DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists - PubMed (original) (raw)
. 2007 Jul;35(Web Server issue):W169-75.
doi: 10.1093/nar/gkm415. Epub 2007 Jun 18.
Affiliations
- PMID: 17576678
- PMCID: PMC1933169
- DOI: 10.1093/nar/gkm415
DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists
Da Wei Huang et al. Nucleic Acids Res. 2007 Jul.
Abstract
All tools in the DAVID Bioinformatics Resources aim to provide functional interpretation of large lists of genes derived from genomic studies. The newly updated DAVID Bioinformatics Resources consists of the DAVID Knowledgebase and five integrated, web-based functional annotation tool suites: the DAVID Gene Functional Classification Tool, the DAVID Functional Annotation Tool, the DAVID Gene ID Conversion Tool, the DAVID Gene Name Viewer and the DAVID NIAID Pathogen Genome Browser. The expanded DAVID Knowledgebase now integrates almost all major and well-known public bioinformatics resources centralized by the DAVID Gene Concept, a single-linkage method to agglomerate tens of millions of diverse gene/protein identifiers and annotation terms from a variety of public bioinformatics databases. For any uploaded gene list, the DAVID Resources now provides not only the typical gene-term enrichment analysis, but also new tools and functions that allow users to condense large gene lists into gene functional groups, convert between gene/protein identifiers, visualize many-genes-to-many-terms relationships, cluster redundant and heterogeneous terms into groups, search for interesting and related genes or terms, dynamically view genes from their lists on bio-pathways and more. With DAVID (http://david.niaid.nih.gov), investigators gain more power to interpret the biological mechanisms associated with large gene lists.
Figures
Figure 1.
A DAVID gene constructed by a single linkage algorithm. Two UniRef100 clusters, two NRef 100 clusters and one Entrez Gene cluster were systematically found sharing one or more protein identifiers with each other. The single-linkage rule can further iteratively agglomerate them as a whole into one DAVID gene. Thus, for this particular example of tyrosine-protein phosphatase non-receptor type 21 (PTPN21), the resulting DAVID gene is able to collect and integrate all gene/protein identifiers more comprehensively than each original gene cluster.
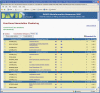
Figure 2.
An HTML report from the Functional Annotation Clustering. The annotation cluster 1 in the example shows that GO term cytokine activity, KEGG pathway cytokine–cytokine receptor interaction, and GO term receptor binding, etc. are grouped together. Thus, the different biological aspects regarding a relevant biology can be explored at the same time.
Figure 3.
A roadmap to choose appropriate DAVID functions and tools.
Similar articles
- DAVID Knowledgebase: a gene-centered database integrating heterogeneous gene annotation resources to facilitate high-throughput gene functional analysis.
Sherman BT, Huang da W, Tan Q, Guo Y, Bour S, Liu D, Stephens R, Baseler MW, Lane HC, Lempicki RA. Sherman BT, et al. BMC Bioinformatics. 2007 Nov 2;8:426. doi: 10.1186/1471-2105-8-426. BMC Bioinformatics. 2007. PMID: 17980028 Free PMC article. - Extracting biological meaning from large gene lists with DAVID.
Huang da W, Sherman BT, Zheng X, Yang J, Imamichi T, Stephens R, Lempicki RA. Huang da W, et al. Curr Protoc Bioinformatics. 2009 Sep;Chapter 13:Unit 13.11. doi: 10.1002/0471250953.bi1311s27. Curr Protoc Bioinformatics. 2009. PMID: 19728287 - Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources.
Huang da W, Sherman BT, Lempicki RA. Huang da W, et al. Nat Protoc. 2009;4(1):44-57. doi: 10.1038/nprot.2008.211. Nat Protoc. 2009. PMID: 19131956 - Interpreting microarray results with gene ontology and MeSH.
Osborne JD, Zhu LJ, Lin SM, Kibbe WA. Osborne JD, et al. Methods Mol Biol. 2007;377:223-42. doi: 10.1007/978-1-59745-390-5_14. Methods Mol Biol. 2007. PMID: 17634620 Review. - Integration of bioinformatics resources for functional analysis of gene expression and proteomic data.
Huang H, Hu ZZ, Arighi CN, Wu CH. Huang H, et al. Front Biosci. 2007 Sep 1;12:5071-88. doi: 10.2741/2449. Front Biosci. 2007. PMID: 17569631 Review.
Cited by
- Berberine Improves Inflammatory Responses of Diabetes Mellitus in Zucker Diabetic Fatty Rats and Insulin-Resistant HepG2 Cells through the PPM1B Pathway.
Wu YS, Li ZM, Chen YT, Dai SJ, Zhou XJ, Yang YX, Lou JS, Ji LT, Bao YT, Xuan L, Lin LN, Li CY. Wu YS, et al. J Immunol Res. 2020 Aug 18;2020:2141508. doi: 10.1155/2020/2141508. eCollection 2020. J Immunol Res. 2020. PMID: 32908938 Free PMC article. - Visual analysis of biological data-knowledge networks.
Vehlow C, Kao DP, Bristow MR, Hunter LE, Weiskopf D, Görg C. Vehlow C, et al. BMC Bioinformatics. 2015 Apr 29;16:135. doi: 10.1186/s12859-015-0550-z. BMC Bioinformatics. 2015. PMID: 25925016 Free PMC article. - Combined serial analysis of gene expression and transcription factor binding site prediction identifies novel-candidate-target genes of Nr2e1 in neocortex development.
Schmouth JF, Arenillas D, Corso-Díaz X, Xie YY, Bohacec S, Banks KG, Bonaguro RJ, Wong SH, Jones SJ, Marra MA, Simpson EM, Wasserman WW. Schmouth JF, et al. BMC Genomics. 2015 Jul 24;16(1):545. doi: 10.1186/s12864-015-1770-3. BMC Genomics. 2015. PMID: 26204903 Free PMC article. - Bioinformatics analysis of prognosis-related long non-coding RNAs in invasive breast carcinoma.
Hu Y, Gu X, Duan Y, Shen Y, Xie X. Hu Y, et al. Oncol Lett. 2020 Jul;20(1):113-122. doi: 10.3892/ol.2020.11558. Epub 2020 Apr 21. Oncol Lett. 2020. PMID: 32565939 Free PMC article. - Dynamic dysregulation of transcriptomic networks in brainstem autonomic nuclei during hypertension development in the female spontaneously hypertensive rat.
Moss A, Kuttippurathu L, Srivastava A, Schwaber JS, Vadigepalli R. Moss A, et al. Physiol Genomics. 2024 Mar 1;56(3):283-300. doi: 10.1152/physiolgenomics.00073.2023. Epub 2023 Dec 25. Physiol Genomics. 2024. PMID: 38145287 Free PMC article.
References
- Dennis G, Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003;4:P3. - PubMed
- Maere S, Heymans K, Kuiper M. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics. 2005;21:3448–3449. - PubMed
- Berriz GF, King OD, Bryant B, Sander C, Roth FP. Characterizing gene sets with FuncAssociate. Bioinformatics. 2003;19:2502–2504. - PubMed
- Bluthgen N, Brand K, Cajavec B, Swat M, Herzel H, Beule D. Biological profiling of gene groups utilizing Gene Ontology. Genome Inform. 2005;16:106–115. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources