Mgr3p and Mgr1p are adaptors for the mitochondrial i-AAA protease complex - PubMed (original) (raw)
Mgr3p and Mgr1p are adaptors for the mitochondrial i-AAA protease complex
Cory D Dunn et al. Mol Biol Cell. 2008 Dec.
Abstract
By screening yeast knockouts for their dependence upon the mitochondrial genome, we identified Mgr3p, a protein that associates with the i-AAA protease complex in the mitochondrial inner membrane. Mgr3p and Mgr1p, another i-AAA-interacting protein, form a subcomplex that bind to the i-AAA subunit Yme1p. We find that loss of Mgr3p, like the lack of Mgr1p, reduces proteolysis by Yme1p. Mgr3p and Mgr1p can bind substrate even in the absence of Yme1p, and both proteins are needed for maximal binding of an unfolded substrate by the i-AAA complex. We speculate that Mgr3p and Mgr1p function in an adaptor complex that targets substrates to the i-AAA protease for degradation.
Figures
Figure 1.
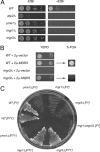
Mgr3p is required for mtDNA-independent growth. (A) Yeast cells lacking Mgr3p are petite-negative. Wild-type strain BY4742, _atp2_Δ strain RJ2092, _yme1_Δ strain RJ2051, _mgr1_Δ strain RJ2058, and _mgr3_Δ strain RJ2122 were grown for 23 h in YEPD medium containing 25 μg/ml ethidium bromide (+EtBr) or for 8 h in YEPD medium (−EtBr) at 30°C. Then, 10-fold serial dilutions were made from cultures at OD600 of 0.02, and 4 μl of each dilution was spotted onto YEPD medium with or without EtBr. Plates were incubated for 2d at 30°C. (B) _mgr3_Δ lethality upon EtBr treatment is due to mtDNA loss. Wild-type strain BY4742 and _mgr3_Δ strain RJ2122 were transformed with empty vector pRS426 or a 2μ plasmid carrying the MGR3 gene. The resulting transformants were grown overnight in SD medium lacking uracil and then transferred to YEPD medium containing EtBr and grown for ∼24 h at 30°C. Cells were diluted to OD600 of 0.02 and 0.002, and 4 μl of each dilution were spotted to YEPD medium lacking EtBr and grown for a further 2 d at 30°C (YEPD). Next, the resulting rho0 WT and _mgr3_Δ strains containing the 2μ-MGR3 plasmid were grown in liquid medium for a further 1 d in YEPD medium and then diluted to OD600 of 0.02. Four microliters of each dilution was spotted to complete medium containing 5-fluoroorotic acid and grown for a further 1 d at 30°C (5-FOA). (C) The petite-negative phenotype of _mgr3_Δ is background specific. The following strains were patched twice to YEPD medium containing 25 μg/ml EtBr and then struck to YEPD medium for 2 d at 30°C. In Winston strain background FY (Winston et al., 1995): parental strain RJ1962 [_WT_], RJ1974 [_yme1_Δ], RJ1961 [_mgr1_Δ], RJ2150 [_mgr3_Δ], RJ2152 [_mgr1_Δ_mgr3_Δ]; and in Thorsness strain background PTY: strain PTY33 [_WT_], strain RJ2001 [_mgr1_Δ], strain RJ2326 [_mgr3_Δ], and strain RJ2040 [_yme1_Δ].
Figure 2.
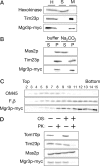
Mgr3p-myc is a mitochondrial inner membrane protein with its carboxy terminus exposed to the IMS. (A) Mgr3p is a mitochondrial protein. Strain RJ2165, which expresses Mgr3p-myc, was homogenized (H) and separated into a mitochondrial pellet fraction (M) and crude cytosolic supernatant fraction (S) by centrifugation. Cell equivalents were then subjected to immunoblotting and decorated with antibodies to hexokinase, Tim23p, or the myc epitope (Mgr3p-myc). (B) Mgr3p is an integral membrane protein. Mitochondria from strain RJ2165 were resuspended in buffer or 0.1 M Na2CO3. After centrifugation, equal amounts of the supernatant (S) or pellet (P) fractions were analyzed by immunoblotting using antibodies to Mas2p, Tim23p, or the myc epitope (Mgr3p-myc). (C) Mgr3p is associated with the inner membrane. Mitochondria containing Mgr3p-myc were sonicated and membrane vesicles separated on sucrose step gradients. Fractions were analyzed by immunoblotting with antibodies to OM45, F1β, or the myc epitope (Mgr3p-myc). Fraction 2 represents the top of the gradient. (D) The carboxy terminus of Mgr3p-myc faces the IMS. Intact mitochondria, or mitochondria with their outer membranes disrupted by OS, were digested with 50 μg/ml proteinase K (PK) for 30 min, and then analyzed by immunoblotting using antibodies to Tom70p, Tim23p, Mas2p, or the myc epitope (Mgr3p-myc).
Figure 3.
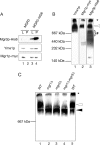
Mgr3p is part of the i-AAA machinery. (A) Yme1p and Mgr1p coprecipitate with Mgr3p. Mitochondria isolated from cells expressing both Mgr1p-myc and Mgr3p-His6 (MGR3-HIS6; strain RJ2161) were solubilized, and the Mgr3p-His6 protein precipitated with Ni-NTA agarose. Ten percent of the starting lysate (L) and 100% of the pellet (P) were immunoblotted using antibodies to the His6 tag (Mgr3p-His6), the myc tag (Mgr1p-myc), or to Yme1p. As a control, mitochondria from cells expressing Mgr1p-myc, but lacking Mgr3p-His6 (MGR3; RJ1970), were used. (B) Mgr1p, Mgr3p, and Yme1p comigrate during BN-PAGE. Mitochondria were isolated from RJ2161, expressing Mgr3p-His6 and Mgr1p-myc. After solubilization in digitonin buffer, protein complexes were separated by BN-PAGE on a 3–13% gradient gel, and separate lanes from the same gel were immunoblotted with antiserum to Yme1p, the myc epitope of Mgr1p-myc, or the His6 epitope of Mgr3p-His6. (C) Loss of Mgr3p alters the distribution of Yme1p in the i-AAA complexes. Mitochondria were isolated from WT (RJ1962), _mgr1_Δ (RJ1961), _mgr3_Δ (RJ2150), and _mgr1_Δ _mgr3_Δ (RJ2152) cells and solubilized. Protein complexes were separated on a 3 to 10.5% gradient blue native gel and analyzed with antibodies to Yme1p. Molecular weight standards run in an adjacent lane are indicated. The white and black arrows denote the larger and smaller forms of the i-AAA complex, respectively. Asterisk (*) marks protein not resolvable by BN-PAGE.
Figure 4.
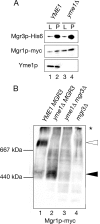
Mgr3p and Mgr1p form a subcomplex in the absence of Yme1p. (A) Mgr3p and Mgr1p coprecipitate in mitochondria lacking Yme1p. Mitochondria isolated from YME1 (RJ2161) and _yme1_Δ (RJ2183) cells, which also express Mgr3p-His6 and Mgr1p-myc, were solubilized and Mgr3p-His6 precipitated using Ni-NTA beads. Ten percent of the starting lysate (L) and 100% of the pellet (P) were immunoblotted using antibodies to the His6 tag (Mgr3p-His6), the myc tag (Mgr1p-myc), or to Yme1p. (B) Mgr1p migrates at ∼450-kDa without Yme1p. Mitochondria were isolated from a series of strains expressing Mgr1p-myc: otherwise wild-type strain RJ1970, _yme1_Δ strain RJ1972, _yme1_Δ _mgr3_Δ strain RJ2214, and _mgr3_Δ strain RJ2148. After solubilization, protein complexes were separated by BN-PAGE on a 3–13% gradient gel and immunoblotted with antiserum recognizing Mgr1p-myc. The white arrow marks the Mgr1p-containing complex when Yme1p and Mgr3p are present, and the black arrow marks the ∼450-kDa complex in Yme1p's absence.
Figure 5.
Mgr1p links the Mgr3p-Mgr1p subcomplex to Yme1p. (A) Mgr1p and Yme1p coprecipitate from mitochondria lacking Mgr3p. Mitochondria from MGR3 (RJ1970) and _mgr3_Δ (RJ2148) cells expressing Mgr1p-myc were solubilized, and Mgr1p-myc was precipitated using beads coupled to anti-myc antibodies. Ten percent of the starting lysate (L) and 100% of the pellet (P) were immunoblotted using antibodies to the myc tag (Mgr1p-myc) or to Yme1p. (B) Mgr3p and Yme1p do not interact in the absence of Mgr1p. Mitochondria from MGR1 (RJ2175) and _mgr1_Δ (RJ2186) cells expressing Mgr3p-His6 were solubilized, and Mgr3p-His6 precipitated with Ni-NTA beads. Ten percent of L and 100% of P were immunoblotted using antibodies to the myc tag (Mgr3p-His6) or to Yme1p.
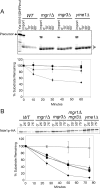
Figure 6.
Mgr3p is required for full activity of the i-AAA protease. (A) Degradation of Yta10(161)-DHFRMUT is inhibited in _mgr3_Δ mitochondria. 35S-labeled Yta10(161)- DHFRMUT was imported into mitochondria isolated from WT (RJ1962; diamonds), _mgr1_Δ (RJ1961; squares), _mgr3_Δ (RJ2150; triangles), and _yme1_Δ (RJ1974; circles) and incubated for the indicated times. Mitochondria from each time point were subjected to SDS-PAGE and analyzed by phosphorimaging. The two processed forms of Yta10(161)-DHFRMUT (asterisk) were quantified and plotted. Values shown are the average of three independent experiments. Error bars represent SD. (B) Cells lacking Mgr3p are defective in Nde1p-HA turnover. WT (RJ2077; diamonds), _mgr1_Δ (RJ2078; squares), _mgr3_Δ (RJ2153; triangles), _mgr1_Δ_mgr3_Δ (RJ2163; circles), and _yme1_Δ (RJ2079; asterisks) strains, each expressing Nde1p-HA, were grown to mid-log phase, and then protein synthesis was inhibited with cycloheximide. At the indicated times, cells were lysed, and Nde1p-HA was assayed by immunoblotting with antibodies to the HA-epitope. Values shown are the average of three independent experiments. Error bars represent SD.
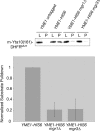
Figure 7.
Yme1p binding to substrate is reduced in mitochondria lacking Mgr1p or Mgr3p. 35S-labeled Yta10(161)-DHFRMUT was imported into mitochondria isolated from otherwise wild-type strain RJ2195, _mgr1_Δ strain RJ2199, or _mgr3_Δ strain RJ2200, each expressing Yme1p-His6. Mitochondria from strain RJ1962, which lacks Yme1p-His6, were used as a control. After incubation, mitochondria were solubilized and Yme1p-His6 precipitated using Ni-NTA agarose. Ten percent of the starting lysate (L) and 100% of the pellet (P) were subjected to SDS-PAGE, and the 35S-labeled, processed substrate [m-Yta10(161)-DHFRMUT] that coprecipitated with Yme1p-His6 was detected by phosphorimaging. After subtraction of nonspecific precipitation from control mitochondria, the amount of m-Yta10(161)-DHFRMUT in the pellet from YME1-HIS6 mitochondria was set to 1, and error bars represent the SD of five independent experiments.
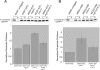
Figure 8.
Mgr1p and Mgr3p bind substrate in the absence of Yme1p. (A) Mgr1p binding. 35S-labeled Yta10(161)-DHFRMUT was imported into mitochondria isolated from the following strains expressing Mgr1p-myc: otherwise, wild-type strain RJ1970, _mgr3_Δ strain RJ2148, _yme1_Δ strain RJ1972, or _yme1_Δ _mgr3_Δ strain RJ2214. After incubation, mitochondria were solubilized, and Mgr1p-myc was precipitated. Ten percent of the mitochondrial lysate (L) and 100% of the pellet fraction (P) were subjected to SDS-PAGE, and the coprecipitated, processed m-Yta10(161)-DHFRMUT, and visualized by phosphorimaging. The amount of substrate in the pellet fraction from MGR1-MYC mitochondria was set to 1 after subtraction of nonspecific substrate precipitation from mitochondria isolated from control strain RJ1962, and error bars represent the SD of three independent experiments. (B) Mgr3p binding. Yta10(161)-DHFRMUT was imported into mitochondria isolated from the following strains expressing Mgr3p-His6: otherwise, wild-type strain RJ2161, _yme1_Δ strain RJ2183, and _yme1_Δ _mgr1_Δ strain RJ2217. Strains expressing Mgr1p contained the myc-tagged variant. The relative amount of m-Yta10(161)-DHFRMUT precipitated by Mgr3p-His6 was graphed, with the amount in the pellet fraction from MGR3-H6 mitochondria set to 1 after subtraction of nonspecific precipitation from RJ1962 control mitochondria. Error bars represent the SD of three independent experiments.
Similar articles
- A genomewide screen for petite-negative yeast strains yields a new subunit of the i-AAA protease complex.
Dunn CD, Lee MS, Spencer FA, Jensen RE. Dunn CD, et al. Mol Biol Cell. 2006 Jan;17(1):213-26. doi: 10.1091/mbc.e05-06-0585. Epub 2005 Nov 2. Mol Biol Cell. 2006. PMID: 16267274 Free PMC article. - Substrate recognition by AAA+ ATPases: distinct substrate binding modes in ATP-dependent protease Yme1 of the mitochondrial intermembrane space.
Graef M, Seewald G, Langer T. Graef M, et al. Mol Cell Biol. 2007 Apr;27(7):2476-85. doi: 10.1128/MCB.01721-06. Epub 2007 Jan 29. Mol Cell Biol. 2007. PMID: 17261594 Free PMC article. - A AAA ATPase Cdc48 with a cofactor Ubx2 facilitates ubiquitylation of a mitochondrial fusion-promoting factor Fzo1 for proteasomal degradation.
Nahar S, Chowdhury A, Ogura T, Esaki M. Nahar S, et al. J Biochem. 2020 Mar 1;167(3):279-286. doi: 10.1093/jb/mvz104. J Biochem. 2020. PMID: 31804690 - ATP-dependent proteases controlling mitochondrial function in the yeast Saccharomyces cerevisiae.
Van Dyck L, Langer T. Van Dyck L, et al. Cell Mol Life Sci. 1999 Nov 30;56(9-10):825-42. doi: 10.1007/s000180050029. Cell Mol Life Sci. 1999. PMID: 11212342 Free PMC article. Review. - Mitochondrial AAA proteases--towards a molecular understanding of membrane-bound proteolytic machines.
Gerdes F, Tatsuta T, Langer T. Gerdes F, et al. Biochim Biophys Acta. 2012 Jan;1823(1):49-55. doi: 10.1016/j.bbamcr.2011.09.015. Epub 2011 Oct 6. Biochim Biophys Acta. 2012. PMID: 22001671 Review.
Cited by
- Current Understanding of Temperature Stress-Responsive Chloroplast FtsH Metalloproteases.
Luo S, Kim C. Luo S, et al. Int J Mol Sci. 2021 Nov 9;22(22):12106. doi: 10.3390/ijms222212106. Int J Mol Sci. 2021. PMID: 34829988 Free PMC article. Review. - Multifunctional Mitochondrial AAA Proteases.
Glynn SE. Glynn SE. Front Mol Biosci. 2017 May 22;4:34. doi: 10.3389/fmolb.2017.00034. eCollection 2017. Front Mol Biosci. 2017. PMID: 28589125 Free PMC article. Review. - Quality control of mitochondrial proteostasis.
Baker MJ, Tatsuta T, Langer T. Baker MJ, et al. Cold Spring Harb Perspect Biol. 2011 Jul 1;3(7):a007559. doi: 10.1101/cshperspect.a007559. Cold Spring Harb Perspect Biol. 2011. PMID: 21628427 Free PMC article. Review. - Barth syndrome mutations that cause tafazzin complex lability.
Claypool SM, Whited K, Srijumnong S, Han X, Koehler CM. Claypool SM, et al. J Cell Biol. 2011 Feb 7;192(3):447-62. doi: 10.1083/jcb.201008177. J Cell Biol. 2011. PMID: 21300850 Free PMC article. - Stress-regulated translational attenuation adapts mitochondrial protein import through Tim17A degradation.
Rainbolt TK, Atanassova N, Genereux JC, Wiseman RL. Rainbolt TK, et al. Cell Metab. 2013 Dec 3;18(6):908-19. doi: 10.1016/j.cmet.2013.11.006. Cell Metab. 2013. PMID: 24315374 Free PMC article.
References
- Adams A., Gottschling D., Kaiser C., Stearns T. Methods in yeast genetics. Plainview, NY: Cold Spring Harbor Laboratory Press; 1997.
- Arnold I., Wagner-Ecker M., Ansorge W., Langer T. Evidence for a novel mitochondria-to-nucleus signalling pathway in respiring cells lacking i-AAA protease and the ABC-transporter Mdl1. Gene. 2006;367:74–88. - PubMed
- Attardi G., Schatz G. Biogenesis of mitochondria. Annu. Rev. Cell Biol. 1988;4:289–333. - PubMed
- Augustin S., Nolden M., Muller S., Hardt O., Arnold I., Langer T. Characterization of peptides released from mitochondria: evidence for constant proteolysis and peptide efflux. J. Biol. Chem. 2005;280:2691–2699. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM054021/GM/NIGMS NIH HHS/United States
- T32 GM007445/GM/NIGMS NIH HHS/United States
- R01 GM-54021/GM/NIGMS NIH HHS/United States
- T32GM07445/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases