque assays for the resistant and wild-type viruses were performed on Vero cells in the absence of PPMOs. The results of three independent selections for 5′-end-PPMO- and 3′ CSI PPMO-resistant viruses are presented. (C) Resistance phenotypes of the PPMO escape isolates. The left and right panels represent results for 5′-end-PPMO-resistant and 3′ CSI PPMO-resistant viruses, respectively. The results are expressed as the means and standard deviations derived from three independent experiments. See the experimental details in Materials and Methods. (D) Growth kinetics of the PPMO-resistant and wild-type (WT) viruses in the presence and absence of PPMO. Vero cells were infected (MOI of 0.1) for 1 h, washed with PBS, and incubated for the indicated times in medium with or without PPMO at 7.5 μM. The averages of results from three experiments are presented.
FIG. 3.
Analysis of mutations recovered from 5′-end-PPMO-resistant viruses. (A) Plaque morphologies of wild-type and recombinant G9U G19A viruses. (B) Resistance analyses of wild-type virus, R-5′ end virus from selection I, and recombinant G9U G19A virus. See Materials and Methods for experimental details.
FIG. 4.
Verification of 10957U-ins as a determinant of 3′ CSI PPMO resistance. (A) Resistance phenotypes of wild-type virus, R-3′ CSI virus from selection I, and three recombinant viruses (C7891U, 10957U-ins, and C7891U 10957U-ins viruses). The experimental procedures were as described in Materials and Methods. The data are means ± standard deviations (n ≥ 3). (B) Effects of 10957U-ins on the thermodynamically predicted 5′ UAR-3′ UAR interaction. (C) 10957U-ins-mediated disturbance of an internal stem region of the 3′ stem-loop structure of the WNV genome. The 3′ CSI PPMO and the 3′ CSI are indicated by thick and thin lines, respectively. In panels B and C, regions containing the 10957U-ins-mediated structural changes are highlighted in gray.
FIG. 5.
Effects of ΔG11024 on WNV resistance to the 3′ CSI PPMO. (A) Resistance profiles of R-3′ CSI virus from selection II and four recombinant viruses (G8074A, G10965U, ΔG11024, and G8074A G10965U viruses). See Materials and Methods for details. (B) Effects of ΔG11024 on the MFold-predicted 3′ stem-loop structure of the WNV genome. Structural changes caused by ΔG11024 are highlighted in gray.
FIG. 6.
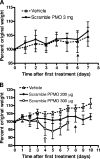
Body weights of mice treated with PMOs or PPMOs. Six-week-old C3H female mice were treated, weighed, and observed daily. Averages of the percentages of original weights are shown, and error bars represent standard deviations of ±1. (A) Mice were inoculated i.p. daily with the vehicle alone (n = 4) or 3 mg of the scramble PMO (n = 8) for seven treatments. The solid arrow indicates the last day of treatment. (B) Mice were inoculated i.p. daily with the vehicle alone (n = 4) for nine treatments, 200 μg of the scramble PPMO (n = 8) for nine treatments, or 300 μg of the scramble PPMO (n = 8) for five treatments. The solid arrow indicates the last day of treatment for groups receiving the vehicle alone and 200 μg of the scramble PPMO. The hollow arrow indicates the last day of treatment for the group receiving 300 μg of the scramble PPMO, which was discontinued after only five treatments due to an average loss among the animals of more than 5% of body weight.
FIG. 7.
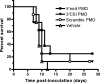
Unconjugated PMOs do not protect mice from WNV disease. Six-week-old C3H female mice were inoculated s.c. in the left rear footpad with 103 PFU of WNV on day 0. Groups of eight mice were treated with 3 mg of the 5′-end PMO, 3 mg of the 3′ CSI PMO, 3 mg of the scramble PMO, or the vehicle alone. Treatments were administered i.p. daily on days 0 to 6.
FIG. 8.
PPMOs partially protect mice from WNV disease. Six-week-old C3H female mice were inoculated s.c. in the left rear footpad with 103 PFU of WNV on day 0. Groups of eight mice were treated i.p. with 100 or 200 μg of PPMOs or the vehicle alone. (A and B) Survival rates among mice treated daily on days 0 to 8 with the 5′-end PPMO and the 3′ CSI PPMO, respectively. The survival curves for groups treated with the vehicle alone and the scramble PPMO are the same in panels A and B and are repeated for ease of comparison. The asterisk indicates that the survival curve for the group treated with the 5′-end PPMO at 200 μg was significantly different from that for the group treated with the scramble PPMO at 200 μg (log rank test; P = 0.04). (C) Average clinical scores for groups treated daily with the indicated PPMOs on days 0 to 8. Clinical scores were assigned a scale of 0 (normal) to 6 (dead or euthanized). The scale of clinical scores is further defined in Materials and Methods. (D) Survival rates of mice treated daily from days 5 to 15 after viral inoculation.
Similar articles
- Co-selection of West Nile virus nucleotides that confer resistance to an antisense oligomer while maintaining long-distance RNA/RNA base pairings.
Zhang B, Dong H, Stein DA, Shi PY. Zhang B, et al. Virology. 2008 Dec 5;382(1):98-106. doi: 10.1016/j.virol.2008.08.044. Epub 2008 Oct 7. Virology. 2008. PMID: 18842280 Free PMC article. - Inhibition of RNA virus infections with peptide-conjugated morpholino oligomers.
Stein DA. Stein DA. Curr Pharm Des. 2008;14(25):2619-34. doi: 10.2174/138161208786071290. Curr Pharm Des. 2008. PMID: 18991679 Review. - Inhibition of alphavirus infection in cell culture and in mice with antisense morpholino oligomers.
Paessler S, Rijnbrand R, Stein DA, Ni H, Yun NE, Dziuba N, Borisevich V, Seregin A, Ma Y, Blouch R, Iversen PL, Zacks MA. Paessler S, et al. Virology. 2008 Jul 5;376(2):357-70. doi: 10.1016/j.virol.2008.03.032. Epub 2008 May 12. Virology. 2008. PMID: 18468653 Free PMC article. - Inhibition of flavivirus infections by antisense oligomers specifically suppressing viral translation and RNA replication.
Deas TS, Binduga-Gajewska I, Tilgner M, Ren P, Stein DA, Moulton HM, Iversen PL, Kauffman EB, Kramer LD, Shi PY. Deas TS, et al. J Virol. 2005 Apr;79(8):4599-609. doi: 10.1128/JVI.79.8.4599-4609.2005. J Virol. 2005. PMID: 15795246 Free PMC article. - Cell-penetrating peptide-morpholino conjugates alter pre-mRNA splicing of DMD (Duchenne muscular dystrophy) and inhibit murine coronavirus replication in vivo.
Moulton HM, Fletcher S, Neuman BW, McClorey G, Stein DA, Abes S, Wilton SD, Buchmeier MJ, Lebleu B, Iversen PL. Moulton HM, et al. Biochem Soc Trans. 2007 Aug;35(Pt 4):826-8. doi: 10.1042/BST0350826. Biochem Soc Trans. 2007. PMID: 17635157 Review.
Cited by
- Splice-Switching Therapy for Spinal Muscular Atrophy.
Meijboom KE, Wood MJA, McClorey G. Meijboom KE, et al. Genes (Basel). 2017 Jun 12;8(6):161. doi: 10.3390/genes8060161. Genes (Basel). 2017. PMID: 28604635 Free PMC article. Review. - Functional Information Stored in the Conserved Structural RNA Domains of Flavivirus Genomes.
Fernández-Sanlés A, Ríos-Marco P, Romero-López C, Berzal-Herranz A. Fernández-Sanlés A, et al. Front Microbiol. 2017 Apr 3;8:546. doi: 10.3389/fmicb.2017.00546. eCollection 2017. Front Microbiol. 2017. PMID: 28421048 Free PMC article. Review. - Evolution of resistance to fluoroquinolones by dengue virus serotype 4 provides insight into mechanism of action and consequences for viral fitness.
Scroggs SLP, Gass JT, Chinnasamy R, Widen SG, Azar SR, Rossi SL, Arterburn JB, Vasilakis N, Hanley KA. Scroggs SLP, et al. Virology. 2021 Jan 2;552:94-106. doi: 10.1016/j.virol.2020.09.004. Epub 2020 Oct 1. Virology. 2021. PMID: 33120225 Free PMC article. - Reverse genetics technology for Rift Valley fever virus: current and future applications for the development of therapeutics and vaccines.
Bouloy M, Flick R. Bouloy M, et al. Antiviral Res. 2009 Nov;84(2):101-18. doi: 10.1016/j.antiviral.2009.08.002. Epub 2009 Aug 12. Antiviral Res. 2009. PMID: 19682499 Free PMC article. Review. - Advanced morpholino oligomers: a novel approach to antiviral therapy.
Warren TK, Shurtleff AC, Bavari S. Warren TK, et al. Antiviral Res. 2012 Apr;94(1):80-8. doi: 10.1016/j.antiviral.2012.02.004. Epub 2012 Feb 14. Antiviral Res. 2012. PMID: 22353544 Free PMC article. Review.
References
- Bredenbeek, P. J., E. A. Kooi, B. Lindenbach, N. Huijkman, C. M. Rice, and W. J. Spaan. 2003. A stable full-length yellow fever virus cDNA clone and the role of conserved RNA elements in flavivirus replication. J. Gen. Virol. 841261-1268. - PubMed
- Brinton, M. A., and J. H. Dispoto. 1988. Sequence and secondary structure analysis of the 5′-terminal region of flavivirus genome RNA. Virology 162290-299. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U54-AI057158/AI/NIAID NIH HHS/United States
- R43 AI065156/AI/NIAID NIH HHS/United States
- U01 AI061193/AI/NIAID NIH HHS/United States
- U54 AI057158/AI/NIAID NIH HHS/United States
- 1R43AI065156/AI/NIAID NIH HHS/United States
- N01 AI025490/AI/NIAID NIH HHS/United States
- 1U01AI061193/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources