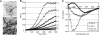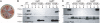The curli nucleator protein, CsgB, contains an amyloidogenic domain that directs CsgA polymerization - PubMed (original) (raw)
The curli nucleator protein, CsgB, contains an amyloidogenic domain that directs CsgA polymerization
Neal D Hammer et al. Proc Natl Acad Sci U S A. 2007.
Abstract
Curli are functional amyloid fibers assembled by enteric bacteria such as Escherichia coli and Salmonella spp. In E. coli, the polymerization of the major curli fiber subunit protein CsgA into an amyloid fiber depends on the minor curli subunit protein, CsgB. The outer membrane-localized CsgB protein shares approximately 30% sequence identity with the amyloid-forming protein CsgA, suggesting that CsgB might also have amyloidogenic properties. Here, we characterized the biochemical properties of CsgB and the molecular basis for CsgB-mediated nucleation of CsgA. Deletion analysis revealed that a CsgB molecule missing 19 amino acids from its C terminus (CsgB(trunc)) was not outer membrane-associated, but secreted away from the cell. CsgB(trunc) was overexpressed and purified from the extracellular milieu of cells as an SDS-soluble, nonaggregated protein. Soluble CsgB(trunc) assembled into fibers that bound to the amyloid-specific dyes Congo red and thioflavin-T. CsgB(trunc) fibers were able to seed soluble CsgA polymerization in vitro. CsgB(trunc) displayed modest nucleator activity in vivo, as demonstrated by its ability to convert extracellular CsgA into an amyloid fiber. Unlike WT CsgB, CsgB(trunc) was only able to act as a nucleator when cells were genetically manipulated to secrete higher concentrations of CsgA. This work represents a unique demonstration of functional amyloid nucleation and it suggests an elegant model for how E. coli guides efficient amyloid fiber formation on the cell surface.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Fig. 1.
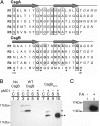
Biochemical characterization of CsgB and CsgBtrunc. (A) The imperfect repeating units in CsgA and CsgB (R1–R5) are predicted to form β-strand-loop-β-strand structures. Amino acids comprising the β-strand are located below the arrows, and amino acids predicted to comprise the loops are italicized. Bolded letters represent amino acids conserved in CsgB and CsgA at each position relative to the start of each repeating unit. Boxed letters represent amino acids conserved throughout the repeating units in both proteins. Positively charged amino acids in the fifth repeating unit of CsgB are marked with asterisks. (B) Western blot analysis of WT CsgB (*) and CsgBtrunc (**) expressed in the Δ_csg_ strain LSR12. The csgB constructs were coexpressed with the isopropyl β-
d
-thiogalactoside-inducible csgG containing plasmid pMC1 or the corresponding empty vector, pTrc99A. Cultures were grown in LB to an OD600 of 0.9 and induced with 250 μM isopropyl β-
d
-thiogalactoside. Postinduction cell suspensions were normalized by OD600 and separated by centrifugation into cell (C) and supernatant fractions (S). The blot was probed by using αCsgB peptide antisera. Lane 1 contains LSR12/pTrc99A/pNH3 cells, and lane 2 contains corresponding supernatant; lanes 3 (C) and 5 (S) contain LSR12/pTrc99A/pNH4; lanes 4 (C) and 6 (S) contain LSR12/pMC1/pNH4; lanes 7 (C) and 9 (S) contain LSR12/pTrc99A/pNH2; and lanes 8 (C) and 10 (S) contain LSR12/pMC1/pNH2. (C) Western blot of purified CsgBtrunc probed with αCsgB peptide antibody. Two samples containing equal concentrations of purified soluble CsgBtrunc were incubated at room temperature overnight before centrifugation for 1 min in an Eppendorf microcentrifuge (Eppendorf-5 Prime, Boulder, CO). The supernatant was decanted and the pellet was resuspended in 2× SDS loading buffer (lane 1) or pretreated with 90% formic acid (FA) and dried before resuspension in 2× SDS loading buffer (lane 2).
Fig. 2.
Amyloid-like properties of CsgBtrunc aggregates. (A) TEM analysis of soluble CsgBtrunc incubated at room temperature overnight reveals fibrous aggregates. (Scale bars: 500 nm.) (B) Freshly purified CsgBtrunc was mixed with the amyloid-specific dye thioflavin T and incubated at room temperature for 600 min at the following concentrations: 43 μM (open diamonds), 33 μM (open circles), 22 μM (filled diamonds), 16 μM (filled circles), 11 μM (open triangles), 8 μM (“x”s), 5 μM (filled triangles), 0 μM (solid line). Relative florescence emitted at 495 nm was measured every 10 min after excitation at 438 nm. (C) Far-UV spectral analysis of 22 μM CsgBtrunc immediately after purification (open circles), after incubation for 24 h (open squares) at room temperature, or after incubation at room temperature for 15 days (filled squares). The potassium-phosphate-buffer (KPi)-only control is represented by a solid black line.
Fig. 3.
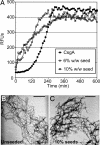
CsgBtrunc can nucleate CsgA in vitro. (A) Thioflavin T kinetic plot monitoring the polymerization of 30 μM CsgA (filled circles) and 30 μM CsgA in the presence of 10% wt/wt (open squares) or 6% wt/wt (open circles) CsgBtrunc aggregates. (B) TEM of 30 μM CsgA. (C) TEM of the fibers produced by polymerization of 30 μM CsgA seeded by 10% CsgBtrunc. (Scale bars: 500 nm.)
Fig. 4.
In vivo nucleation properties of CsgBtrunc. (A) Congo red-containing yeast extract casamino acids (YESCA) plates comparing the Congo red binding phenotypes of csgB and csgBF harboring an empty vector control (pLR2), WT csgB on a plasmid (pLR8), or csgBtrunc (pNH1). Cells: area 1, WT/pLR2; area 2, csgB/pLR2; area 3, csgB/pLR8; area 4, csgB/pNH1; area 5, csgBF/pNH1; area 6, csgBF/pNH1; area 7, csgBF/pLR8; area 8, csgBF/pLR2. All strains contain pTrc99A except area 6, which contains pCsgF. (B Upper) WC lysate samples probed with αCsgB peptide antiserum reveal WT CsgB (*) and CsgBtrunc (**). Samples were treated with (+) or without (−) formic acid (FA). (B Lower) WCs and the underlying agar (plugs) were collected from YESCA plates and probed for CsgB. (C Upper) Western blots of WCs probed with αCsgA antiserum. (C Lower) Western blots of WCs and underlying agar (plug) samples probing for CsgA. The WT cells in lanes 1 and 2 contain empty vectors pLR2 and pTrc99A. All other strains in A and B also contain the empty vector pTrc99A in addition to either pLR2 (pEmpty) or pNH1 (pCsgBtrunc).
Fig. 5.
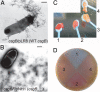
Characterization of fibers nucleated by CsgBtrunc in vivo. (A) TEM of csgB/pLR8. (B) TEM of csgBF/pNH1. (Scale bars: 500 nm.) (C) Cells were mechanically scraped off of a Congo red-containing YESCA plate by using disposable inoculating loops to compare the Congo red-binding phenotypes of WT/pLR2 (empty vector; labeled “1”), csgBF/pNH1 (csgBtrunc; labeled “2”), csgFBA/pNH1 (csgBtrunc; labeled “3”), or csgFBA/pLR2 (empty vector; labeled “4”). (D) The Congo red-containing YESCA plate used to cultivate cells in Fig. 4_C_ after cells have been removed. In D, the numbers indicate the region of the Congo red plate where the following strains had grown: 1, WT/pLR2; 2, csgBF/pNH1; 3, csgFBA/pNH1; and 4, csgFBA/pLR2.
Similar articles
- The C-terminal repeating units of CsgB direct bacterial functional amyloid nucleation.
Hammer ND, McGuffie BA, Zhou Y, Badtke MP, Reinke AA, Brännström K, Gestwicki JE, Olofsson A, Almqvist F, Chapman MR. Hammer ND, et al. J Mol Biol. 2012 Sep 21;422(3):376-89. doi: 10.1016/j.jmb.2012.05.043. Epub 2012 Jun 7. J Mol Biol. 2012. PMID: 22684146 Free PMC article. - Intrinsic aggregation propensity of the CsgB nucleator protein is crucial for curli fiber formation.
Louros NN, Bolas GMP, Tsiolaki PL, Hamodrakas SJ, Iconomidou VA. Louros NN, et al. J Struct Biol. 2016 Aug;195(2):179-189. doi: 10.1016/j.jsb.2016.05.012. Epub 2016 May 28. J Struct Biol. 2016. PMID: 27245712 - Role of Escherichia coli curli operons in directing amyloid fiber formation.
Chapman MR, Robinson LS, Pinkner JS, Roth R, Heuser J, Hammar M, Normark S, Hultgren SJ. Chapman MR, et al. Science. 2002 Feb 1;295(5556):851-5. doi: 10.1126/science.1067484. Science. 2002. PMID: 11823641 Free PMC article. - Curli provide the template for understanding controlled amyloid propagation.
Wang X, Chapman MR. Wang X, et al. Prion. 2008 Apr-Jun;2(2):57-60. doi: 10.4161/pri.2.2.6746. Epub 2008 Apr 5. Prion. 2008. PMID: 19098444 Free PMC article. Review. - Bacterial amyloid formation: structural insights into curli biogensis.
Van Gerven N, Klein RD, Hultgren SJ, Remaut H. Van Gerven N, et al. Trends Microbiol. 2015 Nov;23(11):693-706. doi: 10.1016/j.tim.2015.07.010. Epub 2015 Oct 1. Trends Microbiol. 2015. PMID: 26439293 Free PMC article. Review.
Cited by
- CsgA gatekeeper residues control nucleation but not stability of functional amyloid.
Olsen WP, Courtade G, Peña-Díaz S, Nagaraj M, Sønderby TV, Mulder FAA, Malle MG, Otzen DE. Olsen WP, et al. Protein Sci. 2024 Oct;33(10):e5178. doi: 10.1002/pro.5178. Protein Sci. 2024. PMID: 39302107 Free PMC article. - Oleanolic Acid Promotes the Formation of Probiotic Escherichia coli Nissle 1917 (EcN) Biofilm by Inhibiting Bacterial Motility.
Liu D, Liu J, Ran L, Yang Z, He Y, Yang H, Yu Y, Fu L, Zhu M, Chen H. Liu D, et al. Microorganisms. 2024 May 29;12(6):1097. doi: 10.3390/microorganisms12061097. Microorganisms. 2024. PMID: 38930479 Free PMC article. - Structural insight into Escherichia coli CsgA amyloid fibril assembly.
Bu F, Dee DR, Liu B. Bu F, et al. mBio. 2024 Apr 10;15(4):e0041924. doi: 10.1128/mbio.00419-24. Epub 2024 Mar 19. mBio. 2024. PMID: 38501920 Free PMC article. - Fluoropyrimidines affect de novo pyrimidine synthesis impairing biofilm formation in Escherichia coli.
Ravishankar S, Baldelli V, Angeletti C, Raffaelli N, Landini P, Rossi E. Ravishankar S, et al. Biofilm. 2024 Feb 7;7:100180. doi: 10.1016/j.bioflm.2024.100180. eCollection 2024 Jun. Biofilm. 2024. PMID: 38370152 Free PMC article. - The role of filamentous matrix molecules in shaping the architecture and emergent properties of bacterial biofilms.
Böhning J, Tarafder AK, Bharat TAM. Böhning J, et al. Biochem J. 2024 Feb 21;481(4):245-263. doi: 10.1042/BCJ20210301. Biochem J. 2024. PMID: 38358118 Free PMC article. Review.
References
- Kikuchi T, Mizunoe Y, Takade A, Naito S, Yoshida S. Microbiol Immunol. 2005;49:875–884. - PubMed
- Bian Z, Brauner A, Li Y, Normark S. J Infect Dis. 2000;181:602–612. - PubMed
- Bian Z, Yan ZQ, Hansson GK, Thoren P, Normark S. J Infect Dis. 2001;183:612–619. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P50-A608671/PHS HHS/United States
- AI073847-01/AI/NIAID NIH HHS/United States
- R01 AI073847-02/AI/NIAID NIH HHS/United States
- R01 AI073847/AI/NIAID NIH HHS/United States
- R56 AI073847/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous