Mapping the genetic architecture of gene expression in human liver - PubMed (original) (raw)
doi: 10.1371/journal.pbio.0060107.
Cliona Molony, Eugene Chudin, Ke Hao, Xia Yang, Pek Y Lum, Andrew Kasarskis, Bin Zhang, Susanna Wang, Christine Suver, Jun Zhu, Joshua Millstein, Solveig Sieberts, John Lamb, Debraj GuhaThakurta, Jonathan Derry, John D Storey, Iliana Avila-Campillo, Mark J Kruger, Jason M Johnson, Carol A Rohl, Atila van Nas, Margarete Mehrabian, Thomas A Drake, Aldons J Lusis, Ryan C Smith, F Peter Guengerich, Stephen C Strom, Erin Schuetz, Thomas H Rushmore, Roger Ulrich
Affiliations
- PMID: 18462017
- PMCID: PMC2365981
- DOI: 10.1371/journal.pbio.0060107
Mapping the genetic architecture of gene expression in human liver
Eric E Schadt et al. PLoS Biol. 2008.
Abstract
Genetic variants that are associated with common human diseases do not lead directly to disease, but instead act on intermediate, molecular phenotypes that in turn induce changes in higher-order disease traits. Therefore, identifying the molecular phenotypes that vary in response to changes in DNA and that also associate with changes in disease traits has the potential to provide the functional information required to not only identify and validate the susceptibility genes that are directly affected by changes in DNA, but also to understand the molecular networks in which such genes operate and how changes in these networks lead to changes in disease traits. Toward that end, we profiled more than 39,000 transcripts and we genotyped 782,476 unique single nucleotide polymorphisms (SNPs) in more than 400 human liver samples to characterize the genetic architecture of gene expression in the human liver, a metabolically active tissue that is important in a number of common human diseases, including obesity, diabetes, and atherosclerosis. This genome-wide association study of gene expression resulted in the detection of more than 6,000 associations between SNP genotypes and liver gene expression traits, where many of the corresponding genes identified have already been implicated in a number of human diseases. The utility of these data for elucidating the causes of common human diseases is demonstrated by integrating them with genotypic and expression data from other human and mouse populations. This provides much-needed functional support for the candidate susceptibility genes being identified at a growing number of genetic loci that have been identified as key drivers of disease from genome-wide association studies of disease. By using an integrative genomics approach, we highlight how the gene RPS26 and not ERBB3 is supported by our data as the most likely susceptibility gene for a novel type 1 diabetes locus recently identified in a large-scale, genome-wide association study. We also identify SORT1 and CELSR2 as candidate susceptibility genes for a locus recently associated with coronary artery disease and plasma low-density lipoprotein cholesterol levels in the process.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures
Figure 1. Local Networks for Rps26 and Erbb3 Derived from Causal, Probabilistic Whole-Gene Networks Constructed from the Liver, Adipose, Muscle, and Brain Gene Expression Data Generated from the BXH/wt and BXC Mouse Crosses
(A) The Rps26 subnetwork includes a number of known T1D associated genes (green nodes), and RPS26 in this subnetwork is directly linked to H2-Eb1, a mouse ortholog of HLA-DRB1, a previously identified T1D susceptibility gene that is also strongly associated with a cis eSNP in the HLC (Table 2). The known T1D genes annotated by the Gene Ontology are significantly enriched in this subnetwork (Table 3). (B) The Erbb3 subnetwork is not associated with any pathways known or predicted to be involved in T1D.
Figure 2. PSRC1, CELSR2, and SORT1 Liver Expression Is Associated with a CAD Risk Allele and Plasma LDL Cholesterol Levels
The CAD risk allele for SNP rs599839 was established in a previous WTCCC study [16] (lilac panel). In the HLC, this same SNP is strongly associated with PSRC1, CELSR2, and SORT1 expression, with the CAD risk allele associated with lower relative expression (pink panel). In the BXH/wt cross designed to study metabolic traits that increase cardiovascular risk (green panel), all three of these expression traits were strongly correlated with plasma LDL cholesterol levels, a major CAD risk factor (scatter plots associated with the green panel). Given the association of these genes to plasma LDL-cholesterol levels, we examined whether rs599839 was associated with LDL cholesterol in a previously published GWAS [35] and found this SNP was significantly associated with LDL cholesterol levels, where the CAD risk allele was associated with higher LDL cholesterol levels in this cohort. Lower levels of CELSR2 and SORT1 expression were associated with the risk allele in humans, and with higher LDL cholesterol levels in mouse, making them ideal candidate susceptibility genes for the CAD and LDL cholesterol associations to this locus. On the other hand, lower levels of PSRC1 expression were associated with the risk allele in humans, but with lower LDL cholesterol levels in mouse, suggesting that PSRC1 is not the gene increasing CAD risk, but instead may be acting to protect against it.
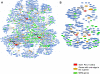
Figure 3. Local Networks for PSRC1, CELSR2, and SORT1 Derived from Causal, Probabilistic Whole-Gene Networks in Mouse and Human
(A) Mouse network for Psrc1, Celsr2, and Sort1 derived from the liver, adipose, muscle, and brain gene expression data generated from the BXH/wt and BXC mouse crosses. (B) Human network for PSRC1, CELSR2, and SORT1 derived from the HLC and from a previously published adipose and blood tissue cohort [21].
Similar articles
- Integrating genome-wide genetic variations and monocyte expression data reveals trans-regulated gene modules in humans.
Rotival M, Zeller T, Wild PS, Maouche S, Szymczak S, Schillert A, Castagné R, Deiseroth A, Proust C, Brocheton J, Godefroy T, Perret C, Germain M, Eleftheriadis M, Sinning CR, Schnabel RB, Lubos E, Lackner KJ, Rossmann H, Münzel T, Rendon A; Cardiogenics Consortium; Erdmann J, Deloukas P, Hengstenberg C, Diemert P, Montalescot G, Ouwehand WH, Samani NJ, Schunkert H, Tregouet DA, Ziegler A, Goodall AH, Cambien F, Tiret L, Blankenberg S. Rotival M, et al. PLoS Genet. 2011 Dec;7(12):e1002367. doi: 10.1371/journal.pgen.1002367. Epub 2011 Dec 1. PLoS Genet. 2011. PMID: 22144904 Free PMC article. - Integrative analysis of liver-specific non-coding regulatory SNPs associated with the risk of coronary artery disease.
Selvarajan I, Toropainen A, Garske KM, López Rodríguez M, Ko A, Miao Z, Kaminska D, Õunap K, Örd T, Ravindran A, Liu OH, Moreau PR, Jawahar Deen A, Männistö V, Pan C, Levonen AL, Lusis AJ, Heikkinen S, Romanoski CE, Pihlajamäki J, Pajukanta P, Kaikkonen MU. Selvarajan I, et al. Am J Hum Genet. 2021 Mar 4;108(3):411-430. doi: 10.1016/j.ajhg.2021.02.006. Epub 2021 Feb 23. Am J Hum Genet. 2021. PMID: 33626337 Free PMC article. - Integromic analysis of genetic variation and gene expression identifies networks for cardiovascular disease phenotypes.
Yao C, Chen BH, Joehanes R, Otlu B, Zhang X, Liu C, Huan T, Tastan O, Cupples LA, Meigs JB, Fox CS, Freedman JE, Courchesne P, O'Donnell CJ, Munson PJ, Keles S, Levy D. Yao C, et al. Circulation. 2015 Feb 10;131(6):536-49. doi: 10.1161/CIRCULATIONAHA.114.010696. Epub 2014 Dec 22. Circulation. 2015. PMID: 25533967 Free PMC article. - Genome-wide studies of gene expression relevant to coronary artery disease.
Hsu J, Smith JD. Hsu J, et al. Curr Opin Cardiol. 2012 May;27(3):210-3. doi: 10.1097/HCO.0b013e3283522198. Curr Opin Cardiol. 2012. PMID: 22476029 Free PMC article. Review. - Functional genomics and assays of regulatory activity detect mechanisms at loci for lipid traits and coronary artery disease.
Roman TS, Mohlke KL. Roman TS, et al. Curr Opin Genet Dev. 2018 Jun;50:52-59. doi: 10.1016/j.gde.2018.02.004. Epub 2018 Feb 20. Curr Opin Genet Dev. 2018. PMID: 29471259 Free PMC article. Review.
Cited by
- Genome-wide significant loci: how important are they? Systems genetics to understand heritability of coronary artery disease and other common complex disorders.
Björkegren JLM, Kovacic JC, Dudley JT, Schadt EE. Björkegren JLM, et al. J Am Coll Cardiol. 2015 Mar 3;65(8):830-845. doi: 10.1016/j.jacc.2014.12.033. J Am Coll Cardiol. 2015. PMID: 25720628 Free PMC article. Review. - Candidate gene studies of a promising intermediate phenotype: failure to replicate.
Hart AB, de Wit H, Palmer AA. Hart AB, et al. Neuropsychopharmacology. 2013 Apr;38(5):802-16. doi: 10.1038/npp.2012.245. Epub 2012 Dec 3. Neuropsychopharmacology. 2013. PMID: 23303064 Free PMC article. Clinical Trial. - Genetic risk factors for type 2 diabetes: a trans-regulatory genetic architecture?
Elbein SC, Gamazon ER, Das SK, Rasouli N, Kern PA, Cox NJ. Elbein SC, et al. Am J Hum Genet. 2012 Sep 7;91(3):466-77. doi: 10.1016/j.ajhg.2012.08.002. Am J Hum Genet. 2012. PMID: 22958899 Free PMC article. - Multiple sclerosis risk variant HLA-DRB1*1501 associates with high expression of DRB1 gene in different human populations.
Alcina A, Abad-Grau Mdel M, Fedetz M, Izquierdo G, Lucas M, Fernández O, Ndagire D, Catalá-Rabasa A, Ruiz A, Gayán J, Delgado C, Arnal C, Matesanz F. Alcina A, et al. PLoS One. 2012;7(1):e29819. doi: 10.1371/journal.pone.0029819. Epub 2012 Jan 13. PLoS One. 2012. PMID: 22253788 Free PMC article.
References
- Edwards AO, Ritter R, 3rd, Abel KJ, Manning A, Panhuysen C, et al. Complement factor H polymorphism and age-related macular degeneration. Science. 2005;308:421–424. - PubMed
- Haines JL, Hauser MA, Schmidt S, Scott WK, Olson LM, et al. Complement factor H variant increases the risk of age-related macular degeneration. Science. 2005;308:419–421. - PubMed
- Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T, et al. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science. 2007;316:1491–1493. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA090426/CA/NCI NIH HHS/United States
- HL28481/HL/NHLBI NIH HHS/United States
- P01 HL028481/HL/NHLBI NIH HHS/United States
- R37 CA090426/CA/NCI NIH HHS/United States
- HL30568/HL/NHLBI NIH HHS/United States
- R37CA090426/CA/NCI NIH HHS/United States
- P01 HL030568/HL/NHLBI NIH HHS/United States
- DK072206/DK/NIDDK NIH HHS/United States
- R01 DK072206/DK/NIDDK NIH HHS/United States
- P30 ES000267/ES/NIEHS NIH HHS/United States
- P30ES000267/ES/NIEHS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials