Network features of the mammalian circadian clock - PubMed (original) (raw)
Network features of the mammalian circadian clock
Julie E Baggs et al. PLoS Biol. 2009.
Abstract
The mammalian circadian clock is a cell-autonomous system that drives oscillations in behavior and physiology in anticipation of daily environmental change. To assess the robustness of a human molecular clock, we systematically depleted known clock components and observed that circadian oscillations are maintained over a wide range of disruptions. We developed a novel strategy termed Gene Dosage Network Analysis (GDNA) in which small interfering RNA (siRNA)-induced dose-dependent changes in gene expression were used to build gene association networks consistent with known biochemical constraints. The use of multiple doses powered the analysis to uncover several novel network features of the circadian clock, including proportional responses and signal propagation through interacting genetic modules. We also observed several examples where a gene is up-regulated following knockdown of its paralog, suggesting the clock network utilizes active compensatory mechanisms rather than simple redundancy to confer robustness and maintain function. We propose that these network features act in concert as a genetic buffering system to maintain clock function in the face of genetic and environmental perturbation.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures
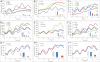
Figure 1. Functional Effects on Oscillations Following Knockdown of Circadian Clock Components in U-2 OS Cells
U-2 OS cells were transfected with pools of four to five siRNAs for each gene, and oscillations in _Bmal1_-luciferase were measured for 5 d following synchronization with dexamethasone. Knockdown of each component was validated by quantitative RT-PCR from RNA isolated from a replicate sample. The relative amount of expression of each gene compared to the negative siRNA control sample is shown (insets). Results are representative of three independent biological replicates.
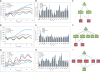
Figure 2. GDNA Following Genetic Perturbation of Bmal1, Clock, and Per1
(A, D, and G) Oscillations in _Bmal1_-luciferase were measured from cells transfected with increasing amounts of siRNAs targeting Bmal1, Clock, or Per1. (B, E, and H) RNA was isolated from replicate samples collected at time 0 (before synchronization) and expression of clock genes was determined by quantitative real time PCR. (C, F, and I) GDNAs were generated for siBmal1 (C), siClock (F), and siPer1(I) using expression data from each knockdown condition. Edges between genes were determined using nonparametric Pearson correlation (_p_-value < 0.10) and biochemical constraints, and gene expression changes are denoted as increases (red) and decreases (green). The gene being depleted is located at the top of network, with first order responses below as restricted by biochemistry, i.e. genes that decrease when Clock and Bmal1 activators are decreased (C and F) and genes that are increased with the Per1 repressor is decreased (I). Second-order responses are defined as those with correlated edges with the first-order responders. Black lines indicated published edges, and blue lines denote unpublished relationships.
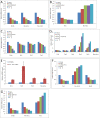
Figure 3. Signal Propagation of Proportional Gene Expression Changes Is Observed upon Dose- Dependent Knockdown of Clock Components
(A–C) Examples of linear, proportional gene expression changes following knockdown of Bmal1 (A), Per1 (B), and Clock (C). (D and E) Disproportional responses are observed in Per2 gene expression following knockdown of Cry1 and Cry2 (D), and similar changes are observed in MEFs derived from Cry1/Cry2 double-knockout mice (E). (F) Signal propagation following knockdown of Per1 through a Repressor/Repressor module where knockdown of Per1 leads to increase of _Rev-erb_s, which in turn causes a decrease in Bmal1 expression. (G) An Activator/Repressor module relays the signal following knockdown of Bmal1, which leads to a decrease in Rev-erb alpha and a subsequent increase in Per2.
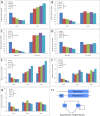
Figure 4. Unidirectional Paralog Compensation in the Circadian Repressors Cry1, Rev-erb beta, and Per1
(A, C, and E) U-2 OS cells were transfected with doses of siRNA as indicated, and gene expression was measured from samples collected 48 h after transfection. Knockdown of Cry1 (A), Rev-erb beta (C), and Per1 (E) leads to increased expression of the gene paralogs Cry2, Rev-erb alpha, and Per2 and Per3, respectively. (B, D, F, and G) The response is unidirectional, as a decrease of Cry2 (B), Rev-erb alpha (D), Per2 (F), or Per3 (G) does not result in an increase of respective gene paralogs. (H) Transcriptional repressors could directly regulate the expression of their gene paralog through response elements (RE) in upstream regulatory regions (H) (modified from [25]). In the case of the circadian clock network, we propose direct regulation of Cry1, Per1, and Rev-erb beta (Repressor 1) by Cry2, Per2/3, or Rev-erb alpha (Repressor 2), respectively.
Figure 5. Compensation in Nonparalogous Genes May Contribute to Oscillator Robustness
(A) U-2 OS cells transfected with siRNAs targeting Bmal1 or Cry1, individually or in combination as indicated, were synchronized, and luminescence levels were measured over 5 d. Data are representative of three independent biological replicates. Note that two examples are shown for the siCry1/siBmal1 combination knockdown. (B) Median and ranges of amplitude following single or combinatoric perturbation. The amplitude of the circadian signal was estimated using continuous wavelet decomposition and averaged across three replicates in three independent experiments. The median and range of the fold changes relative to the negative siRNA control are plotted. Both individual siRNA treatments are significantly down-regulated relative to the negative siRNA controls (p < 0.05, Mann-Whitney test), and the siCry1/siBmal1 double knockdown is also significantly down-regulated relative to siCry1 (p < 0.05, Mann-Whitney test).
Similar articles
- MicroRNA-mediated regulation in the mammalian circadian rhythm.
Liu K, Wang R. Liu K, et al. J Theor Biol. 2012 Jul 7;304:103-10. doi: 10.1016/j.jtbi.2012.03.037. Epub 2012 Apr 8. J Theor Biol. 2012. PMID: 22554948 - High-throughput chemical screen identifies a novel potent modulator of cellular circadian rhythms and reveals CKIα as a clock regulatory kinase.
Hirota T, Lee JW, Lewis WG, Zhang EE, Breton G, Liu X, Garcia M, Peters EC, Etchegaray JP, Traver D, Schultz PG, Kay SA. Hirota T, et al. PLoS Biol. 2010 Dec 14;8(12):e1000559. doi: 10.1371/journal.pbio.1000559. PLoS Biol. 2010. PMID: 21179498 Free PMC article. - [Synchronization and genetic redundancy in circadian clocks].
Dardente H. Dardente H. Med Sci (Paris). 2008 Mar;24(3):270-6. doi: 10.1051/medsci/2008243270. Med Sci (Paris). 2008. PMID: 18334175 Review. French. - Invited review: regulation of mammalian circadian clock genes.
Albrecht U. Albrecht U. J Appl Physiol (1985). 2002 Mar;92(3):1348-55. doi: 10.1152/japplphysiol.00759.2001. J Appl Physiol (1985). 2002. PMID: 11842077 Review. - The mammalian circadian clock: a network of gene expression.
Albrecht U. Albrecht U. Front Biosci. 2004 Jan 1;9:48-55. doi: 10.2741/1196. Front Biosci. 2004. PMID: 14766343 Review.
Cited by
- Circadian regulation of olfaction and an evolutionarily conserved, nontranscriptional marker in Caenorhabditis elegans.
Olmedo M, O'Neill JS, Edgar RS, Valekunja UK, Reddy AB, Merrow M. Olmedo M, et al. Proc Natl Acad Sci U S A. 2012 Dec 11;109(50):20479-84. doi: 10.1073/pnas.1211705109. Epub 2012 Nov 26. Proc Natl Acad Sci U S A. 2012. PMID: 23185015 Free PMC article. - Protein sequestration versus Hill-type repression in circadian clock models.
Kim JK. Kim JK. IET Syst Biol. 2016 Aug;10(4):125-35. doi: 10.1049/iet-syb.2015.0090. IET Syst Biol. 2016. PMID: 27444022 Free PMC article. Review. - Systemic PPARγ deletion impairs circadian rhythms of behavior and metabolism.
Yang G, Jia Z, Aoyagi T, McClain D, Mortensen RM, Yang T. Yang G, et al. PLoS One. 2012;7(8):e38117. doi: 10.1371/journal.pone.0038117. Epub 2012 Aug 10. PLoS One. 2012. PMID: 22899986 Free PMC article. - Propofol improves sleep deprivation-induced sleep structural and cognitive deficits via upregulating the BMAL1 expression and suppressing microglial M1 polarization.
Liu H, Yang C, Wang X, Yu B, Han Y, Wang X, Wang Z, Zhang M, Wang H. Liu H, et al. CNS Neurosci Ther. 2024 Jul;30(7):e14798. doi: 10.1111/cns.14798. CNS Neurosci Ther. 2024. PMID: 39015099 Free PMC article. - Bayesian experts in exploring reaction kinetics of transcription circuits.
Yoshida R, Saito MM, Nagao H, Higuchi T. Yoshida R, et al. Bioinformatics. 2010 Sep 15;26(18):i589-95. doi: 10.1093/bioinformatics/btq389. Bioinformatics. 2010. PMID: 20823326 Free PMC article.
References
- Stratmann M, Schibler U. Properties, entrainment, and physiological functions of mammalian peripheral oscillators. J Biol Rhythms. 2006;21:494–506. - PubMed
- Ko CH, Takahashi JS. Molecular components of the mammalian circadian clock. Hum Mol Genet. 2006;15(Spec No 2):R271–277. - PubMed
- Reppert SM, Weaver DR. Coordination of circadian timing in mammals. Nature. 2002;418:935–941. - PubMed
- Waddington CH. Canalization of development and the inheritance of acquired characters. Nature. 1942;150:563–565. - PubMed
- Gallego M, Virshup DM. Post-translational modifications regulate the ticking of the circadian clock. Nat Rev Mol Cell Biol. 2007;8:139–148. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 1 U54-RR-023567/RR/NCRR NIH HHS/United States
- R01 NS054794-02/NS/NINDS NIH HHS/United States
- F32 GM082083/GM/NIGMS NIH HHS/United States
- R01 NS054794/NS/NINDS NIH HHS/United States
- EY016807/EY/NEI NIH HHS/United States
- P50 MH074924/MH/NIMH NIH HHS/United States
- R01 EY016807/EY/NEI NIH HHS/United States
- 1 F32 GM082083-01/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases