Mitochondrial dysfunction leads to nuclear genome instability via an iron-sulfur cluster defect - PubMed (original) (raw)
Mitochondrial dysfunction leads to nuclear genome instability via an iron-sulfur cluster defect
Joshua R Veatch et al. Cell. 2009.
Abstract
Mutations and deletions in the mitochondrial genome (mtDNA), as well as instability of the nuclear genome, are involved in multiple human diseases. Here, we report that in Saccharomyces cerevisiae, loss of mtDNA leads to nuclear genome instability, through a process of cell-cycle arrest and selection we define as a cellular crisis. This crisis is not mediated by the absence of respiration, but instead correlates with a reduction in the mitochondrial membrane potential. Analysis of cells undergoing this crisis identified a defect in iron-sulfur cluster (ISC) biogenesis, which requires normal mitochondrial function. We found that downregulation of nonmitochondrial ISC protein biogenesis was sufficient to cause increased genomic instability in cells with intact mitochondrial function. These results suggest mitochondrial dysfunction stimulates nuclear genome instability by inhibiting the production of ISC-containing protein(s), which are required for maintenance of nuclear genome integrity. For a video summary of this article, see the PaperFlick file available with the online Supplemental Data.
Figures
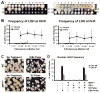
Figure 1. Loss or damage to the mtDNA results in nuclear genomic instability
(A-B) Forty mother cells were analyzed by pedigree analysis. (A) left: Colonies produced by the 1st through 37th daughter of a single representative mother cell are shown (daughter 13 did not produce a colony). Right: The colonies from the left were replicated to media containing lead nitrate, which permits colony color development. Chromosome IV LOH results in a brown color, and chromosome XII LOH results in a red color. (B) The fraction of colonies that had at least a 1/8 sector of colony color are presented as a function of the number of cell divisions the mother cell went through. The data were classified as to whether the colony was normal or petite. Data are shown for LOH events that occurred on either the right arm of chromosome XII or IV. The difference in the LOH frequency between petite and normal cells for divisions 11-30 had a significance of p<0.0001 from chromosome XII and p<0.001 for chromosome IV. (C) A wild-type (WT) strain was treated with or without a transient pulse of ethidium bromide (EtBr), and colonies grown from these cells were transferred to medium allowing visualization of LOH events. A derivative of the wild-type strain containing the dominant negative PGAL1-MIP1DN allele under control of the Gal4-EBD-VP16 (GEV) fusion protein was treated with or without a transient pulse of estradiol (E2), and grown as described above. (D) The treatments described in C were carried out on the WT strain or derivatives of this strain that contain the GEV activator, and/or the dominant negative PGAL1-MIP1DN as indicated. The percentage of colonies showing at least a 1/8 sector of LOH on chromosome XII and IV are presented. All error bars represent 95% confidence intervals and were calculated using the Poisson distribution.
Figure 2. Loss of the mtDNA leads to a progressive decline in growth rate, cell cycle arrest, and selection for suppressor clones
(A) Growth rate of isogenic strains containing the GEV transcriptional activator, PGAL1-MIP1DN or both, following transient treatment with estradiol (E2). (B) Growth rate of the wild-type strain treated with or without a transient pulse of ethidium bromide. (C) The percentage of cells from (A) and (B) able to form a colony of at least 1mm in diameter after 8 days is presented on the left. The percentage of cells arrested as unbudded cells is presented on the right. Error bars represent 95% confidence intervals calculated using the binomial distribution. (D) Spontaneous [ρ0] haploid suppressor mutants were back-crossed to isogenic cells with intact mtDNA. After sporulation, haploid [ρ+] progeny of these crosses containing each suppressor allele were treated transiently with ethidium bromide and the growth rate was monitored as before.
Figure 3. The absence of respiration is not sufficient to cause the crisis and LOH
(A) Wild-type cells (UCC1899) and derivatives with homozygous deletions for either COX4, RIP1 or CAT5 were treated as described in Figure 2A. (B-D) Growth rate of cells with or without transient treatment with ethidium bromide. Wild-type cells are compared to deletions of CAT5 (B), RIP1 (C) or COX4 (D).
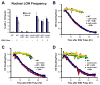
Figure 4. A hyperactive F1 ATP synthase partially restores mitochondrial protein import and prevents the growth defect and nuclear genomic instability that results from mtDNA loss
(A) Growth rates of a wild-type diploid and a the ATP1-111/ATP1-111 derivative, with or without transient ethidium bromide treatment. (B) The frequency of LOH events in the strains from (A). (C) Localization of proteins to the outer mitochondrial membrane (Tom70-GFP, green) or the mitochondrial matrix (preCox4-Cherry, red) in wild-type cells or ATP1-111/ATP1-111 derivative cells are presented.
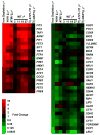
Figure 5. Cells show a transcriptional signature of iron starvation following mtDNA loss
Levels of mRNA from WT cells 3, 11, 19 and 27 hours after mtDNA loss were compared to cells with intact mtDNA. mRNA from ATP1-111 mutants collected 27 hours after mtDNA loss was compared to mRNA of the same strain with intact mtDNA. mRNA was isolated from the NAR1 repressible strain 27 hours after a shift from 50 nM estradiol to 2 nM estradiol and compared to the same strain grown in 50 nM estradiol. Genes are arranged in order of their reported magnitude of induction (red) or repression (green) by iron starvation (Puig et al., 2005). Comparisons are made to published array data for depletion of the YAH1 and ATM1 genes (Hausmann et al., 2008).
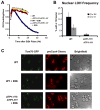
Figure 6. Reducing ISC protein loading activity in the cytoplasm leads to nuclear genome instability
(A) A strain (UCC3970) with NAR1 expression controlled by estradiol (E2) was plated to media containing either 50 nM (NAR1 expressed) or 2 nM estradiol (NAR1 down-regulated) followed by replication to LOH indicator media. (B) The frequency of colonies showing at least 1/8 sector of LOH following plating of the strain in A (NAR1EST) and an isogenic strain without the estradiol regulated promoter (WT) to either low (2nM) or high (50nM) estradiol (E2). Error bars represent 95% confidence intervals calculated using the Poisson distribution.
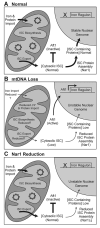
Figure 7. Model to explain nuclear genomic instability following mtDNA loss
A) In normal cells, iron is packaged into iron-sulfur clusters (ISCs) in the mitochondria, and some ISCs are exported into the cytoplasm. The cytoplasmic ISCs inhibit iron regulon induction and are required for the function of proteins that maintain nuclear genome integrity. (B) Upon loss of the mtDNA, the mitochondrial membrane potential (Δψ) is reduced, which compromises mitochondrial iron import, ISC packaging, and/or export. This results in iron regulon induction and nuclear genome instability. (C) Artificially reduced Nar1 function limits packaging of ISCs into proteins involved in nuclear DNA metabolism, leading to nuclear genome instability. In this case mitochondrial function is intact and the iron regulon is not induced.
Comment in
- Ironing out a midlife crisis.
Vergara SV, Thiele DJ. Vergara SV, et al. Cell. 2009 Jun 26;137(7):1179-81. doi: 10.1016/j.cell.2009.06.005. Cell. 2009. PMID: 19563748
Similar articles
- Depletion of thiol reducing capacity impairs cytosolic but not mitochondrial iron-sulfur protein assembly machineries.
Braymer JJ, Stümpfig M, Thelen S, Mühlenhoff U, Lill R. Braymer JJ, et al. Biochim Biophys Acta Mol Cell Res. 2019 Feb;1866(2):240-251. doi: 10.1016/j.bbamcr.2018.11.003. Epub 2018 Nov 10. Biochim Biophys Acta Mol Cell Res. 2019. PMID: 30419257 - Zim17/Tim15 links mitochondrial iron-sulfur cluster biosynthesis to nuclear genome stability.
Díaz de la Loza Mdel C, Gallardo M, García-Rubio ML, Izquierdo A, Herrero E, Aguilera A, Wellinger RE. Díaz de la Loza Mdel C, et al. Nucleic Acids Res. 2011 Aug;39(14):6002-15. doi: 10.1093/nar/gkr193. Epub 2011 Apr 21. Nucleic Acids Res. 2011. PMID: 21511814 Free PMC article. - Mitochondria-nucleus network for genome stability.
Kaniak-Golik A, Skoneczna A. Kaniak-Golik A, et al. Free Radic Biol Med. 2015 May;82:73-104. doi: 10.1016/j.freeradbiomed.2015.01.013. Epub 2015 Jan 30. Free Radic Biol Med. 2015. PMID: 25640729 Review. - Mechanisms of iron-sulfur protein maturation in mitochondria, cytosol and nucleus of eukaryotes.
Lill R, Dutkiewicz R, Elsässer HP, Hausmann A, Netz DJ, Pierik AJ, Stehling O, Urzica E, Mühlenhoff U. Lill R, et al. Biochim Biophys Acta. 2006 Jul;1763(7):652-67. doi: 10.1016/j.bbamcr.2006.05.011. Epub 2006 May 23. Biochim Biophys Acta. 2006. PMID: 16843540 Review. - Mechanisms of Mitochondrial Iron-Sulfur Protein Biogenesis.
Lill R, Freibert SA. Lill R, et al. Annu Rev Biochem. 2020 Jun 20;89:471-499. doi: 10.1146/annurev-biochem-013118-111540. Epub 2020 Jan 14. Annu Rev Biochem. 2020. PMID: 31935115 Review.
Cited by
- Human DNA polymerase ε is phosphorylated at serine-1940 after DNA damage and interacts with the iron-sulfur complex chaperones CIAO1 and MMS19.
Moiseeva TN, Gamper AM, Hood BL, Conrads TP, Bakkenist CJ. Moiseeva TN, et al. DNA Repair (Amst). 2016 Jul;43:9-17. doi: 10.1016/j.dnarep.2016.04.007. Epub 2016 May 7. DNA Repair (Amst). 2016. PMID: 27235625 Free PMC article. - Mechanisms by which different functional states of mitochondria define yeast longevity.
Beach A, Leonov A, Arlia-Ciommo A, Svistkova V, Lutchman V, Titorenko VI. Beach A, et al. Int J Mol Sci. 2015 Mar 11;16(3):5528-54. doi: 10.3390/ijms16035528. Int J Mol Sci. 2015. PMID: 25768339 Free PMC article. Review. - Iron-sulfur cluster synthesis, iron homeostasis and oxidative stress in Friedreich ataxia.
Vaubel RA, Isaya G. Vaubel RA, et al. Mol Cell Neurosci. 2013 Jul;55:50-61. doi: 10.1016/j.mcn.2012.08.003. Epub 2012 Aug 11. Mol Cell Neurosci. 2013. PMID: 22917739 Free PMC article. Review. - Cell organelles and yeast longevity: an intertwined regulation.
Banerjee R, Joshi N, Nagotu S. Banerjee R, et al. Curr Genet. 2020 Feb;66(1):15-41. doi: 10.1007/s00294-019-01035-0. Epub 2019 Sep 18. Curr Genet. 2020. PMID: 31535186 Review. - The good and the bad of being connected: the integrons of aging.
Dillin A, Gottschling DE, Nyström T. Dillin A, et al. Curr Opin Cell Biol. 2014 Feb;26:107-12. doi: 10.1016/j.ceb.2013.12.003. Epub 2013 Dec 30. Curr Opin Cell Biol. 2014. PMID: 24529252 Free PMC article. Review.
References
- Alseth I, Eide L, Pirovano M, Rognes T, Seeberg E, Bjoras M. The Saccharomyces cerevisiae homologues of endonuclease III from Escherichia coli, Ntg1 and Ntg2, are both required for efficient repair of spontaneous and induced oxidative DNA damage in yeast. Mol Cell Biol. 1999;19:3779–3787. - PMC - PubMed
- Baker KP, Schatz G. Mitochondrial proteins essential for viability mediate protein import into yeast mitochondria. Nature. 1991;349:205–208. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 AG023779/AG/NIA NIH HHS/United States
- T32 AG00057/AG/NIA NIH HHS/United States
- AG023779/AG/NIA NIH HHS/United States
- T32 AG000057/AG/NIA NIH HHS/United States
- R37 AG023779/AG/NIA NIH HHS/United States
- GM43893/GM/NIGMS NIH HHS/United States
- R01 GM043893/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases