Coactivation of G protein signaling by cell-surface receptors and an intracellular exchange factor - PubMed (original) (raw)
Coactivation of G protein signaling by cell-surface receptors and an intracellular exchange factor
Michael J Lee et al. Curr Biol. 2008.
Abstract
G protein-coupled receptors (GPCRs) mediate responses to a broad range of chemical and environmental signals. In yeast, a pheromone-binding GPCR triggers events leading to the fusion of haploid cells. In general, GPCRs function as guanine-nucleotide exchange factors (GEFs); upon agonist binding, the receptor induces a conformational change in the G protein alpha subunit, resulting in exchange of guanine diphosphate (GDP) for guanine triphosphate (GTP) and in signal initiation. Signaling is terminated when GTP is hydrolyzed to GDP [1]. This well-established paradigm has in recent years been revised to include new components that rates of GDP release, GTP binding [2-8], and GTP hydrolysis[9, 10]. Here we report the discovery of a nonreceptor GEF, Arr4. Like receptors, Arr4 binds directly to the G protein,accelerates guanine-nucleotide exchange, and stabilizes the nucleotide-free state of the a subunit. Moreover, Arr4 promotes G protein-dependent cellular responses, including mitogen-activated protein kinase (MAPK) phosphorylation,new-gene transcription, and mating. In contrast to knownGPCRs, however, Arr4 is not a transmembrane receptor,but rather a soluble intracellular protein. Our data suggest that intracellular proteins function in cooperation with mating pheromones to amplify G protein signaling, thereby leading to full pathway activation.
Figures
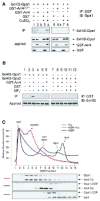
Figure 1. Arr4 as a candidate GEF
(A) Purification of Arr4-TAP from yeast. Cells expressing Arr4-TAP were transformed with vector (pAD4M) containing either no insert, GPA1, _GPA1_Q323L or _GPA1_N388D. TAP fusion protein was purified (IP) using Calmodulin-Sepharose resin, eluted in SDS-PAGE sample buffer, and resolved by immunoblotting (IB) with Gpa1 antibody. Gpa1 typically migrates as a doublet of 52 and 54 kDa, representing the myristoylated and unmyristoylated forms of the protein, respectively [27]. (B) Densitometry of data in panel A expressed as Gpa1 bound relative to total Gpa1 expressed. Data are mean ± SD of 3 separate experiments. (C) Mating gene transcription-reporter assay: effect of ARR4 over-expression. Cells co-expressing _FUS1_-lacZ and vector (pAD4M) containing either no insert, ARR4, or dimerization-deficient mutant arr4CCTT were monitored for β-galactosidase activity. (D) Effect of known mating components on Arr4 activity. Experiment performed as in C, except that receptor (ste2) or MAPK kinase (ste7) deletion strains were used when noted. Results for C and D are the mean ± SEM for 3 individual experiments each performed in triplicate.
Figure 2. Arr4 binds directly to Gpa1 and stabilizes the nucleotide-free state of the G protein
(A) and (B), Direct in vitro binding using recombinant purified components (E. coli). 100 nM of each protein was combined, purified with GST-Sepharose resin (IP), resolved by SDS-PAGE and probed with penta-HIS, Gpa1, or GST antibodies (IB). Note that the same protein preparations were used in the functional assays presented in Fig. 3. (A) Binding with or without 150 nM CuSO4 (Cu) when noted. (B) Arr4 binding to Gpa1 versus Gpa2 was performed as in panel A except that CuSO4 was present in all lanes. (C) Arr4-Gpa1 complex formation using purified components. 4 mg of 6xHIS-Arr4 and 2 mg of 6xHIS-Gpa1 were combined and resolved by steric-exclusion chromatography in the presence or absence of excess GDP and CuSO4, as indicated. Note that the void volume elutes 200 minutes after sample loading. Top panel, A260nm chromatogram. The peak of UV absorbance in the void volume is evidently due to a non-protein buffer component, as determined by Coomassie staining as well as by immunoblotting with penta-His antibodies. Bottom panels, immunoblots using penta-HIS antibody. 20 μl of each 7 ml elution fraction were loaded and resolved by SDS-PAGE. All data are representative of 3 separate experiments.
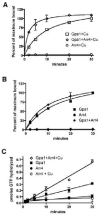
Figure 3. Arr4 is a GEF for Gpa1
(A) and (B), Single turnover GTP binding assay. Time-course of [35S]GTPγS binding to 100 nM purified 6xHIS-Gpa1 in the presence of 200 nM GST-Arr4. Results are the mean ± SEM of duplicate samples, and are presented as percent of maximum bound (saturated binding occurred between 50-75%). (A) with 500 nM CuSO4 added. (B) Same as in A, but no copper added to the reaction. (C) Steady state GTP hydrolysis assay. Time-course of Pi released in the presence or absence of 6xHIS-Gpa1 (250 nM), GST-Arr4 (500 nM), and copper (500 nM). Results are the mean ± SEM of duplicate samples.
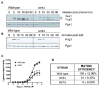
Figure 4. Arr4 is necessary for maximal transmission of the mating signal
(A) Phospho-mating MAPK time course. Yeast cells were treated with 3 μM α-factor mating pheromone and samples were removed at times indicated. MAPK activation was determined by immunoblotting using a phospho-p44/42 MAPK antibody. Pgk1, loading control. (B) Phospho-Hog1 MAPK time course. Performed as in (A) except cells were stimulated with 500 mM KCl, and probed with phospho-p38 MAPK antibody. (C) Transcription-reporter dose-response. Cells expressing a _FUS1_-lacZ reporter were treated with the indicated concentrations of mating pheromone for 90 mins. Results are the mean ± SEM for 3 individual experiments each performed in triplicate. (D) Mating efficiency assay. DC17 _MAT_α cells were mixed with BY4741 (wild-type MATa cells), arr4 deletion, or ste7 deletion as a control. Mating was performed by co-incubation of cells on nitrocellulose filters. Maximum mating efficiency of the wild-type cells was approximately 75%. Percent mating efficiency was calculated as number of diploids/total number of MATa cells. Data are mean ± SD of 3 separate experiments.
Similar articles
- Pheromone signalling in Saccharomyces cerevisiae requires the small GTP-binding protein Cdc42p and its activator CDC24.
Zhao ZS, Leung T, Manser E, Lim L. Zhao ZS, et al. Mol Cell Biol. 1995 Oct;15(10):5246-57. doi: 10.1128/MCB.15.10.5246. Mol Cell Biol. 1995. PMID: 7565673 Free PMC article. - Mu Receptors.
Herman TF, Cascella M, Muzio MR. Herman TF, et al. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31855381 Free Books & Documents. - Role for the Rho-family GTPase Cdc42 in yeast mating-pheromone signal pathway.
Simon MN, De Virgilio C, Souza B, Pringle JR, Abo A, Reed SI. Simon MN, et al. Nature. 1995 Aug 24;376(6542):702-5. doi: 10.1038/376702a0. Nature. 1995. PMID: 7651520 - Pheromone signaling pathways in yeast.
Dohlman HG, Slessareva JE. Dohlman HG, et al. Sci STKE. 2006 Dec 5;2006(364):cm6. doi: 10.1126/stke.3642006cm6. Sci STKE. 2006. PMID: 17148787 Review. - Heterotrimeric G Protein-coupled Receptor Signaling in Yeast Mating Pheromone Response.
Alvaro CG, Thorner J. Alvaro CG, et al. J Biol Chem. 2016 Apr 8;291(15):7788-95. doi: 10.1074/jbc.R116.714980. Epub 2016 Feb 23. J Biol Chem. 2016. PMID: 26907689 Free PMC article. Review.
Cited by
- GIV/Girdin transmits signals from multiple receptors by triggering trimeric G protein activation.
Garcia-Marcos M, Ghosh P, Farquhar MG. Garcia-Marcos M, et al. J Biol Chem. 2015 Mar 13;290(11):6697-704. doi: 10.1074/jbc.R114.613414. Epub 2015 Jan 20. J Biol Chem. 2015. PMID: 25605737 Free PMC article. Review. - Role of phosphatidylinositol phosphate signaling in the regulation of the filamentous-growth mitogen-activated protein kinase pathway.
Adhikari H, Cullen PJ. Adhikari H, et al. Eukaryot Cell. 2015 Apr;14(4):427-40. doi: 10.1128/EC.00013-15. Epub 2015 Feb 27. Eukaryot Cell. 2015. PMID: 25724886 Free PMC article. - Comparative Analysis of Transmembrane Regulators of the Filamentous Growth Mitogen-Activated Protein Kinase Pathway Uncovers Functional and Regulatory Differences.
Adhikari H, Caccamise LM, Pande T, Cullen PJ. Adhikari H, et al. Eukaryot Cell. 2015 Sep;14(9):868-83. doi: 10.1128/EC.00085-15. Epub 2015 Jun 26. Eukaryot Cell. 2015. PMID: 26116211 Free PMC article. - New Aspects of Invasive Growth Regulation Identified by Functional Profiling of MAPK Pathway Targets in Saccharomyces cerevisiae.
Vandermeulen MD, Cullen PJ. Vandermeulen MD, et al. Genetics. 2020 Sep;216(1):95-116. doi: 10.1534/genetics.120.303369. Epub 2020 Jul 14. Genetics. 2020. PMID: 32665277 Free PMC article. - Identification of C. elegans ASNA-1 domains and tissue requirements that differentially influence platinum sensitivity and growth control.
Raj D, Podraza-Farhanieh A, Gallego P, Kao G, Naredi P. Raj D, et al. PLoS Genet. 2022 Dec 8;18(12):e1010538. doi: 10.1371/journal.pgen.1010538. eCollection 2022 Dec. PLoS Genet. 2022. PMID: 36480541 Free PMC article.
References
- Sprang SR. G protein mechanisms: insights from structural analysis. Annu Rev Biochem. 1997;66:639–678. - PubMed
- Cismowski MJ, Takesono A, Ma C, Lizano JS, Xie X, Fuernkranz H, Lanier SM, Duzic E. Genetic screens in yeast to identify mammalian nonreceptor modulators of G-protein signaling. Nat Biotechnol. 1999;17:878–883. - PubMed
- Tall GG, Krumins AM, Gilman AG. Mammalian Ric-8A (synembryn) is a heterotrimeric Galpha protein guanine nucleotide exchange factor. J Biol Chem. 2003;278:8356–8362. - PubMed
- Afshar K, Willard FS, Colombo K, Johnston CA, McCudden CR, Siderovski DP, Gonczy P. RIC-8 is required for GPR-1/2-dependent Galpha function during asymmetric division of C. elegans embryos. Cell. 2004;119:219–230. - PubMed
- Willard FS, Kimple RJ, Siderovski DP. Return of the GDI: the GoLoco motif in cell division. Annu Rev Biochem. 2004;73:925–951. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P01 GM065533/GM/NIGMS NIH HHS/United States
- R01 GM080739/GM/NIGMS NIH HHS/United States
- F31 MH081472/MH/NIMH NIH HHS/United States
- F31-MH081472/MH/NIMH NIH HHS/United States
- GM065533/GM/NIGMS NIH HHS/United States
- R01 GM073180/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases