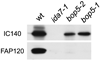A novel ankyrin-repeat protein interacts with the regulatory proteins of inner arm dynein f (I1) of Chlamydomonas reinhardtii - PubMed (original) (raw)
A novel ankyrin-repeat protein interacts with the regulatory proteins of inner arm dynein f (I1) of Chlamydomonas reinhardtii
Kazuho Ikeda et al. Cell Motil Cytoskeleton. 2009 Aug.
Abstract
How ciliary and flagellar motility is regulated is a challenging problem. The flagellar movement in Chlamydomonas reinhardtii is in part regulated by phosphorylation of a 138 kD intermediate chain (IC138) of inner arm dynein f (also called I1). In the present study, we found that the axoneme of mutants lacking dynein f lacks a novel protein having ankyrin repeat motifs, registered as FAP120 in the flagellar proteome database. FAP120 is also missing or decreased in the axonemes of bop5, a mutant that has a mutation in the structural gene of IC138 but assembles the dynein f complex. Intriguingly, the amounts of FAP120 in the axonemes of different alleles of bop5 and several dynein f-lacking mutants roughly parallel their contents of IC138. These results suggest a weak but stoichiometric interaction between FAP120 and IC138. We propose that FAP120 functions in the regulatoryprocess as part of a protein complex involving IC138. Cell Motil. Cytoskeleton 2008. (c) 2008 Wiley-Liss, Inc.
Figures
Fig. 1
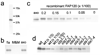
Identification of the 42 kD protein as FAP120. (a)–(c) Separation of the 42 kD protein by 2D electrophoresis. Silver-stained gel patterns of 0.1% Sarkosyl extracts from wild-type (a) and ida1 (b) axonemes. A region covering approximately 30–60 kD is shown. A spot at ~42 kD indicated by the arrowhead is missing in the ida1 pattern (b). Duplicate gel of wild-type gel pattern (a) was transferred to a PVDF membrane and analyzed by immunoblotting (c). The 42 kD spot reacted with the antibody. This band was cut out from the gel and analyzed by mass spectrometry. (d) The determined FAP120 cDNA sequence was analyzed using the programs SMART (
http://smart.embl-heidelberg.de/
) and Coils (
http://www.ch.embnet.org/software/COILS\_form.html
). The analysis indicates that FAP120, a 294-amino acid protein with a molecular weight of 31861.12 and a pI of 4.94, contains four ankyrin repeat motifs in its N terminus half and a putative coiled-coil structure in its C terminus half.
Fig. 2
Immunoblot analysis of wild-type and mutant axonemes using the FAP120-specific antibody. (a) Western analysis of wild-type axonemes with an affinity purified FAP120 antibody. Only a single band of ~42 kDa is recognized. (b) Immunoblot analysis of isolated flagella (fla), the membrane and matrix fraction (M&M) and axonemes (axo) prepared from the same amount of flagella. FAP120 is barely detectable in the membrane and matrix fraction. (c) Estimation of the amount of FAP120 present in the wild-type axoneme by immunoblot analysis using a dilution series of recombinant FAP120 as a standard. Comparison of the immuno-blot signal intensities of FAP120 in 3 µg of the wild-type axoneme (axo) and the diluted series of recombinant FAP120 (diluted from 6 ng to 0 ng; correponding to 0.2% to 0% (weight/weight) of the total mass of the axoneme run on lane 1) yields an estimate of FAP120 to be ~0.18% of the total axonemal proteins. The number above each lane indicates the dilution factor expressed as the percentage of the total weight of the axoneme (axo.)(× 1/100). Because the recombinant FAP120 has a 6×His tag and an additional linker, its mobility on the western blot differs from that of the native FAP120 in the axoneme. (d) Quantification of FAP120 present in the mutant axonemes by immunoblot analysis. The amount of FAP120 in the axonemes of mutants lacking dynein f was estimated by densitometry using a series of diluted wild-type axonemes as a standard (lane 1 (wt), 100%; lane 2 (wt × 1/3), 33%; lane 3 (wt × 1/10), 10%; and lane 4 (wt × 1/30), 3.3%). FAP120 is present in mutant axonemes in small amounts ranging from < 1% to ~18% of the wild-type level.
Fig. 3
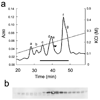
FAP120 does not co-elute with dynein f from ion-exchange column. (a) The elution pattern of the crude dynein extract from oda1 axonemes by anion-exchange chromatography on a Mono Q column (solid line, absorbance at 280 nm; broken line, KCl conc.). a–g, the peak fractions that contain the designated inner arm dynein complex as a major component (Kagami and Kamiya, 1992). The horizontal bar indicates the fractions for which immuno-blot analysis was carried out. (b) Immuno-blot detection of FAP120 in the indicated fractions. The peak of FAP120 distribution (marked by the arrowhead in a) does not correspond to any particular dynein peak.
Fig. 4
Immunofluorescence microscopy of wild-type and ida1 cells. Top, differential interference contrast images. Bottom, indirect immunofluorescence localization of FAP120. Fluorescent signal is observed along the length of the axonemes of wild-type, whereas it is only negligibly present in the ida1 axoneme.
Fig. 5
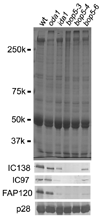
The amounts of IC138 roughly correlate with those of FAP120 in alleles of bop5 mutants Immunoblot analyses on the axonemes of three bop5 alleles (bop5-3, bop5-4, bop5-6), mutants that have mutations in the IC138 gene, were performed using antibodies against IC138, IC97, p28 and FAP120. p28 is a subunit of single-headed inner arm dyneins and used as a control. Two alleles of bop5 (bop5-3, bop5-4) have extremely small amounts of FAP120 while one allele (bop5-6) has a reduced amount of FAP120. Mutant strains that have small amounts of IC138 such as bop5-6 and ida1 also have small amounts of FAP120 and IC97 in their flagella. These observations suggest that FAP120, as well as IC97, interacts with IC138.
Fig. 6
bop5-1, a mutant expressing a truncated IC138 (Hendrickson et al., 2004), has no FAP120 in its flagella. Immunoblot analysis of axonemes from wild-type, ida7-1, bop5-1 and bop5-2 (6F5) using the anti-FAP120 antibody and the anti-IC140 antibody. Bop5-1 and bop5-2 axonemes contain IC140 but have only negligible amounts of FAP120. The Tropix detection system was used for this blot.
Similar articles
- Composition and function of ciliary inner-dynein-arm subunits studied in Chlamydomonas reinhardtii.
Yamamoto R, Hwang J, Ishikawa T, Kon T, Sale WS. Yamamoto R, et al. Cytoskeleton (Hoboken). 2021 Mar;78(3):77-96. doi: 10.1002/cm.21662. Epub 2021 Apr 28. Cytoskeleton (Hoboken). 2021. PMID: 33876572 Free PMC article. Review. - IC138 defines a subdomain at the base of the I1 dynein that regulates microtubule sliding and flagellar motility.
Bower R, VanderWaal K, O'Toole E, Fox L, Perrone C, Mueller J, Wirschell M, Kamiya R, Sale WS, Porter ME. Bower R, et al. Mol Biol Cell. 2009 Jul;20(13):3055-63. doi: 10.1091/mbc.e09-04-0277. Epub 2009 May 6. Mol Biol Cell. 2009. PMID: 19420135 Free PMC article. - bop5 Mutations reveal new roles for the IC138 phosphoprotein in the regulation of flagellar motility and asymmetric waveforms.
VanderWaal KE, Yamamoto R, Wakabayashi K, Fox L, Kamiya R, Dutcher SK, Bayly PV, Sale WS, Porter ME. VanderWaal KE, et al. Mol Biol Cell. 2011 Aug 15;22(16):2862-74. doi: 10.1091/mbc.E11-03-0270. Epub 2011 Jun 22. Mol Biol Cell. 2011. PMID: 21697502 Free PMC article. - The IC138 and IC140 intermediate chains of the I1 axonemal dynein complex bind directly to tubulin.
Hendrickson TW, Goss JL, Seaton CA, Rohrs HW. Hendrickson TW, et al. Biochim Biophys Acta. 2013 Dec;1833(12):3265-3271. doi: 10.1016/j.bbamcr.2013.09.011. Epub 2013 Sep 27. Biochim Biophys Acta. 2013. PMID: 24080090 - Functional and structural significance of the inner-arm-dynein subspecies d in ciliary motility.
Yamamoto R, Kon T. Yamamoto R, et al. Cytoskeleton (Hoboken). 2024 Nov;81(11):569-577. doi: 10.1002/cm.21828. Epub 2024 Jan 12. Cytoskeleton (Hoboken). 2024. PMID: 38214410 Review.
Cited by
- The ciliary inner dynein arm, I1 dynein, is assembled in the cytoplasm and transported by IFT before axonemal docking.
Viswanadha R, Hunter EL, Yamamoto R, Wirschell M, Alford LM, Dutcher SK, Sale WS. Viswanadha R, et al. Cytoskeleton (Hoboken). 2014 Oct;71(10):573-86. doi: 10.1002/cm.21192. Epub 2014 Oct 30. Cytoskeleton (Hoboken). 2014. PMID: 25252184 Free PMC article. - Cryoelectron tomography reveals doublet-specific structures and unique interactions in the I1 dynein.
Heuser T, Barber CF, Lin J, Krell J, Rebesco M, Porter ME, Nicastro D. Heuser T, et al. Proc Natl Acad Sci U S A. 2012 Jul 24;109(30):E2067-76. doi: 10.1073/pnas.1120690109. Epub 2012 Jun 25. Proc Natl Acad Sci U S A. 2012. PMID: 22733763 Free PMC article. - Composition and function of ciliary inner-dynein-arm subunits studied in Chlamydomonas reinhardtii.
Yamamoto R, Hwang J, Ishikawa T, Kon T, Sale WS. Yamamoto R, et al. Cytoskeleton (Hoboken). 2021 Mar;78(3):77-96. doi: 10.1002/cm.21662. Epub 2021 Apr 28. Cytoskeleton (Hoboken). 2021. PMID: 33876572 Free PMC article. Review. - IC138 defines a subdomain at the base of the I1 dynein that regulates microtubule sliding and flagellar motility.
Bower R, VanderWaal K, O'Toole E, Fox L, Perrone C, Mueller J, Wirschell M, Kamiya R, Sale WS, Porter ME. Bower R, et al. Mol Biol Cell. 2009 Jul;20(13):3055-63. doi: 10.1091/mbc.e09-04-0277. Epub 2009 May 6. Mol Biol Cell. 2009. PMID: 19420135 Free PMC article. - A dynein-associated photoreceptor protein prevents ciliary acclimation to blue light.
Kutomi O, Yamamoto R, Hirose K, Mizuno K, Nakagiri Y, Imai H, Noga A, Obbineni JM, Zimmermann N, Nakajima M, Shibata D, Shibata M, Shiba K, Kita M, Kigoshi H, Tanaka Y, Yamasaki Y, Asahina Y, Song C, Nomura M, Nomura M, Nakajima A, Nakachi M, Yamada L, Nakazawa S, Sawada H, Murata K, Mitsuoka K, Ishikawa T, Wakabayashi KI, Kon T, Inaba K. Kutomi O, et al. Sci Adv. 2021 Feb 26;7(9):eabf3621. doi: 10.1126/sciadv.abf3621. Print 2021 Feb. Sci Adv. 2021. PMID: 33637535 Free PMC article.
References
- Brokaw CJ, Kamiya R. Bending patterns of Chlamydomonas flagella: IV. Mutants with defects in inner and outer dynein arms indicate differences in dynein arm function. Cell Motil Cytoskel. 1987;8:68–75. - PubMed
- DiBella LM, Smith EF, Patel-King RS, Wakabayashi K, King SM. A novel Tctex2-related light chain is required for stability of inner dynein arm I1 and motor function in the Chlamydomonas flagellum. J Biol Chem. 2004;279:21666–21676. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM051173/GM/NIGMS NIH HHS/United States
- GM55667/GM/NIGMS NIH HHS/United States
- GM51173/GM/NIGMS NIH HHS/United States
- R37 GM055667/GM/NIGMS NIH HHS/United States
- R37 GM051173/GM/NIGMS NIH HHS/United States
- R01 GM055667/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources